Abstract
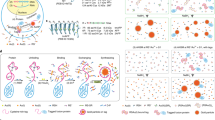
Fluorescence microscopy is an essential tool for the exploration of cell growth, division, transcription and translation in eukaryotes and prokaryotes alike. Despite the rapid development of techniques to study bacteria, the size of these organisms (1–10 μm) and their robust and largely impenetrable cell envelope present major challenges in imaging experiments. Fusion-based strategies, such as attachment of the protein of interest to a fluorescent protein or epitope tag, are by far the most common means for examining protein localization and expression in prokaryotes. While valuable, the use of genetically encoded tags can result in mislocalization or altered activity of the desired protein, does not provide a readout of the catalytic state of enzymes and cannot enable visualization of many other important cellular components, such as peptidoglycan, lipids, nucleic acids or glycans. Here, we highlight the use of biomolecule-specific small-molecule probes for imaging in bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Shapiro, L., McAdams, H.H. & Losick, R. Why and how bacteria localize proteins. Science 326, 1225–1228 (2009).
Rudner, D.Z. & Losick, R. Protein subcellular localization in bacteria. Cold Spring Harb. Perspect. Biol. 2, a000307 (2010).
Feilmeier, B.J., Iseminger, G., Schroeder, D., Webber, H. & Phillips, G.J. Green fluorescent protein functions as a reporter for protein localization in Escherichia coli. J. Bacteriol. 182, 4068–4076 (2000).
Jarvik, J.W. & Telmer, C.A. Epitope tagging. Annu. Rev. Genet. 32, 601–618 (1998).
Brizzard, B. Epitope tagging. Biotechniques 44, 693–695 (2008).
Dean, K.M. & Palmer, A.E. Advances in fluorescence labeling strategies for dynamic cellular imaging. Nat. Chem. Biol. 10, 512–523 (2014). An overview of the improvements made in fluorescence labeling approaches.
Griffin, B.A., Adams, S.R. & Tsien, R.Y. Specific covalent labeling of recombinant protein molecules inside live cells. Science 281, 269–272 (1998).
Zhang, J., Campbell, R.E., Ting, A.Y. & Tsien, R.Y. Creating new fluorescent probes for cell biology. Nat. Rev. Mol. Cell Biol. 3, 906–918 (2002).
Lang, K. & Chin, J.W. Cellular incorporation of unnatural amino acids and bioorthogonal labeling of proteins. Chem. Rev. 114, 4764–4806 (2014).
Landgraf, D., Okumus, B., Chien, P., Baker, T.A. & Paulsson, J. Segregation of molecules at cell division reveals native protein localization. Nat. Methods 9, 480–482 (2012).
Margolin, W. The price of tags in protein localization studies. J. Bacteriol. 194, 6369–6371 (2012).
Domínguez-Escobar, J. et al. Processive movement of MreB-associated cell wall biosynthetic complexes in bacteria. Science 333, 225–228 (2011).
Garner, E.C. et al. Coupled, circumferential motions of the cell wall synthesis machinery and MreB filaments in B. subtilis. Science 333, 222–225 (2011).
Swulius, M.T. & Jensen, G.J. The helical MreB cytoskeleton in Escherichia coli MC1000/pLE7 is an artifact of the N-terminal yellow fluorescent protein tag. J. Bacteriol. 194, 6382–6386 (2012).
Huang, B., Bates, M. & Zhuang, X. Super-resolution fluorescence microscopy. Annu. Rev. Biochem. 78, 993–1016 (2009).
Koch, A.L. What size should a bacterium be? A question of scale. Annu. Rev. Microbiol. 50, 317–348 (1996).
Erickson, H.P., Taylor, D.W., Taylor, K.A. & Bramhill, D. Bacterial cell division protein FtsZ assembles into protofilament sheets and minirings, structural homologs of tubulin polymers. Proc. Natl. Acad. Sci. USA 93, 519–523 (1996).
Fu, G. et al. In vivo structure of the E. coli FtsZ-ring revealed by photoactivated localization microscopy (PALM). PLoS One 5, e12682 (2010).
Buss, J. et al. In vivo organization of the FtsZ-ring by ZapA and ZapB revealed by quantitative super-resolution microscopy. Mol. Microbiol. 89, 1099–1120 (2013).
Holden, S.J. et al. High throughput 3D super-resolution microscopy reveals Caulobacter crescentus in vivo Z-ring organization. Proc. Natl. Acad. Sci. USA 111, 4566–4571 (2014).
Strauss, M.P. et al. 3D-SIM super resolution microscopy reveals a bead-like arrangement for FtsZ and the division machinery: implications for triggering cytokinesis. PLoS Biol. 10, e1001389 (2012).
Chang, P.V. & Bertozzi, C.R. Imaging beyond the proteome. Chem. Commun. (Camb.) 48, 8864–8879 (2012).
Stracy, M., Uphoff, S., Garza de Leon, F. & Kapanidis, A.N. In vivo single-molecule imaging of bacterial DNA replication, transcription, and repair. FEBS Lett. 588, 3585–3594 (2014).
Song, W., Strack, R.L. & Jaffrey, S.R. Imaging bacterial protein expression using genetically encoded RNA sensors. Nat. Methods 10, 873–875 (2013).
Schimak, M.P. et al. MiL-FISH: Multilabeled oligonucleotides for fluorescence in situ hybridization improve visualization of bacterial cells. Appl. Environ. Microbiol. 82, 62–70 (2016).
Spahn, C., Endesfelder, U. & Heilemann, M. Super-resolution imaging of Escherichia coli nucleoids reveals highly structured and asymmetric segregation during fast growth. J. Struct. Biol. 185, 243–249 (2014).
Schoen, I., Ries, J., Klotzsch, E., Ewers, H. & Vogel, V. Binding-activated localization microscopy of DNA structures. Nano Lett. 11, 4008–4011 (2011).
Chan, J., Dodani, S.C. & Chang, C.J. Reaction-based small-molecule fluorescent probes for chemoselective bioimaging. Nat. Chem. 4, 973–984 (2012).
Vollmer, W., Blanot, D. & de Pedro, M.A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 (2008).
Vollmer, W. & Seligman, S.J. Architecture of peptidoglycan: more data and more models. Trends Microbiol. 18, 59–66 (2010).
Matias, V.R. & Beveridge, T.J. Cryo-electron microscopy reveals native polymeric cell wall structure in Bacillus subtilis 168 and the existence of a periplasmic space. Mol. Microbiol. 56, 240–251 (2005).
Bugg, T.D., Braddick, D., Dowson, C.G. & Roper, D.I. Bacterial cell wall assembly: still an attractive antibacterial target. Trends Biotechnol. 29, 167–173 (2011).
Typas, A., Banzhaf, M., Gross, C.A. & Vollmer, W. From the regulation of peptidoglycan synthesis to bacterial growth and morphology. Nat. Rev. Microbiol. 10, 123–136 (2012).
Lovering, A.L., Safadi, S.S. & Strynadka, N.C. Structural perspective of peptidoglycan biosynthesis and assembly. Annu. Rev. Biochem. 81, 451–478 (2012).
Tipper, D.J. & Strominger, J.L. Mechanism of action of penicillins: a proposal based on their structural similarity to acyl-D-alanyl-D-alanine. Proc. Natl. Acad. Sci. USA 54, 1133–1141 (1965).
Blumberg, P.M., Yocum, R.R., Willoughby, E. & Strominger, J.L. Binding of [14C]penicillin G to the membrane-bound and the purified D-alanine carboxypeptidases from Bacillus stearothermophilus and Bacillus subtilis and its release. J. Biol. Chem. 249, 6828–6835 (1974).
Zhao, G., Meier, T.I., Kahl, S.D., Gee, K.R. & Blaszczak, L.C. BOCILLIN FL, a sensitive and commercially available reagent for detection of penicillin-binding proteins. Antimicrob. Agents Chemother. 43, 1124–1128 (1999).
Gee, K.R., Kang, H.C., Meier, T.I., Zhao, G. & Blaszcak, L.C. Fluorescent bocillins: synthesis and application in the detection of penicillin-binding proteins. Electrophoresis 22, 960–965 (2001).
Heal, W.P. & Tate, E.W. Application of activity-based protein profiling to the study of microbial pathogenesis. Top. Curr. Chem. 324, 115–135 (2012).
Puri, A.W. & Bogyo, M. Applications of small molecule probes in dissecting mechanisms of bacterial virulence and host responses. Biochemistry 52, 5985–5996 (2013).
Staub, I. & Sieber, S.A. Beta-lactams as selective chemical probes for the in vivo labeling of bacterial enzymes involved in cell wall biosynthesis, antibiotic resistance, and virulence. J. Am. Chem. Soc. 130, 13400–13409 (2008).
Böttcher, T. & Sieber, S.A. Beta-lactones as specific inhibitors of ClpP attenuate the production of extracellular virulence factors of Staphylococcus aureus. J. Am. Chem. Soc. 130, 14400–14401 (2008).
Böttcher, T. & Sieber, S.A. Structurally refined beta-lactones as potent inhibitors of devastating bacterial virulence factors. ChemBioChem 10, 663–666 (2009).
Zeiler, E., Korotkov, V.S., Lorenz-Baath, K., Böttcher, T. & Sieber, S.A. Development and characterization of improved β-lactone-based anti-virulence drugs targeting ClpP. Bioorg. Med. Chem. 20, 583–591 (2012).
Kocaoglu, O. et al. Selective penicillin-binding protein imaging probes reveal substructure in bacterial cell division. ACS Chem. Biol. 7, 1746–1753 (2012). First example of selective PBP imaging using an activity-based probe. This study provides precedent for the use of β-lactam antibiotics to facilitate microscopy-based study of the PBPs.
Kocaoglu, O. & Carlson, E.E. Penicillin-binding protein imaging probes. Curr. Protoc. Chem. Biol. 5, 239–250 (2013).
Kocaoglu, O. & Carlson, E.E. Profiling of β-lactam selectivity for penicillin-binding proteins in Escherichia coli strain DC2. Antimicrob. Agents Chemother. 59, 2785–2790 (2015).
Kocaoglu, O., Tsui, H.-C.T., Winkler, M.E. & Carlson, E.E. Profiling of β-lactam selectivity for penicillin-binding proteins in Streptococcus pneumoniae D39. Antimicrob. Agents Chemother. 59, 3548–3555 (2015).
Daniel, R.A. & Errington, J. Control of cell morphogenesis in bacteria: two distinct ways to make a rod-shaped cell. Cell 113, 767–776 (2003). Beautiful demonstration of the utility of vancomycin as an imaging agent to visualize nascent peptidoglycan.
Tiyanont, K. et al. Imaging peptidoglycan biosynthesis in Bacillus subtilis with fluorescent antibiotics. Proc. Natl. Acad. Sci. USA 103, 11033–11038 (2006).
Gautam, S. et al. An activity-based probe for studying crosslinking in live bacteria. Angew. Chem. Int. Edn Engl. 54, 10492–10496 (2015).
Liu, H., Sadamoto, R., Sears, P.S. & Wong, C.H. An efficient chemoenzymatic strategy for the synthesis of wild-type and vancomycin-resistant bacterial cell-wall precursors: UDP-N-acetylmuramyl-peptides. J. Am. Chem. Soc. 123, 9916–9917 (2001).
Sadamoto, R. et al. Cell-wall engineering of living bacteria. J. Am. Chem. Soc. 124, 9018–9019 (2002).
Olrichs, N.K. et al. A novel in vivo cell-wall labeling approach sheds new light on peptidoglycan synthesis in Escherichia coli. ChemBioChem 12, 1124–1133 (2011).
Kuru, E. et al. In Situ probing of newly synthesized peptidoglycan in live bacteria with fluorescent D-amino acids. Angew. Chem. Int. Edn Engl. 51, 12519–12523 (2012).
Siegrist, M.S. et al. D-Amino acid chemical reporters reveal peptidoglycan dynamics of an intracellular pathogen. ACS Chem. Biol. 8, 500–505 (2013).
Shieh, P., Siegrist, M.S., Cullen, A.J. & Bertozzi, C.R. Imaging bacterial peptidoglycan with near-infrared fluorogenic azide probes. Proc. Natl. Acad. Sci. USA 111, 5456–5461 (2014).
Lebar, M.D. et al. Reconstitution of peptidoglycan cross-linking leads to improved fluorescent probes of cell wall synthesis. J. Am. Chem. Soc. 136, 10874–10877 (2014).
Pidgeon, S.E. et al. Metabolic profiling of bacteria by unnatural C-terminated D-amino acids. Angew. Chem. Int. Edn Engl. 54, 6158–6162 (2015).
Schirner, K. et al. Lipid-linked cell wall precursors regulate membrane association of bacterial actin MreB. Nat. Chem. Biol. 11, 38–45 (2015).
Monteiro, J.M. et al. Cell shape dynamics during the staphylococcal cell cycle. Nat. Commun. 6, 8055 (2015).
Garrett, A.J., Harrison, M.J. & Manire, G.P. A search for the bacterial mucopeptide component, muramic acid, in Chlamydia. J. Gen. Microbiol. 80, 315–318 (1974).
Fox, A. et al. Muramic acid is not detectable in Chlamydia psittaci or Chlamydia trachomatis by gas chromatography-mass spectrometry. Infect. Immun. 58, 835–837 (1990).
Pilhofer, M. et al. Discovery of chlamydial peptidoglycan reveals bacteria with murein sacculi but without FtsZ. Nat. Commun. 4, 2856 (2013).
Liechti, G.W. et al. A new metabolic cell-wall labelling method reveals peptidoglycan in Chlamydia trachomatis. Nature 506, 507–510 (2014). Demonstration of the presence of peptidoglycan in Chlamydia trachomatis using fluorescent D -amino acids for the first time.
Wheeler, R., Mesnage, S., Boneca, I.G., Hobbs, J.K. & Foster, S.J. Super-resolution microscopy reveals cell wall dynamics and peptidoglycan architecture in ovococcal bacteria. Mol. Microbiol. 82, 1096–1109 (2011).
Tsui, H.C. et al. Pbp2x localizes separately from Pbp2b and other peptidoglycan synthesis proteins during later stages of cell division of Streptococcus pneumoniae D39. Mol. Microbiol. 94, 21–40 (2014). Beautiful combination of FDAAs, Van-FL, β-lactams and fusion proteins to explore PBP localization during division.
Weidenmaier, C. et al. Lack of wall teichoic acids in Staphylococcus aureus leads to reduced interactions with endothelial cells and to attenuated virulence in a rabbit model of endocarditis. J. Infect. Dis. 191, 1771–1777 (2005).
Tra, V.N. & Dube, D.H. Glycans in pathogenic bacteria—potential for targeted covalent therapeutics and imaging agents. Chem. Commun. (Camb.) 50, 4659–4673 (2014).
Woodruff, P.J. et al. Trehalose is required for growth of Mycobacterium smegmatis. J. Biol. Chem. 279, 28835–28843 (2004).
Backus, K.M. et al. Uptake of unnatural trehalose analogs as a reporter for Mycobacterium tuberculosis. Nat. Chem. Biol. 7, 228–235 (2011). Describes the use of fluorescent unnatural trehalose for sensitive detection of Mycobacterium tuberculosis in mammalian cells.
Swarts, B.M. et al. Probing the mycobacterial trehalome with bioorthogonal chemistry. J. Am. Chem. Soc. 134, 16123–16126 (2012).
Urbanek, B.L. et al. Chemoenzymatic synthesis of trehalose analogues: rapid access to chemical probes for investigating mycobacteria. ChemBioChem 15, 2066–2070 (2014).
Dumont, A., Malleron, A., Awwad, M., Dukan, S. & Vauzeilles, B. Click-mediated labeling of bacterial membranes through metabolic modification of the lipopolysaccharide inner core. Angew. Chem. Int. Edn Engl. 51, 3143–3146 (2012).
Lee, M.K., Rai, P., Williams, J., Twieg, R.J. & Moerner, W.E. Small-molecule labeling of live cell surfaces for three-dimensional super-resolution microscopy. J. Am. Chem. Soc. 136, 14003–14006 (2014).
Gunsolus, I.L. et al. Facile method to stain the bacterial cell surface for super-resolution fluorescence microscopy. Analyst 139, 3174–3178 (2014).
Conley, N.R., Biteen, J.S. & Moerner, W.E. Cy3-Cy5 covalent heterodimers for single-molecule photoswitching. J. Phys. Chem. B 112, 11878–11880 (2008).
Nelson, J.W. et al. A biosynthetic strategy for re-engineering the Staphylococcus aureus cell wall with non-native small molecules. ACS Chem. Biol. 5, 1147–1155 (2010).
Wang, L., Brock, A., Herberich, B. & Schultz, P.G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Charbon, G. et al. Subcellular protein localization by using a genetically encoded fluorescent amino acid. ChemBioChem 12, 1818–1821 (2011).
Beatty, K.E., Xie, F., Wang, Q. & Tirrell, D.A. Selective dye-labeling of newly synthesized proteins in bacterial cells. J. Am. Chem. Soc. 127, 14150–14151 (2005).
Hatzenpichler, R. et al. In situ visualization of newly synthesized proteins in environmental microbes using amino acid tagging and click chemistry. Environ. Microbiol. 16, 2568–2590 (2014).
Mahdavi, A. et al. Identification of secreted bacterial proteins by noncanonical amino acid tagging. Proc. Natl. Acad. Sci. USA 111, 433–438 (2014).
Raulf, A. et al. Click chemistry facilitates direct labeling and super-resolution imaging of nucleic acids and proteins. RCS Adv. 4, 30462–30466 (2014).
Beatty, K.E. et al. Sulfatase-activated fluorophores for rapid discrimination of mycobacterial species and strains. Proc. Natl. Acad. Sci. USA 110, 12911–12916 (2013).
Smith, E.L., Bertozzi, C.R. & Beatty, K.E. An expanded set of fluorogenic sulfatase activity probes. ChemBioChem 15, 1101–1105 (2014).
Gloeckl, S. et al. Identification of a serine protease inhibitor which causes inclusion vacuole reduction and is lethal to Chlamydia trachomatis. Mol. Microbiol. 89, 676–689 (2013).
Böttcher, T. & Sieber, S.A. Showdomycin as a versatile chemical tool for the detection of pathogenesis-associated enzymes in bacteria. J. Am. Chem. Soc. 132, 6964–6972 (2010).
Chauvigné-Hines, L.M. et al. Suite of activity-based probes for cellulose-degrading enzymes. J. Am. Chem. Soc. 134, 20521–20532 (2012).
Sadler, N.C. et al. Live cell chemical profiling of temporal redox dynamics in a photoautotrophic cyanobacterium. ACS Chem. Biol. 9, 291–300 (2014).
Deng, X. et al. Proteome-wide quantification and characterization of oxidation-sensitive cysteines in pathogenic bacteria. Cell Host Microbe 13, 358–370 (2013).
Wilke, K.E., Francis, S. & Carlson, E.E. Activity-based probe for histidine kinase signaling. J. Am. Chem. Soc. 134, 9150–9153 (2012).
Lee, M.K., Williams, J.C., Twieg, R.J., Rao, J. & Moerner, W.E. Enzymatic activation of nitro-aryl fluorogens in live bacterial cells for enzymatic turnover-activated localization microscopy. Chem. Sci. (Camb.) 4, 220–225 (2013).
Gustafsson, M.G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Hell, S.W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Schermelleh, L., Heintzmann, R. & Leonhardt, H. A guide to super-resolution fluorescence microscopy. J. Cell Biol. 190, 165–175 (2010).
Coltharp, C. & Xiao, J. Superresolution microscopy for microbiology. Cell. Microbiol. 14, 1808–1818 (2012). An excellent review comparing super-resolution microscopy techniques for bacterial cell imaging.
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Rust, M.J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Tuson, H.H. & Biteen, J.S. Unveiling the inner workings of live bacteria using super-resolution microscopy. Anal. Chem. 87, 42–63 (2015). Comprehensive review of cutting-edge strategies to examine bacteria using super-resolution microscopy techniques.
Acknowledgements
This work was supported by NIH DP2OD008592 (E.E.C.), a Pew Biomedical Scholar Award (E.E.C.), Sloan Research Fellow Award (E.E.C.) and Indiana University–Bloomington Department of Chemistry Start-Up Funds and a Marvin Carmack Fellowship (O.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kocaoglu, O., Carlson, E. Progress and prospects for small-molecule probes of bacterial imaging. Nat Chem Biol 12, 472–478 (2016). https://doi.org/10.1038/nchembio.2109
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2109
This article is cited by
-
Computerized fluorescence microscopy of microbial cells
World Journal of Microbiology and Biotechnology (2021)
-
The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications
Nature Communications (2019)
-
Towards the generalized iterative synthesis of small molecules
Nature Reviews Chemistry (2018)
-
EzColocalization: An ImageJ plugin for visualizing and measuring colocalization in cells and organisms
Scientific Reports (2018)
-
Illuminating vital surface molecules of symbionts in health and disease
Nature Microbiology (2017)