Abstract
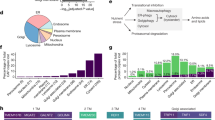
Entry into the cell cycle is regulated by nutrient availability such that cells do not divide when resources are limited. The Skp1–Cul1–F-box (SCF) ubiquitin ligase with the F-box protein Grr1 (SCFGrr1) controls the proteolytic turnover of regulators of cell-cycle entry and a glucose sensor, suggesting that it links the cell cycle with nutrient availability. Here, we show that SCFGrr1 broadly regulates cellular metabolism. We have developed a proteomic screening method that uses high-throughput quantitative microscopy to comprehensively screen for ubiquitin-ligase substrates. Seven new metabolic targets of SCFGrr1 were identified, including two regulators of glycolysis — the transcription factor Tye7 and Pfk27. The latter produces the second messenger fructose-2,6-bisphosphate that activates glycolysis and inhibits gluconeogenesis. We show that SCFGrr1 targets Pfk27 and Tye7 in response to glucose removal. Moreover, Pfk27 is phosphorylated by the kinase Snf1, and unphosphorylatable Pfk27 is stable and inhibits growth in the absence of glucose. These results demonstrate a role for SCFGrr1 in regulating the glycolytic–gluconeogenic switch.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Willems, A. R., Schwab, M. & Tyers, M. A hitchhiker's guide to the cullin ubiquitin ligases: SCF and its kin. Biochim. Biophys. Acta 1695, 133–170 (2004).
Petroski, M. D. & Deshaies, R. J. Function and regulation of cullin–RING ubiquitin ligases. Nature Rev. Mol. Cell Biol. 6, 9–20 (2005).
Cardozo, T. & Pagano, M. The SCF ubiquitin ligase: insights into a molecular machine. Nature Rev. Mol. Cell Biol 5, 739–751 (2004).
Flick, J. S. & Johnston, M. GRR1 of Saccharomyces cerevisiae is required for glucose repression and encodes a protein with leucine-rich repeats. Mol. Cell. Biol. 11, 5101–5112 (1991).
Flick, K. M. et al. Grr1-dependent inactivation of Mth1 mediates glucose-induced dissociation of Rgt1 from HXT gene promoters. Mol. Biol. Cell 14, 3230–3241 (2003).
Spielewoy, N., Flick, K., Kalashnikova, T. I., Walker, J. R. & Wittenberg, C. Regulation and recognition of SCFGrr1 targets in the glucose and amino acid signaling pathways. Mol. Cell. Biol. 24, 8994–9005 (2004).
Andreasson, C. & Ljungdahl, P. O. The N-terminal regulatory domain of Stp1p is modular and, fused to an artificial transcription factor, confers full Ssy1p-Ptr3p-Ssy5p sensor control. Mol. Cell. Biol. 24, 7503–7513 (2004).
Barral, Y., Jentsch, S. & Mann, C. G1 cyclin turnover and nutrient uptake are controlled by a common pathway in yeast. Genes Dev. 9, 399–409 (1995).
Jaquenoud, M., Gulli, M. P., Peter, K. & Peter, M. The Cdc42p effector Gic2p is targeted for ubiquitin-dependent degradation by the SCFGrr1 complex. EMBO J. 17, 5360–5373 (1998).
Blondel, M. et al. Degradation of Hof1 by SCF(Grr1) is important for actomyosin contraction during cytokinesis in yeast. EMBO J. 24, 1440–1452 (2005).
Purnapatre, K., Gray, M., Piccirillo, S. & Honigberg, S. M. Glucose inhibits meiotic DNA replication through SCFGrr1p-dependent destruction of Ime2p kinase. Mol. Cell. Biol. 25, 440–450 (2005).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Eckert-Boulet, N., Regenberg, B. & Nielsen, J. Grr1p is required for transcriptional induction of amino acid permease genes and proper transcriptional regulation of genes in carbon metabolism of Saccharomyces cerevisiae. Curr. Genet. 47, 139–149 (2005).
Gardner, R. G., Nelson, Z. W. & Gottschling, D. E. Degradation-mediated protein quality control in the nucleus. Cell 120, 803–815 (2005).
Kishi, T. & Yamao, F. An essential function of Grr1 for the degradation of Cln2 is to act as a binding core that links Cln2 to Skp1. J. Cell Sci. 111, 3655–3661 (1998).
Hsiung, Y. G. et al. F-box protein Grr1 interacts with phosphorylated targets via the cationic surface of its leucine-rich repeat. Mol. Cell. Biol. 21, 2506–2520 (2001).
Andreasson, C. & Ljungdahl, P. O. Receptor-mediated endoproteolytic activation of two transcription factors in yeast. Genes Dev. 16, 3158–3172 (2002).
Okar, D. A. & Lange, A. J. Fructose-2,6-bisphosphate and control of carbohydrate metabolism in eukaryotes. Biofactors 10, 1–14 (1999).
Nishi, K. et al. The GCR1 requirement for yeast glycolytic gene expression is suppressed by dominant mutations in the SGC1 gene, which encodes a novel basic-helix-loop-helix protein. Mol. Cell. Biol. 15, 2646–2653 (1995).
Moriya, H. & Johnston, M. Glucose sensing and signaling in Saccharomyces cerevisiae through the Rgt2 glucose sensor and casein kinase I. Proc. Natl Acad. Sci. USA 101, 1572–1577 (2004).
Ptacek, J. et al. Global analysis of protein phosphorylation in yeast. Nature 438, 679–684 (2005).
Dale, S., Wilson, W. A., Edelman, A. M. & Hardie, D. G. Similar substrate recognition motifs for mammalian AMP-activated protein kinase, higher plant HMG-CoA reductase kinase-A, yeast SNF1, and mammalian calmodulin-dependent protein kinase I. FEBS Lett. 361, 191–195 (1995).
Goncalves, P. M., Griffioen, G., Bebelman, J. P. & Planta, R. J. Signalling pathways leading to transcriptional regulation of genes involved in the activation of glycolysis in yeast. Mol. Microbiol. 25, 483–493 (1997).
Tang, X. et al. Genome-wide surveys for phosphorylation-dependent substrates of SCF ubiquitin ligases. Methods Enzymol. 399, 433–458 (2005).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Schmitt, M. E., Brown, T. A. & Trumpower, B. L. A rapid and simple method for preparation of RNA from Saccharomyces cerevisiae. Nucleic Acids Res. 18, 3091–3092 (1990).
Espinoza, F. H. et al. Cak1 is required for Kin28 phosphorylation and activation in vivo. Mol. Cell. Biol. 18, 6365–6373 (1998).
Acknowledgements
The authors thank J. Weissman, E. O'Shea, K. Weis, M. Tyers, A. Amon, C. Boone and T. Kishi for reagents; V. Vincent (Cellomics, Pittsburg, PA) and the University of California, San Francisco Laboratory for Cell Analysis for assistance with microscopy; members of the Morgan laboratory for reagents and discussion; and D. Morgan, K. Ashrafi, T. Fazzio and members of the Toczyski laboratory for discussion and comments on the manuscript. J.A.B is supported by the Damon Runyon Cancer Research Foundation. This work was supported by a National Institutes of Health grant to D.P.T.
Author information
Authors and Affiliations
Contributions
J.A.B. and D.P.T. designed the experiments, analysed the data and wrote the paper. J.A.B. carried out the experiments with assistance from S.K.C. and M.C.B.
Corresponding author
Supplementary information
Supplementary Information.
Supplementary Figures S1, S2, S3, S4 and S5, Supplementary Tables S1 and S2, and Methods (PDF 1048 kb)
Rights and permissions
About this article
Cite this article
Benanti, J., Cheung, S., Brady, M. et al. A proteomic screen reveals SCFGrr1 targets that regulate the glycolytic–gluconeogenic switch. Nat Cell Biol 9, 1184–1191 (2007). https://doi.org/10.1038/ncb1639
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1639
This article is cited by
-
Skp2 dictates cell cycle-dependent metabolic oscillation between glycolysis and TCA cycle
Cell Research (2021)
-
Redundant targeting of Isr1 by two CDKs in mitotic cells
Current Genetics (2021)
-
Phylogenetic and Expression Analyses of Cullin Family Members Unveil the Role of PbCUL1.C1 in Pollen Tube Growth Underlying Non-self S-RNase in Pear
Plant Molecular Biology Reporter (2020)
-
Enzyme–substrate relationships in the ubiquitin system: approaches for identifying substrates of ubiquitin ligases
Cellular and Molecular Life Sciences (2017)
-
Isolation of ubiquitinated substrates by tandem affinity purification of E3 ligase–polyubiquitin-binding domain fusions (ligase traps)
Nature Protocols (2016)