Abstract
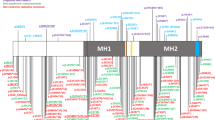
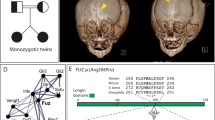
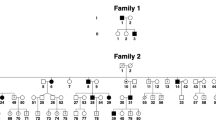
Crouzon syndrome is an autosomal dominant condition causing premature fusion of the cranial sutures (craniosynostosis) and maps to chromosome 10q25–q26. We now present evidence that mutations in the fibroblast growth factor receptor 2 gene (FGFR2) cause Crouzon syndrome. We found SSCP variations in the B exon of FGFR2 in nine unrelated affected individuals as well as complete cosegregation between SSCP variation and disease in three unrelated multigenerational families. In four sporadic cases, the normal parents did not have SSCP variation. Finally, direct sequencing has revealed specific mutations in the B exon in all nine sporadic and familial cases, including replacement of a cysteine in an immunoglobulin-like domain in five patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Cohen, M.M.Jr, Craniosynostosis: diagnosis, evaluation and management. (Raven Press, New York, 1986).
Winter, R.M. & Baraitser, M. The London Dysmorphology Database. (Oxford University Press, Oxford, 1994).
Warman, M.L., Mulliken, J.B., Hayward, P.G. & Muller, U. Newly recognised autosomal dominant craniosynostotic syndrome. Am. J. med. Genet. 46, 444–449 (1993).
Muller, U., Warman, M.L., Mulliken, J.B. & Weber, J. Assignment of a gene locus Involved in craniosynostosis to chromosome 5qter. Hum. molec. Genet. 2, 119–122 (1992).
Jabs, E.W. et al. Mutation in the Homeodomain of the Human MSX2 gene in a family affected with autosomal dominant craniosynostosis. Cell 75, 443–450 (1993).
Brueton, L.A., van Herwerden, L., Chotai, K.A. & Winter, R.M. The mapping of a gene for craniosynostosis: evidence for linkage of the Saethre-Chotzen syndrome to distal chromosome 7p. J. med. Genet. 29, 681–685 (1992).
van Herwerden, L. et al. Evidence for locus heterogeneity in acrocephalosyndactyly: a refined localization for the Saethre-Chotzen syndrome locus on distal chromosome 7p — and exclusion of Jackson-Weiss syndrome from craniosynostosis loci on 7p & 5q. Am. J. hum. Genet. 54, 669–674 (1994).
Lewanda, A.F. et al. Genetic Heterogeneity among craniosynostosis syndromes: mapping the Saethre-Chotzen syndrome locus between D7S513 & D7S516 and exclusion of Jackson-Weiss & Crouzon syndrome loci from 7p. Genomics 19, 115–119 (1994).
Reardon, W., McManus, S.P., Summers, D. & Winter, R.M. Cytogenetic evidence that the Saethre-Chotzen gene maps to 7p21.2. Am. J. med. Genet. 47, 633–636 (1993).
Reid, C.S. et al. Saethre-Chotzen syndrome with familial translocation at chromosome 7p22. Am. J. med. Genet. 47, 637–639 (1993).
Reardon, W., van Herwerden, L., Rose, C., Jones, B., Malcolm, S. & Winter, R.M. Crouzon syndrome is not linked to craniosynostosis loci at 7p & 5qter. J. med. Genet. 31, 219–221 (1994).
Preston, R.A. et al. A gene for Crouzon craniofacial dysostosis maps to the long arm of chromosome 10. Nature Genet. 7, 149–153 (1994).
Johnson, D.E., Lee, P.L., Lu, J. & Williams, L.T. Diverse forms of a receptor for acidic & basic fibroblast growth factors. Molec. Cell. Genet. 10, 4728–4736 (1990).
Kombluth, S., Paulson, K.E. & Hanafusa, H. Novel Tyrosine Kinase identified by Phosphotyrosine antibody screening of cDNA libraries. Molec. Cell. Biol. 8, 5541–5544 (1988).
Lee, P.L., Johnson, D.E., Cousens, L.S., Fried, V.A. & Williams, L.T. Purification and complementary DNA sequence of areceptor for basic Fibroblast Growth Factor. Science 245, 57–60 (1989).
Ruta, M. et al. Receptor for acidic fibroblast growth factor is related to the tyrosine klnase encoded by the fms-like gene (FLG). Proc. natn. Acad. Sci. U.S.A. 86, 8722–8726 (1989).
Reid, H.H., Wilks, A.F. & Bernard, O. Two forms of basic fibroblast growth factor receptor-like mRNA are expressed in the developing mouse brain. Proc. natn. Acad. Sci. U.S.A 87, 1596–1600 (1990).
Mansukhani, A., Moscatelli, D., Talarico, D., Levytska, V. & Basilico, C. A murine fibroblast growth factor (FGF) receptor expressed in CHO Cells is activated by basis FGF & Kaposi FGF. Pro. natn. Acad. Sci. U.S.A. 87, 4378–4382 (1990).
Dionne, C.A. et al. Cloning & expression of two distinct high-affinity receptors cross-reacting with acidic & basic fibroblast growth factors. EMBO J. 8, 2685–2692 (1990).
Houssaint, E. et al. Related fibroblast growth factor receptor genes exist in the human genome. Proc. natn. Acad. Sci. U.S.A. 87, 8180–8184 (1990).
Pasquale, E.B. A distinctive family of embryonic protein-tyroslne kinase receptors. Proc. natn. Acad. Sci. U.S.A. 87, 5812–5816 (1990).
Raz, V., Keiman, Z., Avivi, A., Neufeld, G., Givol, D. & Yarden, Y. PCR-Based identification of new receptors — molecular cloning of a receptor for Fibroblast growth factors. Oncogens 6, 753–760 (1991).
Keegan, K., Johnson, D.E., Williams, L.T. & Hayman, M.J. Isolation of an additional member of the fibroblast growth factor receptor family, FGFR-3. Proc. natn. Acad. Sci. U.S.A 88, 1095–1099 (1993).
Partanen, J. et al. FGFR-4, a novel acidic fibroblast growth factor receptor with a distinct expression pattern. EMBO J. 1347–1354 (1991).
Johnson, D.E., Lu, J., Chen, H., Werner, S. & Williams, L.T. The Human Fibroblast Growth Factor Genes: acommon structural arrangement underlies the mechanisms for generating receptor forms that differ in their third immunoglobulin domain. Molec. Cell. Biol. 11, 4627–4634 (1991).
Miki, T. et al. Determination of ligand-binding specificity by alternative splicing: Two distinct growth factor receptors encoded by a single gene. Proc. natn. Acad. Sci. U.S.A. 89, 246–250 (1992).
Shiang, R. et al. Mutations in the transmembrane domain of FGFR3 cause the most common genetic form of dwarflsm, Achondroplasia. Cell 78, 335–342 (1994).
Mattei, M-G., Moreau, A., Gesnel, M-C., Houssaint, E. & Breathnach, R. Assignment by in situ hybridization of a fibroblast growth factor receptor gene to human chromosome band 10q26. Hum. Genet. 87, 84–86 (1991).
Dionne, C.A., Modi, W.S., Crumley, G., O'Brien, S.J., Schlessinger, J. & Jaye, M. BEK, a receptor for multiple members of the fibroblast growth factor (FGF) family, maps to human chromosome 10q25.3–q26. Cytogenet Cell Genet. 60, 34–36 (1992).
Gilbert, E., Del Gatto, F., Champion-Amaud, P., Gesnel, M-C. & Breathnach, R. Control of Bek and K-Sam splice sites in alternative splicing of the fibroblast growth factor receptor 2 pre-mRNA. Molec. Cell. Biol. 13, 5461–5468 (1993).
Orr-Uretreger, A. et al. Developmental localization of the splicing alternatives of fibroblast growth factor receptor-2 (FGFR-2). Devel. Biol. 158, 475–486 (1993).
Orr-Uretreger, A., Givol, D., Yayon, A., & Lonai, P. Developmental expression of two murine fibroblast growth factor receptors, fig & bek. Development 113, 1419–1434 (1991).
Shapiro, M.B. & Senepathy, P. RNA splice junctions of different classes of eukaryotes: sequence statistics & fundamental implication in gene expression. Nucl. Acids Res. 15, 7155–7174 (1987).
Langer, L.O. et al. Thanatophoric Dysplasia & Cloverteaf Skull. Am. J. med. Genet. Suppl. 3, 167–179 (1987).
Miller, S.A., Dykes, D.D. & Polesky, H.F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucl. Acids Res. 16, 1215 (1988).
Orita, M., Iwahana, H., Kanazawa, M., Hayashi, K. & Sekiya, T. Detection of polymorphisms of human DNA by gel electrophoresis as single strand conformation polymorphisms. Proc. natn. Acad. Sci. U.S.A. 86, 2766–2770 (1987).
Lathrop, G.M., Lalouel, J.M., Julier, C. & Ott, J. Strategies for multilocus linkage analysis In human. Proc. natn. Acad. Sci. U.S.A 81, 3443–3446 (1984).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Reardon, W., Winter, R., Rutland, P. et al. Mutations in the fibroblast growth factor receptor 2 gene cause Crouzon syndrome. Nat Genet 8, 98–103 (1994). https://doi.org/10.1038/ng0994-98
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0994-98
This article is cited by
-
Benchmarking splice variant prediction algorithms using massively parallel splicing assays
Genome Biology (2023)
-
Differential diagnosis of syndromic craniosynostosis: a case series
Archives of Gynecology and Obstetrics (2022)
-
Adapting SureSelect enrichment protocol to the Ion Torrent S5 platform in molecular diagnostics of craniosynostosis
Scientific Reports (2020)
-
The duality of human oncoproteins: drivers of cancer and congenital disorders
Nature Reviews Cancer (2020)