Abstract
As one member of fatty acid binding proteins (FABPs), FABP5 makes a contribution in the occurrence and development of several tumor types, but existing analysis about FABP5 and FABP5-related molecular mechanism remains limited. Meanwhile, some tumor patients showed limited response rates to current immunotherapy, and more potential targets need to be explored for the improvement of immunotherapy. In this study, we made a pan-cancer analysis of FABP5 based on the clinical data from The Cancer Genome Atlas database for the first time. FABP5 overexpression was observed in many tumor types, and was statistically associated with poor prognosis of several tumor types. Additionally, we further explored FABP5-related miRNAs and corresponding lncRNAs. Then, miR-577-FABP5 regulatory network in kidney renal clear cell carcinoma as well as CD27-AS1/GUSBP11/SNHG16/TTC28-AS1-miR-22-3p-FABP5 competing endogenous RNA regulatory network in liver hepatocellular carcinoma were constructed. Meanwhile, Western Blot and reverse transcription quantitative real-time polymerase chain reaction (RT-qPCR) analysis were used to verify miR-22-3p-FABP5 relationship in LIHC cell lines. Moreover, the potential relationships of FABP5 with immune infiltration and six immune checkpoints (CD274, CTLA4, HAVCR2, LAG3, PDCD1 and TIGIT) were discovered. Our work not only deepens the understanding of FABP5’s functions in multiple tumors and supplements existing FABP5-related mechanisms, but also provides more possibilities for immunotherapy.
Similar content being viewed by others
Introduction
Fatty acid binding proteins (FABPs), serving as noncatalytic binding proteins with complex mechanism and functional diversity, have been considered to play an important role in the modulation of lipid fluxes, signal transduction as well as metabolism1,2. As one member of this family, FABP5 presents unique characteristics in various specific cells and tissues including adipocytes, macrophages and tumor cells2. Jing et al. first identified FABP5 as a tumor-promoting gene3,4, and other researchers have also confirmed its role in several tumor types5,6,7,8. Studies have reported that FABP5 promotes cell proliferation, progression and invasion of tumors via fatty acid uptake oxidation and several signaling such as PPAR β/δ and HIF-1α2,9,10. Additionally, the prognostic value of FABP5 in several tumor types has also been revealed by several studies. For example, FABP5 is associated with poor prognosis in clear cell renal cell carcinoma, gastric cancer and triple-negative breast cancer11,12,13.
In addition to FABP5, tumor microenvironment (TME), which is composed of stromal cells, immune cells and tumor cells, also makes great contributions in the tumorigenesis and progression14,15. Tumor cells and other nonmalignant cells interact with each other, building a microenvironment which is conducive to tumor growth16. As a part of stromal cells, cancer associated fibroblasts (CAFs) promote tumor growth and make tumor cells more carcinogenic17. Among those immune cells in TME, effector T cells, dendritic cells (DCs), M1 macrophages and natural killer cells serve as antineoplastic factors, while immunosuppressive cells, including regulatory T cells (Tregs) and M2 macrophages, are tumor-promoting factors15,18. Tumor immune escape is one of vital mechanisms in tumor occurrence and progression. One of main factors in this mechanism is that the above-mentioned immunosuppressive cells influence the immune tolerance of tumor cells19. Another factor is that the regulation of immune checkpoints’ expression inhibits effector T cells’ activation19. Targeting immune checkpoints has been a hot direction in the current clinical research and considered as one of effective antitumor therapies. Up to date, three immune checkpoints, namely programmed cell death 1 (PD-1/PDCD1), programmed cell death ligand 1 (PD-L1/CD274) and cytotoxic T lymphocyte-associated antigen 4 (CTLA4) have been frequently targeted. Nevertheless, patients under the therapy targeting above immune checkpoints have limited response rates. Therefore, the exploration of potential targets is needed to obtain higher response rates and improve the antitumor therapeutic effectiveness. There have been several novel immune checkpoints, including T cell immunoreceptor with Ig and ITIM domains (TIGIT), T cell immunoglobulin domain and mucin domain 3 (TIM-3/HAVCR2) and lymphocyte activation gene 3 (LAG3) under clinical trials. LAG3 and HAVCR2 have been reported to associate with T cell exhaustion, and TIGIT plays a vital role in the limitation of T cell inflammation20,21,22.
A report has defined a new regulatory network, namely competing endogenous RNA (ceRNA), constituted by the competition between mRNAs, long non-coding RNAs (lncRNAs), pseudogene transcripts and circRNAs for the binding of microRNA (miRNA)23. This novel regulatory network has been confirmed to have a great role in tumor occurrence and development24,25,26. Among different kinds of ceRNA networks currently discovered, a ceRNA network targeting FABP5, namely circ-ABCB10/miR-620/FABP5, have been proved to participate in the occurrence and development of glioma27. Although the potential oncogenic role of FABP5 has been confirmed in multiple tumor types, there is a lack of more comprehensive analyses about FABP5 and corresponding molecular mechanism. For further understanding of FABP5-related molecular mechanism in multiple tumor types, more ceRNA regulatory networks need to be explored.
Taken together, our study made a pan-cancer analysis of FABP5 on the basis of the clinical data from The Cancer Genome Atlas (TCGA) database for the first time, and observed novel FABP5-related regulatory networks in kidney renal clear cell carcinoma (KIRC) and liver hepatocellular carcinoma (LIHC). Meanwhile, miR-22-3p-FABP5 relationship was verified in LIHC cell lines for the first time. Besides, we evaluated the association between FABP5 and immune infiltration, and further analyzed the association between FABP5 and six immune checkpoints. Our work mainly aimed to supplement and improve existing FABP5-related molecular and immunological mechanisms, and provide more possibilities for immunotherapy.
Results
Expression analysis of FABP5
We conducted expression analysis of FABP5 in multiple tumor types with sufficient normal tissues (more than five cases) in TCGA project. Compared with normal tissues, we observed the up-regulated expression of FABP5 in the tumor tissues of cholangiocarcinoma (CHOL), esophageal carcinoma (ESCA), glioblastoma multiforme (GBM), head and neck squamous cell carcinoma (HNSC), KIRC, kidney renal papillary cell carcinoma (KIRP), LIHC, prostate adenocarcinoma (PRAD) and uterine corpus endometrial carcinoma (UCEC) (ESCA p < 0.05, GBM and HNSC p < 0.01, and other tumor types p < 0.001), but down-regulated expression in tissues of breast invasive carcinoma (BRCA), colon adenocarcinoma (COAD), kidney chromophobe (KICH), lung adenocarcinoma (LUAD) and thyroid carcinoma (THCA) (COAD and KICH p < 0.05, and other tumor types p < 0.001) (Fig. 1A).
Then we added normal tissues from GTEx database as controls, and further analyzed the difference of FABP5 expression between the normal and tumor tissues in other thirteen tumor types (Fig. 1B). Compared with normal tissues, we found that FABP5 expression was up-regulated in the tumor tissues of cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC), lymphoid neoplasm diffuse large B-cell lymphoma (DLBC), acute myeloid leukemia (LAML), ovarian serous cystadenocarcinoma (OV), pancreatic adenocarcinoma (PAAD), pheochromocytoma and paraganglioma (PCPG), testicular germ cell tumors (TGCT), Thymoma (THYM) and uterine carcinosarcoma (UCS), but down-regulated in brain lower grade glioma (LGG) and skin cutaneous melanoma (SKCM) (all p < 0.05).
Furthermore, we investigated the potential association between FABP5 expression and pathological stages of tumors. FABP5 expression was associated with pathological stages of Adrenocortical carcinoma (ACC), BLCA, CESC and KIRC (ACC p = 0.011, BLCA p = 0.001, CESC p = 0.034 and KIRC p = 0.009, Fig. 1C). As the tumor progresses, FABP5 expression increased in the cases of CESC and KIRC, but decreased in BLCA.
Survival analysis of FABP5
Survival analysis was conducted by GEPIA to investigate the prognostic value of FABP5. High expression of FABP5 was associated with poor overall survival (OS) in ACC, GBM, KIRC, LAML, LGG, LIHC, LUAD, SKCM and uveal melanoma (UVM) (ACC p = 0.025, GBM p = 0.014, KIRC p = 0.001, LAML p = 0.024, LIHC p = 0.001, LUAD p = 0.001, SKCM p = 0.017, and other tumor types p < 0.001, Fig. 2A). Additionally, the results of disease-free survival (DFS) analysis showed that high FABP5 expression was statistically associated with poor prognosis in ACC, BRCA, KIRC, LGG, LUAD and UVM, but favorable prognosis in lung squamous cell carcinoma (LUSC) (ACC p = 0.012, BRCA p = 0.022, KIRC p = 0.024, LGG p = 0.003, LUAD p = 0.038, LUSC p = 0.045, UVM p = 0.007, Fig. 2B).
Survival analysis was also conducted by the Kaplan–Meier survival curve and Log-rank test in those tumor types with FABP5 overexpression. We found that high FABP5 expression was associated with poor OS prognosis by the Kaplan–Meier survival curve and Log-rank test simultaneously in only three tumor types, namely GBM, KIRC and LIHC (Table 1).
Prediction and analysis of upstream miRNAs
Considering that there is no corresponding miRNA transcriptome data of GBM available in the TCGA database, we narrowed our study to two tumor types, namely KIRC and LIHC. Using ENCORI approach, we predicted potential upstream miRNAs, all of which have already been confirmed by CLIP-Seq experiment to have interactions with FABP5. Seventeen predicted miRNAs were shown in Table 2. In KIRC, there were only one miRNA, namely miR-577, significantly negatively correlated with FABP5 (R = − 0.22, p < 0.001, Fig. 3A). Down-regulated miR-577 expression was observed in KIRC tissues compared with normal tissues (p < 0.001, Fig. 3B). Besides, KIRC patients with low miR-577 expression had poor OS (p = 0.077, Fig. 3C). In LIHC, we also found only one miRNA, namely miR-22-3p, having significantly negative correlation with FABP5 (R = − 0.26, p < 0.001, Fig. 3D). MicroRNA-22-3p expression was down-regulated in tumor tissues compared with normal tissues (p < 0.001, Fig. 3E), and low miR-22-3p expression was associated with poor OS in LIHC (p < 0.001, Fig. 3F).
(A) Correlation analysis between FABP5 and miR-577 in KIRC. (B) Expression analysis of miR-577 in KIRC. (C) Overall survival analysis of miR-577 in KIRC. (D) Correlation analysis of FABP5 with miR-22-3p in LIHC. (E) Expression analysis of miR-22-3p in LIHC. (F) Overall survival analysis of miR-22-3p in LIHC.
Primary validation of the predicted association between miR-22-3p and FABP5 in LIHC
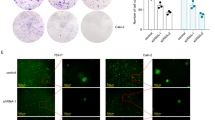
Binding sites of miR-22-3p on FABP5 predicted by PITA, miRmap and miRanda were shown in Fig. 4A. To test the accuracy of the miRNA-mRNA interactions predicted via ENCORI approach and ensure the innovation of the study, we chose LIHC cell lines for the validation of the potential miR-22-3p-FABP5 association that has not been reported in any tumor type yet. Our study transfected miR-22-3p mimics into two different LIHC cell lines (Hep3B and HepG2). The results of reverse transcription quantitative real-time polymerase chain reaction (RT-qPCR) presented higher miR-22-3p expression in miR-22-3p mimics group compared with miR negative control (NC) group in both Hep3B and HepG2 cell lines (Fig. 4B), reflecting the transfection effectiveness of miR-22-3p. Then we detected the FABP5 protein expression in each group, and found lower FABP5 protein expression in miR-22-3p mimics group compared with miR NC group (Fig. 4C). Combining the results of RT-qPCR and Western Blot, we discovered that miR-22-3p reduced FABP5 protein expression in both two LIHC cell lines.
Prediction and analysis of upstream lncRNAs
To further investigate the molecular biological mechanisms of FABP5 in KIRC and LIHC, we explored the upstream lncRNAs of miR-577 and miR-22-3p, and then made corresponding analysis. In KIRC, although several lncRNAs had negative associations with miR-577, all of those associations were not significant (− 0.2 < cor < 0, Table 3). In LIHC, SNHG16, GUSBP11, FGD5-AS1, TTC28-AS1, LINC00630, CD27-AS1, LINC02381, H19 and LINC00997 presented notably negative correlations with miR-22-3p (Table 4). Among those lncRNAs, only seven lncRNAs, namely CD27-AS1, FGD5-AS1, GUSBP11, LINC00630, LINC00997, SNHG16 and TTC28-AS1, showed higher expression in LIHC tissues compared with normal tissues (all p < 0.001, Fig. 5A). Then we conducted correlation analysis of FABP5 with those seven lncRNAs. The results showed that only four lncRNAs, including CD27-AS1 (R = 0.23), GUSBP11 (R = 0.23), SNHG16 (R = 0.19) and TTC28-AS1 (R = 0.19), were positively associated with FABP5 (all p < 0.001, Fig. 5B). Survival analysis of above four lncRNAs (Fig. 5C) showed that high expression of GUSBP11 (p = 0.002), SNHG16 (p = 0.006) and TTC28-AS1 (p = 0.028) was statistically related to poor prognosis of LIHC patients.
(A) Expression analysis of CD27-AS1, FGD5-AS1, GUSBP11, LINC00630, LINC00997, SNHG16 and TTC28-AS1 in LIHC. (B) Correlation analysis of FABP5 with CD27-AS1, GUSBP11, SNHG16 and TTC28-AS1 in LIHC. (C) Overall survival analysis of CD27-AS1, GUSBP11, SNHG16 and TTC28-AS1 in LIHC. (D) FABP5-related regulatory network in KIRC. (E) FABP5-related ceRNA regulatory network in LIHC.
Combining those findings, we obtained two novel regulatory networks of FABP5, namely miR-577-FABP5 regulatory network in KIRC (Fig. 5D) and CD27-AS1/GUSBP11/SNHG16/TTC28-AS1-miR-22-3p-FABP5 ceRNA regulatory network in LIHC (Fig. 5E).
Correlation analysis of FABP5 with immune infiltration
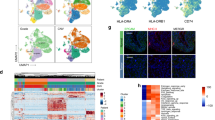
Using TIMER2 approach, we explored the potential relationship between FABP5 expression and immune infiltrations in different tumor types. Most of algorithms showed a positive association between B cell infiltration and FABP5 expression in KIRP, LIHC, PAAD and PRAD, but a negative relationship in BLCA, CESC, ESCA, LUSC, SKCM, SKCM-Metastasis, SKCM-Primary and TGCT (Fig. 6A). As for CD4+ T-cells, there was a positive correlation between the infiltration and FABP5 expression in LIHC and PAAD, but a negative correlation in HNSC, HNSC-HPV−, Sarcoma (SARC) and TGCT based on EPIC and TIMER (Fig. 6B). CD8+ T cell infiltration showed a positive association with FABP5 expression in CHOL, LGG, LIHC, LUAD, PAAD and UVM, but negative association in BLCA, LUSC, SKCM and SKCM-Metastasis based on most of algorithms (Fig. 6C). As shown in Fig. 6D, DCs’ infiltration was positively associated with FABP5 in BRCA, BRCA-LumA, BRCA-LumB, CHOL, KICH, KIRC, KIRP, LGG, LIHC, LUAD, PAAD, PCPG and THCA, but negatively in BLCA, LUSC, SKCM, TGCT and UCEC. Based on EPIC, TIMER and XCELL algorithms, infiltration of macrophages presented a positive relationship with FABP5 expression in BRCA-LumA, KIRP, LGG, LUAD and PAAD, but a negative relationship in BLCA, ESCA, LUSC and STAD (Fig. 6E). Among different types of macrophages, infiltration of M1 macrophages was positively associated with FABP5 expression in BRCA, BRCA-LumA, CESC, CHOL, KIRC, KIRP, LGG, LIHC, LUAD, THCA and UVM, but negatively in LUSC, SKCM and SKCM-Metastasis based on most of algorithms (Fig. 6F). As shown in Fig. 6G, M2 macrophage infiltration was positively related to FABP5 expression in BRCA-LumB and PRAD, but negatively in BLCA, ESCA, HNSC and HNSC-HPV− based on most of algorithms. Most of algorithms showed that infiltration of Tregs had a positive relationship with FABP5 expression in BRCA, BRCA-LumA, KIRC, LIHC, PAAD, PCPG and THCA, but a negative relationship in DLBC, ESCA, HNSC, HNSC-HPV−, HNSC-HPV+, LUSC, SKCM and TGCT (Fig. 6H). Moreover, based on most of algorithms, CAFs’ infiltration presented a positive relationship with FABP5 expression in ACC, BRCA, BRCA-Her2, BRCA-LumA, BRCA-LumB, KICH, KIRC, KIRP, PAAD, SKCM, SKCM-Metastasis, THCA and UVM, but a negative relationship in BLCA, BRCA-Basal, HNSC, HNSC-HPV−, PRAD, READ, STAD and THYM (Fig. 6I).
(A) Correlation between FABP5 and B cell infiltration. (B) Correlation between FABP5 and CD4+ T cell infiltration. (C) Correlation between FABP5 and CD8+ T cell infiltration. (D) Correlation between FABP5 and DC infiltration. (E) Correlation between FABP5 and macrophage infiltration. (F) Correlation between FABP5 and M1 macrophage infiltration. (G) Correlation between FABP5 and M2 macrophage infiltration. (H) Correlation between FABP5 and Treg infiltration. (I) Correlation between FABP5 and CAF infiltration.
Correlation analysis of FABP5 with immune checkpoints
We investigated the potential relationship between FABP5 expression and several immune checkpoints, including CD274, CTLA4, HAVCR2, LAG3, PDCD1 and TIGIT (Fig. 7A). In KIRC, FABP5 expression presented positive associations with the expression of PDCD1 (Rho = 0.104) and TIGIT (Rho = 0.093) (all p < 0.05, Fig. 7B). In LIHC, FABP5 expression was positively associated with the expression of CD274 (Rho = 0.245), CTLA4 (Rho = 0.342), HAVCR2 (Rho = 0.488), LAG3 (Rho = 0.282), PDCD1 (Rho = 0.358) and TIGIT (Rho = 0.379) (all p < 0.05, Fig. 7C). In addition to LIHC, FABP5 expression was positively correlated with expression of the above six immune checkpoints in BRCA, BRCA-LumA, KIRP, PAAD, PCPG, THCA and UVM.
Discussion
FABP5 has been reported to participate in the occurrence and development of different tumors, including KIRC, LIHC, LUAD and cervical cancer5,10,11,28. However, there is no study analyzing the potential roles of FABP5 based on a pan-cancer perspective. Therefore, we conducted a pan-cancer analysis of FABP5 based on the data from TCGA and GTEx database. We verified FABP5 overexpression in many tumor types, and partial results obtained support by previous studies10,29,30,31,32. Moreover, survival analysis showed an association between high FABP5 expression and poor prognosis in several tumor types, suggesting potential prognostic values of FABP5 in those tumors. Some of those results were consistent with the conclusions made by previous studies. High expression level of FABP5 was confirmed to predict poor prognosis in KIRC and LUAD28,31. Besides, FABP5 expression was considered as a prognostic marker of LIHC and UVM33,34.
Existing knowledge about FABP5-related regulatory networks is still limited, so further exploration of corresponding regulatory networks is needed. Additionally, further exploration may contribute to the discovery of new specific diagnostic and prognostic markers. To date, many studies have reported that ceRNA networks play a great role in multiple tumor types, providing a direction for exploring potential FABP5-related molecular mechanism. In this study, we combined the results of expression and survival analysis, and observed a situation. Only in GBM, KIRC and LIHC, FABP5 was overexpressed, and the overexpression was associated with poor OS prognosis shown by both Kaplan–Meier survival curve and Log-rank test. However, there is no corresponding miRNA transcriptome data of GBM available in the TCGA database. Thus, we did not include this tumor type in the scope of this study for the time being, and the subsequent analyses for FABP5-related upstream regulatory networks mainly focused on the other two tumor types. MicroRNAs mainly function as negative regulatory factors of corresponding genes. In this study, FABP5 functioned as an oncogenic factor in KIRC and LIHC, so upstream miRNAs of FABP5 should be antineoplastic and have significantly negative association with FABP5.
In KIRC, several reports have demonstrated the carcinogenic role of FABP57,11,31. However, we failed to find any articles about FABP5-related regulatory networks, so further exploration is really meaningful and necessary. Among those predicted miRNAs, only miR-577 was significantly negatively associated with FABP5 in KIRC. Although studies have confirmed miR-577’s antineoplastic role in many tumor types35,36,37,38, the certain role in KIRC has not been reported up to date. Based on our bioinformatics analysis, the expression of miR-577 was down-regulated, and KIRC patients with higher miR-577 expression had favorite outcomes. Meanwhile, we observed that as KIRC progresses, FABP5 expression increased. Therefore, down-regulated miR-577 may influence KIRC occurrence and progression via the regulation of FABP5 expression. Then we constructed miR-577-FABP5 axis and considered it as a regulator in KIRC occurrence and progression, but more experimental evidence is still needed to further improve the reliability of miR-577’s function and this axis’ regulatory effects in KIRC.
In LIHC, we found only one miRNA, namely miR-22-3p, significantly negatively associated with FABP5. Our bioinformatics analysis showed that miR-22-3p expression was down-regulated and lower expression was associated with poor prognosis in LIHC. Above results reflected the potential anti-tumor roles of miR-22-3p, which also obtained supports by the conclusion based on functional experiments in previous study that miR-22-3p suppressed tumor cell proliferation and arrested cell cycle in LIHC39,40. Combining those results, we constructed miR-22-3p-FABP5 axis, and considered it as one of potential regulatory networks in LIHC. Meanwhile, further experiment research was conducted to validate the potential miR-22-3p-FABP5 association in LIHC. MicroRNA-22-3p was verified to negatively regulate the expression of FABP5 via Western Blot. At present, there has been no report about the potential miR-22-3p-FABP5 relationship in any tumor type available. Our study verified the negative association between miR-22-3p and FABP5 in one tumor type for the first time, presenting a great novelty and providing a new direction for molecular mechanism exploration in other tumor types with high FABP5 expression. Moreover, it has been confirmed that down-regulated miR-22-3p expression promotes cell mobility and invasiveness of LIHC, and miR-22-3p overexpression has opposite effects41, suggesting the potential prognostic and therapeutic value of miR-22-3p. Therefore, we considered miR-22-3p-FABP5 axis as an important factor affecting the prognosis in LIHC, and further exploration of miR-22-3p-FABP5 axis benefits to the therapeutic and prognostic guidance of LIHC.
According to the definition of the ceRNA network, upstream lncRNAs are required to have negative associations with miR-22-3p, but positive associations with FABP5 in LIHC. Among those predicted lncRNAs of miR-22-3p, CD27-AS1, GUSBP11, SNHG16 and TTC28-AS1 met the above requirements, and all of those four lncRNAs were overexpressed in LIHC. Additionally, higher expression of GUSBP11, SNHG16 and TTC28-AS1 was statistically related to poor prognosis, and CD27-AS1 nearly reached statistical significance. A previous study has verified that CD27-AS1, GUSBP11 and SNHG16 serve as regulatory factors in LIHC through miR-22-3p-CCNA2 axis42. Our study came to a consistent conclusion that CD27-AS1, GUSBP11 and SNHG16 took miR-22-3p as a target to participate in the occurrence and progression of LIHC. Different from the previous study, we confirmed that FABP5 was another potential target of miR-22-3p in addition to CCNA2, reflecting that CD27-AS1/GUSBP11/SNHG16-miR-22-3p can affect the occurrence and development of LIHC via different pathways. Therefore, we further constructed CD27-AS1/GUSBP11/SNHG16-miR-22-3p-FABP5 ceRNA network, and considered it as a novel regulatory network in LIHC occurrence and progression. Furthermore, many studies have confirmed that SNHG16 promotes tumorigenesis, invasion and metastasis of LIHC by functioning as a sponge for multiple miRNAs43,44,45,46. The expression of SNHG16, an independent prognostic predictor, is associated with tumor size and sorafenib resistance of LIHC patients47,48,49. Therefore, SNHG16/miR-22-3p/FABP5 axis provides not only more possible prognostic indicators of LIHC, but also potential therapeutic targets for the improvement of chemotherapy’s effectiveness. TTC28-AS1 has been verified to have an association with epithelial–mesenchymal transition in bladder cancer50, but its role in LIHC progression remains uncertain. Thus, the view that TTC28-AS1 has carcinogenic effects in LIHC needs more experimental evidence to confirm.
In addition to FABP5 and FABP5-related regulatory networks, TME plays an important role in the occurrence and progression of tumors, and is involved in drug resistance14,15,51. Therefore, exploration of the potential relationship between FABP5 expression and immune infiltration provides possibilities for supplementing corresponding immunological mechanism and improving the therapeutic effects. Contrary to those antineoplastic immune cells including CD8+ T-cells, DCs and M1 macrophages, M2 macrophages and Tregs are considered to be tumor-promoting15,18,51,52. As for LIHC, we observed a positive relationship between FABP5 expression and the infiltration of Tregs. Due to FABP5 overexpression in LIHC tissues, high FABP5 expression may function as a carcinogenic factor by influencing Treg infiltration in LIHC. Tumor-promoting factors in TME also include CAFs in addition to M2 macrophages and Tregs17,52. In KIRC, FABP5 expression presented positive associations with the infiltration of Tregs and CAFs. Combining the results of expression analysis of FABP5, we considered a possibility that high FABP5 expression contributes to KIRC occurrence and progression by regulating the infiltration of Tregs and CAFs.
Moreover, our study conducted the evaluation of the potential relationship between FABP5 expression and six immune checkpoints. Among those checkpoints, PDCD1, CTLA4, LAG3 and HAVCR2 have been reported to show strong associations with each other, and affect the prognosis of tumors53. The positive relationship between FABP5 expression and all of those four immune checkpoints in BRCA, BRCA-LumA, KIRP, LIHC, PAAD, PCPG, THCA and UVM reflected that FABP5 may interact with those four immune checkpoints to participate in tumor occurrence and development. As for immunotherapy, antibodies targeting PD-1, PD-L1 and CTLA4 have been confirmed to make a significant improvement in the survival outcomes of patients with tumors, and the combination therapy targeting CTLA4 and PD-1 further improves the response rates and median survival of patients with many tumor types54,55. Besides, combination therapy inhibiting TIGIT and PD-1 made a significant improvement of OS in preclinical trials56. In this study, we observed that FABP5 expression was positively related to one of above-mentioned immune checkpoints in multiple tumor types, suggesting a possible synergy between FABP5 and one of above immune checkpoints. Further exploration of potential interactions between FABP5 and those immune checkpoints and corresponding experiments are needed to verify whether FABP5 can be a target for immunotherapy.
Taken together, we made a pan-cancer analysis of FABP5 based on TCGA and GTEx database for the first time, and confirmed FABP5 overexpression in most tumor types. We also explored the potential prognostic values of FABP5, and observed that high FABP5 expression was associated with poor outcomes of patients with most of tumor types. Additionally, we further investigated FABP5-related regulatory networks and constructed miR-577-FABP5 regulatory network in KIRC as well as CD27-AS1/GUSBP11/SNHG16/TTC28-AS1-miR-22-3p-FABP5 ceRNA regulatory network in LIHC. Among those FABP5-related regulatory networks, the association between miR-22-3p and FABP5 in LIHC got discovered and confirmed via miRNA-mRNA interaction prediction and experiments for the first time. Moreover, the potential relationships of FABP5 expression with immune cell infiltration and immune checkpoints in multiple tumor types were investigated. Our work not only helps to further understand functions of FABP5 and supplement existing FABP5-related mechanisms in tumors, but also provides more possibilities for immunotherapy.
Methods
Data source
TCGA (http://cancergenome.nih.gov) database is a cancer genomics program characterizing more than twenty thousand primary cancers and matching normal samples of 33 tumor types. Our study employed UCSC Xena (https://xenabrowser.net/), a browser interactively visualizing cancer genome datasets, to collect gene expression RNA sequence and clinical data across TCGA tumor types57. In several tumor types, less than five normal samples are included in TCGA database, so we analyzed the difference in FABP5 expression between tumor and normal tissues in these tumor types after adding normal samples from Genotype-Tissue Expression (GTEx, http://commonfund.nih.gov/GTEx/) database.
Expression analysis
Perl script and R language were applied for data organization. Using R language, we made expression analysis of FABP5 in tumor types with at least five adjacent normal tissues in TCGA database. |log2FC| > 1 and the adjusted p-value < 0.05 were regarded as the cut-off criterion. As for those tumor types with less than five normal samples in TCGA database, we applied Gene Expression Profiling Interactive Analysis (GEPIA, http://gepia.cancer-pku.cn/) to compare FABP5 expression between tumor and normal tissues based on TCGA and GTEx database. Moreover, the “Stage Plot” module of GEPIA was applied to explore the correlations between the expression level of FABP5 and pathological stages of different tumors.
Survival analysis
Using median FABP5 expression level as the cutoff, tumor samples were divided into the high-expression and low-expression groups. The “Survival Plots” module of GEPIA was applied to compare the OS and DFS between high-expression and low-expression group. Additionally, we used “survival” package in R to make survival analysis by the Kaplan–Meier survival curve and Log-rank test in tumor types that have at least five corresponding normal tissues in TCGA database. Only tumor types with statistically significant results of survival analysis by both Kaplan–Meier survival curve and Log-rank test will be chosen for further exploration of FABP5-related regulatory network. Moreover, we utilized “survival” and “survminer” packages in R to make survival analysis of predicted miRNAs and lncRNAs by Kaplan–Meier survival curve.
Prediction of upstream miRNA and lncRNA
Encyclopedia of RNA Interactomes (ENCORI, http://starbase.sysu.edu.cn/), a database that integrates RNA interactions, was applied to predict miRNA-mRNA and lncRNA-miRNA interactions58. All interactions obtained by ENCORI have been supported by Ago CLIP-seq data. Based on the positive results of at least one miRNA-target predicting databases including microT, miRanda, miRmap, PicTar, PITA, RNA22 and TargetScan, FABP5-related miRNAs were identified. Besides, upstream lncRNAs of FABP5-related miRNAs were predicted by iRanda program.
Analysis of immune infiltration and immune checkpoints
Tumor Immune Estimation Resource Version 2 (TIMER2.0, http://timer.comp-genomics.org/) provides comprehensive analyses of immune infiltrates and visualizes their functions in multiple tumor types59. In this study, TIMER2.0 was used for the evaluation of relationships between FABP5 expression and immune infiltration (B cells, CD4+ T-cells, CD8+ T-cells, DCs, macrophages, Tregs and CAFs) across TCGA tumor types. Besides, the “gene correlation” module of TIMER2.0 was used to explore associations between FABP5 and immune checkpoints (CD274, PDCD1, CTLA-4, LAG-3, HAVCR2 and TIGIT). P-values and partial rho values were obtained by purity-adjusted Spearman’s rank correlation test.
Correlation analysis
R language was applied for evaluating the relationships between FABP5 and FABP5-bound miRNAs, FAPB5-bound miRNAs and corresponding lncRNAs. Spearman’s rank correlation test was used to obtain the p-values and partial rho values.
Cell culture
Two different hepatocarcinoma cell lines (Hep3B and HepG2) were respectively cultured with their own special mediums (Procell, China) in a 37 ℃ incubator with 5% CO2. Preparing for the subsequent transfection assay, cells were plated into 24 mm six-well plates at a density of 3 × 105 cell/ml and cultured for 24 h.
Transfection assay
MicroRNA-22-3p mimics, nonspecific miR-NC oligo and the transfection reagent were obtained from GenePharma (Shanghai, China). Hepatocarcinoma cells in each six-well plate were divided into two groups, miR NC group and miR-22-3p mimics group. Using the transfection reagent, miR NC group got transfected with miR-NC, and miR-22-3p mimics group got transfected with miR-22-3p mimics. All cells after treatments were cultured at 37 ℃, 5% CO2 for 72 h, and then collected for further analysis. Specific sequences were as follows: miR-22-3p mimics sense: 5′-3′AAGCUGCCAGUUGAAGAACUGU and antisense: 5′-3′AGUUCUUCAACUGGCAGCUUUU. miR-NC sense: 5′-3′UUCUCCGAACGUGUCACGUTT and antisense: 5′-3′ACGUGACACGUUCGGAGAATT.
RT-qPCR
Total RNA from cultured cells of different groups was obtained by Ezol reagent (GenePharma, China), and purified via column passing method. Reverse transcription and quantitative real-time PCR (qPCR) were conducted via Hairpin-it™ microRNA and U6 snRNA Normalization RT-PCR Quantitation Kit (GenePharma, China) according to the manufacturer's instructions. The primer sequences for RT-qPCR were as follows: HmiR-22-3p-FO-2: 5′-3′GCGGTCAAGCTGCCAGTT and HmiR-22-3p-RE-2: 5′-3′TATGGTTGTTCACGACTCCTTCAC. U6 forward: 5′-3′CGCTTCGGCAGCACATATAC and reverse: 5′-3′TTCACGAATTTGCGTGTCATC.
Western blot
Total protein from cultured cells of different groups was extracted by using Whole Protein Extraction Kit (Solarbio Life Science, China) according to the manufacturer's instructions. Then, we measured the protein concentration by using the bicinchoninic acid (BCA) protein assay kit (Elabscience, China). According to the molecular weight of FABP5, 12.5% SDS-PAGE gels were prepared by using Omni-Easy™ One-Step PAGE Gel Fast Preparation Kit (Yamay, China) for the separation of proteins. After protein separation, we transferred proteins onto PVDF membranes, and blocked membranes with QuickBlock™ Blocking Buffer (Beyotime, China). Then, membranes got incubated with corresponding primary antibodies, including antibodies against FABP5 (1:1000, Biodragon, China) and β-actin (1:20,000, ABclonal, China), at 4 ℃ overnight. We further incubated membranes with secondary antibodies (1:1000, Beyotime, China) for 2 h after washing them. After treated by ECL chemiluminescence method, the bands of the membranes got revealed.
Data availability
Publicly available datasets were analyzed in this study, these can be found in UCSC Xena (https://xenabrowser.net/). The ID numbers included: TCGA-BLCA.htseq_fpkm.tsv, TCGA-BRCA. htseq_fpkm.tsv, TCGA-CHOL.htseq_fpkm.tsv, TCGA-COAD.htseq_fpkm.tsv, TCGA-ESCA.htseq_fpkm.tsv, TCGA-GBM.htseq_fpkm.tsv, TCGA-HNSC. htseq_fpkm.tsv, TCGA-KICH.htseq_fpkm.tsv, TCGA-KIRC.htseq_fpkm.tsv, TCGA-KIRP.htseq_fpkm.tsv, TCGA-LIHC.htseq_fpkm.tsv, TCGA-LUAD. htseq_fpkm.tsv, TCGA-LUSC.htseq_fpkm.tsv, TCGA-PRAD.htseq_fpkm.tsv, TCGA-READ.htseq_fpkm.tsv, TCGA-STAD.htseq_fpkm.tsv, TCGA-THCA. htseq_fpkm.tsv, TCGA-UCEC.htseq_fpkm.tsv. Some of the data were stored in the Supplementary Data. The authors confirm that the data supporting the findings of this study are available within the article and its Supplementary materials.
References
Hotamisligil, G. S. & Bernlohr, D. A. Metabolic functions of FABPs-mechanisms and therapeutic implications. Nat. Rev. Endocrinol. 11, 592–605. https://doi.org/10.1038/nrendo.2015.122 (2015).
Li, B., Hao, J., Zeng, J. & Sauter, E. R. SnapShot: FABP functions. Cell 182, 1066-1066.e1061. https://doi.org/10.1016/j.cell.2020.07.027 (2020).
Jing, C. et al. Identification of the messenger RNA for human cutaneous fatty acid-binding protein as a metastasis inducer. Cancer Res. 60, 2390–2398 (2000).
Jing, C. et al. Human cutaneous fatty acid-binding protein induces metastasis by up-regulating the expression of vascular endothelial growth factor gene in rat Rama 37 model cells. Cancer Res. 61, 4357–4364 (2001).
Zhang, C. et al. FABP5 promotes lymph node metastasis in cervical cancer by reprogramming fatty acid metabolism. Theranostics 10, 6561–6580. https://doi.org/10.7150/thno.44868 (2020).
Pan, J., Dai, Q., Zhang, T. & Li, C. Palmitate acid promotes gastric cancer metastasis via FABP5/SP1/UCA1 pathway. Cancer Cell Int. 19, 69. https://doi.org/10.1186/s12935-019-0787-0 (2019).
Lv, Q. et al. FABP5 regulates the proliferation of clear cell renal cell carcinoma cells via the PI3K/AKT signaling pathway. Int. J. Oncol. 54, 1221–1232. https://doi.org/10.3892/ijo.2019.4721 (2019).
Kawaguchi, K. et al. The cancer-promoting gene fatty acid-binding protein 5 (FABP5) is epigenetically regulated during human prostate carcinogenesis. Biochem. J. 473, 449–461. https://doi.org/10.1042/BJ20150926 (2016).
Seo, J., Yun, J., Fukuda, J. & Chun, Y. S. Tumor-intrinsic FABP5 is a novel driver for colon cancer cell growth via the HIF-1 signaling pathway. Cancer Genet. 258–259, 151–156. https://doi.org/10.1016/j.cancergen.2021.11.001 (2021).
Seo, J. et al. Fatty-acid-induced FABP5/HIF-1 reprograms lipid metabolism and enhances the proliferation of liver cancer cells. Commun. Biol. 3, 638. https://doi.org/10.1038/s42003-020-01367-5 (2020).
Wu, G. et al. FABP5 is correlated with poor prognosis and promotes tumour cell growth and metastasis in clear cell renal cell carcinoma. Eur. J. Pharmacol. 862, 172637. https://doi.org/10.1016/j.ejphar.2019.172637 (2019).
Wang, W. et al. Downregulation of FABP5 suppresses the proliferation and induces the apoptosis of gastric cancer cells through the hippo signaling pathway. DNA Cell Biol. 40, 1076–1086. https://doi.org/10.1089/dna.2021.0370 (2021).
Liu, R. Z. et al. Association of FABP5 expression with poor survival in triple-negative breast cancer: Implication for retinoic acid therapy. Am. J. Pathol. 178, 997–1008. https://doi.org/10.1016/j.ajpath.2010.11.075 (2011).
Arneth, B. Tumor microenvironment. Medicina (Kaunas). https://doi.org/10.3390/medicina56010015 (2019).
Lei, X. et al. Immune cells within the tumor microenvironment: Biological functions and roles in cancer immunotherapy. Cancer Lett. 470, 126–133. https://doi.org/10.1016/j.canlet.2019.11.009 (2020).
Hui, L. & Chen, Y. Tumor microenvironment: Sanctuary of the devil. Cancer Lett. 368, 7–13. https://doi.org/10.1016/j.canlet.2015.07.039 (2015).
Denton, A. E., Roberts, E. W. & Fearon, D. T. Stromal cells in the tumor microenvironment. Adv. Exp. Med. Biol. 1060, 99–114. https://doi.org/10.1007/978-3-319-78127-3_6 (2018).
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437. https://doi.org/10.1038/nm.3394 (2013).
Jiang, X. et al. Role of the tumor microenvironment in PD-L1/PD-1-mediated tumor immune escape. Mol. Cancer 18, 10. https://doi.org/10.1186/s12943-018-0928-4 (2019).
Andrews, L. P., Marciscano, A. E., Drake, C. G. & Vignali, D. A. LAG3 (CD223) as a cancer immunotherapy target. Immunol. Rev. 276, 80–96. https://doi.org/10.1111/imr.12519 (2017).
Das, M., Zhu, C. & Kuchroo, V. K. Tim-3 and its role in regulating anti-tumor immunity. Immunol. Rev. 276, 97–111. https://doi.org/10.1111/imr.12520 (2017).
Manieri, N. A., Chiang, E. Y. & Grogan, J. L. TIGIT: A key inhibitor of the cancer immunity cycle. Trends Immunol. 38, 20–28. https://doi.org/10.1016/j.it.2016.10.002 (2017).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P. P. A ceRNA hypothesis: The Rosetta Stone of a hidden RNA language?. Cell 146, 353–358. https://doi.org/10.1016/j.cell.2011.07.014 (2011).
Qi, X. et al. ceRNA in cancer: Possible functions and clinical implications. J. Med. Genet. 52, 710–718. https://doi.org/10.1136/jmedgenet-2015-103334 (2015).
Chan, J. J. & Tay, Y. Noncoding RNA:RNA regulatory networks in cancer. Int. J. Mol. Sci. https://doi.org/10.3390/ijms19051310 (2018).
Wang, X. et al. Oncogenic role and potential regulatory mechanism of topoisomerase IIα in a pan-cancer analysis. Sci. Rep. 12, 11161. https://doi.org/10.1038/s41598-022-15205-7 (2022).
Sun, W. Y. et al. Circ-ABCB10 acts as an oncogene in glioma cells via regulation of the miR-620/FABP5 axis. Eur. Rev. Med. Pharmacol. Sci. 24, 6848–6857. https://doi.org/10.26355/eurrev_202006_21674 (2020).
Garcia, K. A. et al. Fatty acid binding protein 5 regulates lipogenesis and tumor growth in lung adenocarcinoma. Life Sci. 301, 120621. https://doi.org/10.1016/j.lfs.2022.120621 (2022).
Jeong, C. Y. et al. Fatty acid-binding protein 5 promotes cell proliferation and invasion in human intrahepatic cholangiocarcinoma. Oncol. Rep. 28, 1283–1292. https://doi.org/10.3892/or.2012.1922 (2012).
Han, J. et al. Identification of potential therapeutic targets in human head & neck squamous cell carcinoma. Head Neck Oncol. 1, 27. https://doi.org/10.1186/1758-3284-1-27 (2009).
Wu, G. et al. Study of FABP’s interactome and detecting new molecular targets in clear cell renal cell carcinoma. J. Cell Physiol. 235, 3776–3789. https://doi.org/10.1002/jcp.29272 (2020).
Carbonetti, G. et al. FABP5 coordinates lipid signaling that promotes prostate cancer metastasis. Sci. Rep. 9, 18944. https://doi.org/10.1038/s41598-019-55418-x (2019).
Xu, Y., Xu, W. H., Yang, X. L., Zhang, H. L. & Zhang, X. F. Fatty acid-binding protein 5 predicts poor prognosis in patients with uveal melanoma. Oncol. Lett. 19, 1771–1780. https://doi.org/10.3892/ol.2020.11301 (2020).
Ohata, T. et al. Fatty acid-binding protein 5 function in hepatocellular carcinoma through induction of epithelial-mesenchymal transition. Cancer Med. 6, 1049–1061. https://doi.org/10.1002/cam4.1020 (2017).
Men, L., Nie, D. & Nie, H. microRNA-577 inhibits cell proliferation and invasion in non-small cell lung cancer by directly targeting homeobox A1. Mol. Med. Rep. 19, 1875–1882. https://doi.org/10.3892/mmr.2019.9804 (2019).
Jiang, H., Ju, H., Zhang, L., Lu, H. & Jie, K. microRNA-577 suppresses tumor growth and enhances chemosensitivity in colorectal cancer. J. Biochem. Mol. Toxicol. https://doi.org/10.1002/jbt.21888 (2017).
Yu, Z., Zhang, W. & Deng, F. MicroRNA-577 inhibits gastric cancer growth by targeting E2F transcription factor 3. Oncol. Lett. 10, 1447–1452. https://doi.org/10.3892/ol.2015.3390 (2015).
Yuan, X., He, J., Sun, F. & Gu, J. Effects and interactions of MiR-577 and TSGA10 in regulating esophageal squamous cell carcinoma. Int. J. Clin. Exp. Pathol. 6, 2651–2667 (2013).
Kong, D., Wang, X., Wang, X., Wang, Z. & Wang, F. Downregulated miRNA-22-3p promotes the progression and leads to poor prognosis of hepatocellular carcinoma through targeting CDKN2C. J. Buon. 26, 409–417 (2021).
Zhou, X. et al. Long noncoding RNA NEAT1 promotes cell proliferation and invasion and suppresses apoptosis in hepatocellular carcinoma by regulating miRNA-22-3p/akt2 in vitro and in vivo. Onco. Targets Ther. 12, 8991–9004. https://doi.org/10.2147/OTT.S224521 (2019).
Zhang, D. Y. et al. Identification and functional characterization of long non-coding RNA MIR22HG as a tumor suppressor for hepatocellular carcinoma. Theranostics 8, 3751–3765. https://doi.org/10.7150/thno.22493 (2018).
Chen, S. et al. The predictive competing endogenous RNA regulatory networks and potential prognostic and immunological roles of cyclin A2 in pan-cancer analysis. Front. Mol. Biosci. 9, 809509. https://doi.org/10.3389/fmolb.2022.809509 (2022).
Lin, Q., Zheng, H., Xu, J., Zhang, F. & Pan, H. LncRNA SNHG16 aggravates tumorigenesis and development of hepatocellular carcinoma by sponging miR-4500 and targeting STAT3. J. Cell Biochem. https://doi.org/10.1002/jcb.28440 (2019).
Xie, X., Xu, X., Sun, C. & Yu, Z. Long intergenic noncoding RNA SNHG16 interacts with miR-195 to promote proliferation, invasion and tumorigenesis in hepatocellular carcinoma. Exp. Cell Res. 383, 111501. https://doi.org/10.1016/j.yexcr.2019.111501 (2019).
Hu, Y. L. et al. SNHG16/miR-605-3p/TRAF6/NF-κB feedback loop regulates hepatocellular carcinoma metastasis. J. Cell Mol. Med. 24, 7637–7651. https://doi.org/10.1111/jcmm.15399 (2020).
Li, W., Xu, W., Song, J. S., Wu, T. & Wang, W. X. LncRNA SNHG16 promotes cell proliferation through miR-302a-3p/FGF19 axis in hepatocellular carcinoma. Neoplasma 66, 397–404. https://doi.org/10.4149/neo_2018_180720N504 (2019).
Guo, Z. et al. Long noncoding RNA (lncRNA) small nucleolar RNA host gene 16 (SNHG16) predicts poor prognosis and sorafenib resistance in hepatocellular carcinoma. Med. Sci. Monit. 25, 2079–2086. https://doi.org/10.12659/MSM.915541 (2019).
Zhong, J. H. et al. The lncRNA SNHG16 affects prognosis in hepatocellular carcinoma by regulating p62 expression. J. Cell Physiol. 235, 1090–1102. https://doi.org/10.1002/jcp.29023 (2020).
Ye, J., Zhang, R., Du, X., Chai, W. & Zhou, Q. Long noncoding RNA SNHG16 induces sorafenib resistance in hepatocellular carcinoma cells through sponging miR-140-5p. Onco. Targets Ther. 12, 415–422. https://doi.org/10.2147/ott.S175176 (2019).
Tong, H. et al. An epithelial-mesenchymal transition-related long noncoding RNA signature correlates with the prognosis and progression in patients with bladder cancer. Biosci. Rep. https://doi.org/10.1042/bsr20203944 (2021).
Khalaf, K. et al. Aspects of the tumor microenvironment involved in immune resistance and drug resistance. Front. Immunol. 12, 656364. https://doi.org/10.3389/fimmu.2021.656364 (2021).
Farhood, B., Najafi, M. & Mortezaee, K. CD8+ cytotoxic T lymphocytes in cancer immunotherapy: A review. J. Cell. Physiol. 234(6), 8509–8521. https://doi.org/10.1002/jcp.27782 (2019).
Tu, L. et al. Assessment of the expression of the immune checkpoint molecules PD-1, CTLA4, TIM-3 and LAG-3 across different cancers in relation to treatment response, tumor-infiltrating immune cells and survival. Int. J. Cancer 147, 423–439. https://doi.org/10.1002/ijc.32785 (2020).
Ghahremanloo, A., Soltani, A., Modaresi, S. M. S. & Hashemy, S. I. Recent advances in the clinical development of immune checkpoint blockade therapy. Cell Oncol. (Dordr.) 42, 609–626. https://doi.org/10.1007/s13402-019-00456-w (2019).
Rotte, A. Combination of CTLA-4 and PD-1 blockers for treatment of cancer. J. Exp. Clin. Cancer Res. 38, 255. https://doi.org/10.1186/s13046-019-1259-z (2019).
Solomon, B. L. & Garrido-Laguna, I. TIGIT: A novel immunotherapy target moving from bench to bedside. Cancer Immunol. Immunother. 67, 1659–1667. https://doi.org/10.1007/s00262-018-2246-5 (2018).
Goldman, M. J. et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 38, 675–678. https://doi.org/10.1038/s41587-020-0546-8 (2020).
Li, J. H., Liu, S., Zhou, H., Qu, L. H. & Yang, J. H. starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 42, D92–D97. https://doi.org/10.1093/nar/gkt1248 (2014).
Liu, X. S. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48, W509–W514. https://doi.org/10.1093/nar/gkaa407 (2020).
Funding
This study was funded by National Natural Science Foundation of China (81800203), Dalian Deng Feng Program (243), and Doctoral Scientific Research Foundation of Liaoning Province (20180540088).
Author information
Authors and Affiliations
Contributions
J.W. and J.S. performed the data analysis and interpreted the data. S.Z., X.W. and M.G. prepared the draft. J.W. and S.Z. performed the visualization and revised the draft. B.T. and J.Y. designed the research and supervised all the work. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, J., Zhao, S., Sun, J. et al. Oncogenic role and potential regulatory mechanism of fatty acid binding protein 5 based on a pan-cancer analysis. Sci Rep 13, 4060 (2023). https://doi.org/10.1038/s41598-023-30695-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-30695-9
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.