Abstract
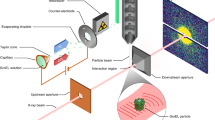
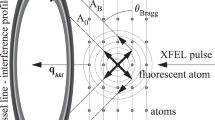
X-ray lasers offer new capabilities in understanding the structure of biological systems, complex materials and matter under extreme conditions1,2,3,4. Very short and extremely bright, coherent X-ray pulses can be used to outrun key damage processes and obtain a single diffraction pattern from a large macromolecule, a virus or a cell before the sample explodes and turns into plasma1. The continuous diffraction pattern of non-crystalline objects permits oversampling and direct phase retrieval2. Here we show that high-quality diffraction data can be obtained with a single X-ray pulse from a non-crystalline biological sample, a single mimivirus particle, which was injected into the pulsed beam of a hard-X-ray free-electron laser, the Linac Coherent Light Source5. Calculations indicate that the energy deposited into the virus by the pulse heated the particle to over 100,000 K after the pulse had left the sample. The reconstructed exit wavefront (image) yielded 32-nm full-period resolution in a single exposure and showed no measurable damage. The reconstruction indicates inhomogeneous arrangement of dense material inside the virion. We expect that significantly higher resolutions will be achieved in such experiments with shorter and brighter photon pulses focused to a smaller area. The resolution in such experiments can be further extended for samples available in multiple identical copies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Neutze, R. et al. Potential for biomolecular imaging with femtosecond X-ray pulses. Nature 406, 752–757 (2000)
Chapman, H. N. et al. Femtosecond diffractive imaging with a soft-X-ray free-electron laser. Nature Phys. 2, 839–843 (2006)
Bergh, M. et al. Feasibility of imaging living cells at sub-nanometer resolution by ultrafast X-ray diffraction. Q. Rev. Biophys. 41, 181–204 (2008)
Nagler, B. et al. Turning solid aluminium transparent by intense soft X-ray photoionization. Nature Phys. 5, 693–696 (2009)
Emma, P. et al. First lasing and operation of an ångstrom-wavelength free-electron laser. Nature Photon. 4, 641–647 (2010)
La Scola, B. et al. A giant virus in amoebae. Science 299, 2033 (2003)
Xiao, C. et al. Structural studies of the giant mimivirus. PLoS Biol. 7, e1000092 (2009)
Claverie, J. M. & Abergel, C. Mimivirus and its virophage. Annu. Rev. Genet. 43, 49–66 (2009)
Raoult, D. et al. The 1.2-megabase genome sequence of mimivirus. Science 306, 1344–1350 (2004)
La Scola, B. et al. The virophage as a unique parasite of the giant mimivirus. Nature 455, 100–104 (2008)
Claverie, J. M. & Abergel, C. Mimivirus: the emerging paradox of quasi-autonomous viruses. Trends Genet. 26, 431–437 (2010)
Strüder, L. et al. Large-format, high-speed, X-ray pnCCDs combined with electron and ion imaging spectrometers in a multipurpose chamber for experiments at 4th generation light sources. Nucl. Instrum. Methods Phys. Res. A 614, 483–496 (2010)
Bozek, J. D. AMO instrumentation for the LCLS X-ray FEL. Eur. Phys. J. Spec. Top. 169, 129–132 (2009)
DePonte, D. P. et al. Gas dynamic virtual nozzle for generation of microscopic droplet streams. J. Phys. D 41, 195505 (2008)
Bogan, M. J. et al. Single particle X-ray diffractive imaging. Nano Lett. 8, 310–316 (2008)
Maia, F. R. N. C., Ekeberg, T., van der Spoel, D. & Hajdu, J. Hawk: the image reconstruction package for coherent X-ray diffractive imaging. J. Appl. Crystallogr. 43, 1535–1539 (2010)
Luke, D. R. Relaxed averaged alternating reflections for diffraction imaging. Inverse Probl. 21, 37–50 (2005)
Marchesini, S. et al. X-ray image reconstruction from a diffraction pattern alone. Phys. Rev. B 68, 140101 (2003)
Thibault, P., Elser, V., Jacobsen, C., Shapiro, D. & Sayre, D. Reconstruction of a yeast cell from X-ray diffraction data. Acta Crystallogr. A 62, 248–261 (2006)
Shapiro, D. et al. Biological imaging by soft X-ray diffraction microscopy. Proc. Natl Acad. Sci. USA 102, 15343–15346 (2005)
Ding, Y. et al. Measurements and simulations of ultralow emittance and ultrashort electron beams in the Linac Coherent Light Source. Phys. Rev. Lett. 102, 254801 (2009)
Young, L. et al. Femtosecond electronic response of atoms to ultra-intense X-rays. Nature 466, 56–61 (2010)
Hau-Riege, S. P. et al. A sacrificial tamper slows down sample explosion in flash diffraction experiments. Phys. Rev. Lett. 104, 064801 (2010)
Chapman, H. N. et al. Femtosecond X-ray protein nanocrystallography. Nature doi:10.1038/nature09750 (this issue).
Huldt, G., Szoke, A. & Hajdu, J. Diffraction imaging of single particles and biomolecules. J. Struct. Biol. 144, 219–227 (2003)
Fung, R. et al. Structure from fleeting illumination of faint spinning objects in flight. Nature Phys. 5, 64–67 (2009)
Loh, N. T. D. & Elser, V. Reconstruction algorithm for single-particle diffraction imaging. Phys. Rev. E 80, 026705 (2009)
Bortel, G., Faigel, G. & Tegze, M. Classification and averaging of random orientation single macromolecular diffraction patterns at atomic resolution. J. Struct. Biol. 166, 226–233 (2009)
Maia, F. R. N. C., Ekeberg, T., Tımneanu, N., van der Spoel, D. & Hajdu, J. Structural variability and the incoherent addition of scattered intensities in single-particle diffraction. Phys. Rev. E 80, 031905 (2009)
Perkins, T. T., Smith, D. E. & Chu, S. Single polymer dynamics in an elongated flow. Science 276, 2016–2021 (1997)
Song, C. et al. Quantitative imaging of single, unstained viruses with coherent X rays. Phys. Rev. Lett. 101, 158101 (2008)
Byrne, D. et al. The polyadenylation site of Mimivirus transcripts obeys a stringent ‘hairpin rule’. Genome Res. 19, 1233–1242 (2009)
Steinbrener, J. et al. Data preparation and evaluation techniques for X-ray diffraction microscopy. Opt. Express 18, 18598–18614 (2010)
Seibert, M. M. et al. Femtosecond diffractive imaging of biological cells. J. Phys. At. Mol. Opt. Phys. 43, 194015 (2010)
Acknowledgements
This work was supported by the following agencies: the Swedish Research Councils; the Swedish Foundation for International Cooperation in Research and Higher Education; Stiftelsen Olle Engkvist Byggmästare; the Swedish University of Agricultural Sciences; the Helmholtz Association (VH-VI-302); the DFG Cluster of Excellence at the Munich Centre for Advanced Photonics; the Centre National de la Recherche Scientifique; Agence Nationale de la Recherche (ANR-BLAN08-0089); the Hamburg Ministry of Science and Research and Joachim Herz Stiftung, as part of the Hamburg Initiative for Excellence in Research (LEXI); the Hamburg School for Structure and Dynamics; the Max Planck Society, the US National Science Foundation (grants MCB 0919195 and MCB-1021557); and the US Department of Energy, through the PULSE Institute. Portions of this research were carried out at the Linac Coherent Light Source, a National User Facility operated by Stanford University on behalf of the US Department of Energy, Office of Basic Energy Sciences. We are grateful to B. Hedman and N. Timneanu for their help and to the scientific and technical staff of the LCLS for their outstanding facility and support.
Author information
Authors and Affiliations
Contributions
J.H. and H.N.C. conceived the experiment. C.A., V.S., J.-M.C., M.S., O.J., A. Rocker, I.A. and D.O. prepared and characterized the samples. J.D.B., C.B. and K.O.H. created the LCLS beamline. J.H., H.N.C., J.S., L.G., A.B., N.C., A.A., A.M., J.K., K.O.H. and S.P.H.-R. developed the imaging concept. S.W.E., R.H., D. Rolles, A. Rudenko, C.S., L.F., N.K., P.H., B.R., B.E., A.H., Ch.R., D.P., G.W., L.S., G.H., H. Gorke, J.U., I.S., S.H., G.S., F.S., H.S., K.-U.K., R.A., C.-D.S., F.K., M. Bott, S. Schorb, D. Rupp, M.A., T.G., H.H., L.G., G.P., H. Graafsma and B.N. designed and set up the CAMP instrument and/or developed and operated the pnCCD detectors. S. Bajt and M. Barthelmess coordinated instrumentation, worked on engineering and prepared filters and calibration samples. J.K., S.P.H.-R., A.B., H.N.C., J.S., A.M. and N.C. characterized the focus. B.I., D.W., M.S., M.M.S. and J.H. built the sample injector. D.P.D., J.H., M.S., D.W., U.W., R.K., M. Hunter, R.B.D. and J.C.H.S. designed and built the nanospray nebulizer. J.H., M.S., M.M.S., F.R.N.C.M., J.A., A.A.M., A. Rocker, M.J.B., C.Y.H., R.G.S., S. Boutet, I.A., O.J., D.S., A.B., J.S., D.P.D., A.A., M.L., J.K., T.A.W., A.M., R.L.S., L.L., M. Barthelmess, J.C.H.S., P.F., I.S., U.W., R.K., M. Hunter, R.B.D., M.F., G.W., M. Bott, S.W.E., B.E., L.F., R.H., N.K., L.L., D. Rupp, B.R., A. Rudenko, R.L.S., L.S., I.S., C.S., J.U. and H.N.C. characterized the imaging apparatus and carried out the experiment. T.E., F.R.N.C.M., M. Hantke, M.M.S., A.B., T.A.W., A.M., A.A., S.M., M.M. and J.H. analysed the data. T.E., F.R.N.C.M., A.B. and S.M. performed image reconstructions. J.H. and T.E. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
PowerPoint slides
Rights and permissions
About this article
Cite this article
Seibert, M., Ekeberg, T., Maia, F. et al. Single mimivirus particles intercepted and imaged with an X-ray laser. Nature 470, 78–81 (2011). https://doi.org/10.1038/nature09748
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09748
This article is cited by
-
In situ label-free X-ray imaging for visualizing the localization of nanomedicines and subcellular architecture in intact single cells
Nature Protocols (2024)
-
Chaotic dynamics in X-ray free-electron lasers with an optical undulator
Scientific Reports (2024)
-
Water layer and radiation damage effects on the orientation recovery of proteins in single-particle imaging at an X-ray free-electron laser
Scientific Reports (2023)
-
Finding the semantic similarity in single-particle diffraction images using self-supervised contrastive projection learning
npj Computational Materials (2023)
-
Density functional tight binding approach utilized to study X-ray-induced transitions in solid materials
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.