Abstract
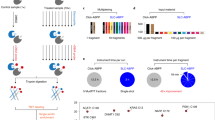
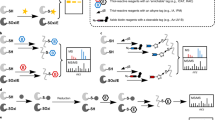
Activity-based protein profiling (ABPP) serves as a chemical proteomic platform to discover and characterize functional amino acids in proteins on the basis of their enhanced reactivity towards small-molecule probes. This approach, to date, has mainly targeted nucleophilic functional groups, such as the side chains of serine and cysteine, using electrophilic probes. Here we show that ‘reverse-polarity’ (RP)-ABPP using clickable, nucleophilic hydrazine probes can capture and identify protein-bound electrophiles in cells. Using this approach, we demonstrate that the pyruvoyl cofactor of S-adenosyl-L-methionine decarboxylase (AMD1) is dynamically controlled by intracellular methionine concentrations. We also identify a heretofore unknown modification—an N-terminally bound glyoxylyl group—in the poorly characterized protein secernin-3. RP-ABPP thus provides a versatile method to monitor the metabolic regulation of electrophilic cofactors and discover novel types of electrophilic modifications on proteins in human cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Walsh, C. T., Garneau-Tsodikova, S. & Gatto, G. J. Jr Protein posttranslational modifications: the chemistry of proteome diversifications. Angew. Chem. Int. Ed. 44, 7342–7372 (2005).
Okeley, N. M. & van der Donk, W. A. Novel cofactors via post-translational modifications of enzyme active sites. Chem. Biol. 7, R159–R171 (2000).
Olsen, J. V. & Mann, M. Status of large-scale analysis of post-translational modifications by mass spectrometry. Mol. Cell. Proteomics 12, 3444–3452 (2013).
Devarie Baez, N. O., Reisz, J. A. & Furdui, C. M. Mass spectrometry in studies of protein thiol chemistry and signaling: opportunities and caveats. Free Radical Biol. Med. 80, 191–211 (2015).
Furdui, C. M. & Poole, L. B. Chemical approaches to detect and analyze protein sulfenic acids. Mass Spectrom. Rev. 33, 126–146 (2014).
Chuh, K. N. & Pratt, M. R. Chemical methods for the proteome-wide identification of posttranslationally modified proteins. Curr. Opin. Chem. Biol. 24, 27–37 (2015).
Tate, E. W., Kalesh, K. A., Lanyon-Hogg, T., Storck, E. M. & Thinon, E. Global profiling of protein lipidation using chemical proteomic technologies. Curr. Opin. Chem. Biol. 24, 48–57 (2015).
Li, X. & Li, X. D. Chemical proteomics approaches to examine novel histone posttranslational modifications. Curr. Opin. Chem. Biol. 24, 80–90 (2015).
Cravatt, B. F., Wright, A. T. & Kozarich, J. W. Activity-based protein profiling: from enzyme chemistry to proteomic chemistry. Annu. Rev. Biochem. 77, 383–414 (2008).
Shannon, D. A. & Weerapana, E. Covalent protein modification: the current landscape of residue-specific electrophiles. Curr. Opin. Chem. Biol. 24, 18–26 (2015).
Klinman, J. P. & Bonnot, F. Intrigues and intricacies of the biosynthetic pathways for the enzymatic quinocofactors: PQQ, TTQ, CTQ, TPQ, and LTQ. Chem. Rev. 114, 4343–4365 (2014).
Phillips, R. S. Chemistry and diversity of pyridoxal-5′-phosphate dependent enzymes. Biochim. Biophys. Acta 1854, 1167–1174 (2015).
Zhang, H., Li, X. J., Martin, D. B. & Aebersold, R. Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat. Biotechnol. 21, 660–666 (2003).
Morgan, R. K. & Cohen, M. S. A clickable aminooxy probe for monitoring cellular ADP-ribosylation. ACS Chem. Biol. 10, 1778–1784 (2015).
Gupta, V. & Carroll, K. S. Sulfenic acid chemistry, detection and cellular lifetime. Biochim. Biophys. Acta 1840, 847–875 (2014).
Madian, A. G. & Regnier, F. E. Proteomic identification of carbonylated proteins and their oxidation sites. J. Proteome Res. 9, 3766–3780 (2010).
Alfaro, J. F. et al. Chemo-enzymatic detection of protein isoaspartate using protein isoaspartate methyltransferase and hydrazine trapping. Anal. Chem. 80, 3882–3889 (2008).
Klaene, J. J., Ni, W., Alfaro, J. F. & Zhou, Z. S. Detection and quantitation of succinimide in intact protein via hydrazine trapping and chemical derivatization. J. Pharm. Sci. 103, 3033–3042 (2014).
Rostovtsev, V. V., Green, L. G., Fokin, V. V. & Sharpless, K. B. A stepwise huisgen cycloaddition process: copper(I)-catalyzed regioselective “ligation” of azides and terminal alkynes. Angew. Chem. Int. Ed. 41, 2596–2599 (2002).
Speers, A. E., Adam, G. C. & Cravatt, B. F. Activity-based protein profiling in vivo using a copper(I)-catalyzed azide-alkyne [3+2] cycloaddition. J. Am. Chem. Soc. 125, 4686–4687 (2003).
Binda, C. et al. Structural and mechanistic studies of arylalkylhydrazine inhibition of human monoamine oxidases A and B. Biochemistry 47, 5616–5625 (2008).
Shantz, L. M., Stanley, B. A., Secrist, J. A. III & Pegg, A. E. Purification of human S-adenosylmethionine decarboxylase expressed in Escherichia coli and use of this protein to investigate the mechanism of inhibition by the irreversible inhibitors, 5′-deoxy-5′-[(3-hydrazinopropyl)methylamino]adenosine and 5′-([(Z)-4-amino-2-butenyl]methylamino)-5′-deoxyadenosine. Biochemistry 31, 6848–6855 (1992).
Washburn, M. P., Wolters, D. & Yates, J. R. III Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
Mann, M. Functional and quantitative proteomics using SILAC. Nat. Rev. Mol. Cell Biol. 7, 952–958 (2006).
Adibekian, A. et al. Click-generated triazole ureas as ultrapotent in vivo-active serine hydrolase inhibitors. Nat. Chem. Biol. 7, 469–478 (2011).
Dalle-Donne, I. et al. Protein carbonylation: 2,4-dinitrophenylhydrazine reacts with both aldehydes/ketones and sulfenic acids. Free Radical Biol. Med. 46, 1411–1419 (2009).
Gupta, V., Paritala, H. & Carroll, K. S. Reactivity, selectivity and stability in sulfenic acid detection: a comparative study of nucleophilic and electropilic probes. Bioconjugate Chem. 27, 1411–1418 (2016).
Tolbert, W. D. et al. The structural basis for substrate specificity and inhibition of human S-adenosylmethionine decarboxylase. Biochemistry 40, 9484–9494 (2001).
McCloskey, D. E. et al. New insights into the design of inhibitors of human S-adenosylmethionine decarboxylase: studies of adenine C8 substitution in structural analogues of S-adenosylmethionine. J. Med. Chem. 52, 1388–1407 (2009).
Shantz, L. M., Holm, I., Jänne, O. A. & Pegg, A. E. Regulation of S-adenosylmethionine decarboxylase activity by alterations in the intracellular polyamine content. Biochem. J. 288, 511–518 (1992).
Zinellu, A. et al. Plasma methionine determination by capillary electrophoresis-UV assay: application on patients affected by retinal venous occlusive disease. Anal. Biochem. 363, 91–96 (2007).
Xiong, H., Stanley, B. A. & Pegg, A. E. Role of cysteine-82 in the catalytic mechanism of human S-adenosylmethionine decarboxylase. Biochemistry 38, 2462–2470 (1999).
Weerapana, E. et al. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 468, 790–795 (2010).
Weerapana, E., Speers, A. E. & Cravatt, B. F. Tandem orthogonal proteolysis-activity-based protein profiling (TOP-ABPP)–a general method for mapping sites of probe modification in proteomes. Nat. Protoc. 2, 1414–1425 (2007).
Pei, J. & Grishin, N. V. Peptidase family U34 belongs to the superfamily of N-terminal nucleophile hydrolases. Protein. Sci. 12, 1131–1135 (2003).
Brannigan, J. A. et al. A protein catalytic framework with an N-terminal nucleophile is capable of self-activation. Nature 378, 416–419 (1995).
Aronson, N. N. Jr Lysosomal glycosylasparaginase: a member of a family of amidases that employ a processed N-terminal threonine, serine or cysteine as a combined base-nucleophile catalyst. Glycobiology 6, 669–675 (1996).
Dirksen, A., Dirksen, S., Hackeng, T. M. & Dawson, P. E. Nucleophilic catalysis of hydrazone formation and transimination: implications for dynamic covalent chemistry. J. Am. Chem. Soc. 128, 15602–15603 (2006).
Dirksen, A. & Dawson, P. E. Rapid oxime and hydrazone ligations with aromatic aldehydes for biomolecular labeling. Bioconjug. Chem. 19, 2543–2548 (2008).
Yerlikaya, A. & Stanley, B. A. S-adenosylmethionine decarboxylase degradation by the 26S proteasome is accelerated by substrate-mediated transamination. J. Biol. Chem. 279, 12469–12478 (2004).
Dall, E., Fegg, J. C., Briza, P. & Brandstetter, H. Structure and mechanism of an aspartimide-dependent peptide ligase in human legumain. Angew. Chem. Int. Ed. 54, 2917–2921 (2015).
Casero, R. A. Jr & Marton, L. J. Targeting polyamine metabolism and function in cancer and other hyperproliferative diseases. Nat. Rev. Drug Discov. 6, 373–390 (2007).
Jaramillo, M. C. & Zhang, D. D. The emerging role of the Nrf2-Keap1 signaling pathway in cancer. Genes Dev. 27, 2179–2191 (2013).
Blennow, K., Mattsson, N., Schöll, M., Hansson, O. & Zetterberg, H. Amyloid biomarkers in Alzheimer's disease. Trends Pharmacol. Sci. 36, 297–309 (2015).
Yang, J. et al. FTO genotype is associated with phenotypic variability of body mass index. Nature 490, 267–272 (2012).
Buckingham, J. The chemistry of arylhydrazones. Q. Rev. Chem. Soc. 23, 37–56 (1969).
Huizinga, E. G. et al. Active site structure of methylamine dehydrogenase: hydrazines identify C6 as the reactive site of the tryptophan-derived quinone cofactor. Biochemistry 31, 9789–9795 (1992).
Appel, M. J. & Bertozzi, C. R. Formylglycine, a post-translationally generated residue with unique catalytic capabilities and biotechnology applications. ACS Chem. Biol. 10, 72–84 (2015).
Xu, Q., Buckley, D., Guan, C. & Guo, H. C. Structural insights into the mechanism of intramolecular proteolysis. Cell 98, 651–661 (1999).
Kabisch, U. C. et al. Identification of D-proline reductase from Clostridium sticklandii as a selenoenzyme and indications for a catalytically active pyruvoyl group derived from a cysteine residue by cleavage of a proprotein. J. Biol. Chem. 274, 8445–8454 (1999).
Andreesen, J. R. Glycine reductase mechanism. Curr. Opin. Chem. Biol. 8, 454–461 (2004).
Mihara, H. & Esaki, N. Bacterial cysteine desulfurases: their function and mechanisms. Appl. Microbiol. Biotechnol. 60, 12–23 (2002).
Scheck, R. A., Dedeo, M. T., Iavarone, A. T. & Francis, M. B. Optimization of a biomimetic transamination reaction. J. Am. Chem. Soc. 130, 11762–11770 (2008).
Acknowledgements
We acknowledge Steven Ealick and Leslie Kinsland for the AdoMetDC bacterial expression plasmid, Keriann Backus for the cleavable tags, Gonzalo González-Páez and Dennis Wolan for TEV protease, Melissa Dix and Jim Moresco for instrument assistance, Armand Cognetta and Kenneth Lum for software and data assistance, Liron Bar-Peled and Stephan Hacker for plasmids and Silvia Ortega-Gutiérrez for helpful discussions. This work was supported by the National Institutes of Health Grants CA132630 (B.F.C.), P41 GM103533 and U54 GM114833-02 (J.R.Y.), an NSF Graduate Research Fellowship DGE-1346837 (E.J.O.) and a Helen Hay Whitney Postdoctoral Fellowship sponsored by Merck (M.L.M).
Author information
Authors and Affiliations
Contributions
M.L.M and B.F.C. conceived the project, designed experiments and composed the manuscript. M.L.M performed all experiments. L.H. wrote software and J.R.Y. provided advice. M.L.M., L.H., B.E.C. and B.F.C. analysed data. M.L.M., E.J.O. and P.E.D. designed peptide standards. M.L.M. and E.J.O. synthesized peptides. M.L.M. and B.D.H. synthesized probes.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 6324 kb)
Supplementary information
Supplementary information (XLSX 9547 kb)
Rights and permissions
About this article
Cite this article
Matthews, M., He, L., Horning, B. et al. Chemoproteomic profiling and discovery of protein electrophiles in human cells. Nature Chem 9, 234–243 (2017). https://doi.org/10.1038/nchem.2645
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2645
This article is cited by
-
Advances in covalent drug discovery
Nature Reviews Drug Discovery (2022)
-
Enterococci enhance Clostridioides difficile pathogenesis
Nature (2022)
-
Reimagining high-throughput profiling of reactive cysteines for cell-based screening of large electrophile libraries
Nature Biotechnology (2021)
-
Global targeting of functional tyrosines using sulfur-triazole exchange chemistry
Nature Chemical Biology (2020)
-
Nature Biotechnology's academic spinouts of 2017
Nature Biotechnology (2018)