Abstract
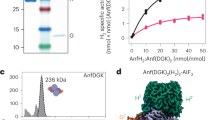
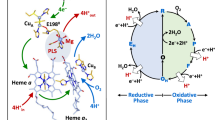
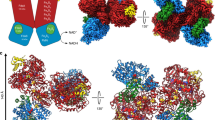
The structure of the respiratory nitrate reductase (NapAB) from Rhodobacter sphaeroides, the periplasmic heterodimeric enzyme responsible for the first step in the denitrification process, has been determined at a resolution of 3.2 Å. The di-heme electron transfer small subunit NapB binds to the large subunit with heme II in close proximity to the [4Fe-4S] cluster of NapA. A total of 57 residues at the N- and C-terminal extremities of NapB adopt an extended conformation, embracing the NapA subunit and largely contributing to the total area of 5,900 Å2 buried in the complex. Complex formation was studied further by measuring the variation of the redox potentials of all the cofactors upon binding. The marked effects observed are interpreted in light of the three-dimensional structure and depict a plasticity that contributes to an efficient electron transfer in the complex from the heme I of NapB to the molybdenum catalytic site of NapA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Blasco, F. et al. The coordination and function of the redox centres of the membrane-bound nitrate reductases. Cell. Mol. Life Sci. 58, 179–193 (2001).
Richardson, D.J., Berks, B.C., Russell, D.A., Spiro, S. & Taylor, C.J. Functional, biochemical and genetic diversity of prokaryotic nitrate reductases. Cell. Mol. Life Sci. 58, 165–178 (2001).
Sabaty, M., Schwintner, C., Cahors, S., Richaud, P. & Verméglio, A. Nitrite and nitrous oxide reductase regulation by nitrogen oxides in Rhodobacter sphaeroides f. sp. denitrificans IL106. J. Bacteriol. 181, 6028–6032 (1999).
Hille, R. Molybdenum and tungsten in biology. Trends Biochem. Sci. 27, 360–367 (2002).
Richardson, D.J. et al. The diversity of redox proteins involved in bacterial heterotrophic nitrification and aerobic denitrification. Biochem. Soc. Trans. 26, 401–408 (1998).
Anderson, G.L., Williams, J. & Hille, R. The purification and characterization of arsenite oxidase from Alcaligenes faecalis, a molybdenum-containing hydroxylase. J. Biol. Chem. 267, 23674–23682 (1992).
Cartron, M.L., Roldan, M.D., Ferguson, S.J., Berks, B.C. & Richardson, D.J. Identification of two domains and distal histidine ligands to the four haems in the bacterial c-type cytochrome NapC; the prototype connector between quinol/quinone and periplasmic oxido-reductases. Biochem. J. 368, 425–432 (2002).
Brigé, A., Leys, D., Meyer, T.E., Cusanovich, M.A. & Van Beeumen, J.J. The 1.25 Å resolution structure of the diheme NapB subunit of soluble nitrate reductase reveals a novel cytochrome c fold with a stacked heme arrangement. Biochemistry 41, 4827–4836 (2002).
Dias, J.M. et al. Crystal structure of the first dissimilatory nitrate reductase at 1.9 Å solved by MAD methods. Struct. Fold. Des. 7, 65–79 (1999).
Sargent, F., Berks, B.C. & Palmer T. Assembly of membrane-bound respiratory complexes by the Tat protein-transport system. Arch. Microbiol. 178, 77–84 (2002).
Bennett, B., Berks, B.C., Ferguson, S.J., Thomson, A.J. & Richardson, D.J. Mo(V) EPR signals from the periplasmic nitrate reductase of Thiosphaera pantotropha. Eur. J. Biochem. 226, 789–798 (1994).
Bursakov, S.A. et al. Isolation and preliminary characterization of a soluble nitrate reductase from the sulfate-reducing organism Desulfovibrio desulfuricans ATCC 27774. Anaerobe 1, 55–60 (1995).
Lo Conte, L., Chothia, C. & Janin, J. The atomic structure of protein–protein recognition sites. J. Mol. Biol. 285, 2177–2198 (1999).
Lange, C. & Hunte, C. Crystal structure of the yeast cytochrome bc1 complex with its bound substrate cytochrome c. Proc. Natl. Acad. Sci. USA 99, 2800–2805 (2002).
Axelrod, H.L. et al. X-ray structure determination of the cytochrome c2: reaction center electron transfer complex from Rhodobacter sphaeroides. J. Mol. Biol. 319, 501–515 (2002).
Page, C.C., Moser, C.C., Chen, X. & Dutton, P.L. Natural engineering principles of electron tunnelling in biological oxidation-reduction. Nature 402, 47–52 (1999).
Pelletier, H. & Kraut, J. Crystal structure of a complex between electron transfer partners, cytochrome c peroxidase and cytochrome c. Science 258, 1748–1755 (1992).
Bamford, V.A. et al. Structural basis for the oxidation of thiosulfate by a sulfur cycle enzyme. EMBO J. 21, 5599–5610 (2002).
Morelli, X. et al. Structural model of the Fe-hydrogenase/cytochrome c553 complex combining transverse relaxation-optimized spectroscopy experiments and soft docking calculations. J. Biol. Chem. 275, 23204–23210 (2000).
Morelli, X. et al. Heteronuclear NMR and soft docking: an experimental approach for a structural model of the cytochrome c553–ferredoxin complex. Biochemistry 39, 2530–2537 (2000).
Deisenhofer, J., Epp, O., Sinning, I. & Michel, H. Crystallographic refinement at 2.3 Å resolution and refined model of the photosynthetic reaction centre from Rhodopseudomonas viridis. J. Mol. Biol. 246, 429–457 (1995).
Dohse, B. et al. Electron transfer from the tetraheme cytochrome to the special pair in the Rhodopseudomonas viridis reaction center: effect of mutations of tyrosine L162. Biochemistry 34, 11335–11343 (1995).
Farchaus, J.W., Wachtveitl, J., Mathis, P. & Oesterhelt, D. Tyrosine 162 of the photosynthetic reaction center L-subunit plays a critical role in the cytochrome c2-mediated rereduction of the photooxidized bacteriochlorophyll dimer in Rhodobacter sphaeroides. 1. Site-directed mutagenesis and initial characterization. Biochemistry 32, 10885–10893 (1993).
Wachtveitl, J., Farchaus, J.W., Mathis, P. & Oesterhelt, D. Tyrosine 162 of the photosynthetic reaction center L-subunit plays a critical role in the cytochrome c2-mediated rereduction of the photooxidized bacteriochlorophyll dimer in Rhodobacter sphaeroides. 2. Quantitative kinetic analysis. Biochemistry 32, 10894–10904 (1993).
Berks, B.C. et al. Purification and characterization of the periplasmic nitrate reductase from Thiosphaera pantotropha. Eur. J. Biochem. 220, 117–124 (1994).
Moura, I., Bursakov, S.A., Costa, C. & Moura, J.J. Nitrate and nitrite utilization in sulfate-reducing bacteria. Anaerobe 3, 279–290 (1997).
Potter, L.C., Millington, P., Griffiths, L., Thomas, G.H. & Cole, J.A. Competition between Escherichia coli strains expressing either a periplasmic or a membrane-bound nitrate reductase: does Nap confer a selective advantage during nitrate-limited growth? Biochem. J. 344, 77–84 (1999).
Reyes, F., Gavira, M., Castillo, F. & Moreno-Vivian, C. Periplasmic nitrate-reducing system of the phototrophic bacterium Rhodobacter sphaeroides DSM 158: transcriptional and mutational analysis of the napKEFDABC gene cluster. Biochem. J. 331, 897–904 (1998).
Buc, J. et al. Enzymatic and physiological properties of the tungsten-substituted molybdenum TMAO reductase from Escherichia coli. Mol. Microbiol. 32, 159–168 (1999).
Sabaty, M., Avazeri, C., Pignol, D. & Verméglio, A. Characterization of the reduction of selenate and tellurite by nitrate reductases. Appl. Environ. Microbiol. 67, 5122–5126 (2001).
Pignol, D., Adriano, J.M., Fontecilla-Camps, J.C. & Sabaty, M. Crystallization and preliminary X-ray analysis of the periplasmic nitrate reductase (NapA–NapB complex) from Rhodobacter sphaeroides f. sp. denitrificans. Acta Crystallogr. D 57, 1900–1902 (2001).
Leslie, A.G.W. Autoindexing of rotation diffraction images and parameter refinement, in Proceedings of the CCP4 Study Weekend: 'Data Collection and Processing' (eds. Sawyer, L., Isaacs, N. & Bailey, S.) 44–51 (SERC, Daresbury Laboratory, Daresbury, UK, 1993).
Collaborative Computational Project, Number 4. The CCP4 Suite: Programs for Protein Crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968).
Navazza, J. AmoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Roussel, A. & Cambillau, C. In Silicon Graphics Geometry Partners Directory (Silicon Graphics, Mountain View, California, USA, 1991).
Murshudov, G.N., Vagin, A.A., Lebedev, A., Wilson, K.S. & Dodson, E.J. Efficient anisotropic refinement of macromolecular structures using FFT. Acta Crystallogr. D 55, 247–255 (1999).
Joliot, P., Béal, D. & Frilley, B. Une nouvelle méthode spectrophotométrique destinée à l'étude des réactions photosynthétiques. J. Chim. Phys. 77, 209–216 (1980).
Acknowledgements
We would like to thank the European Synchrotron Radiation Facility and especially the staff of the BM30 and ID14 beamlines (Grenoble, France) for the provision of data collection and processing facilities, the staff of the Laboratoire de Cristallographie et de Cristallogenèse des Protéines of the Institut de Biologie Structurale for encouragement, the Progamme de Toxicologie Nucléaire of the Comissariat à l'Energie atomique for financial support, and J. Lavergne and A. Verméglio for discussions and reading of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Arnoux, P., Sabaty, M., Alric, J. et al. Structural and redox plasticity in the heterodimeric periplasmic nitrate reductase. Nat Struct Mol Biol 10, 928–934 (2003). https://doi.org/10.1038/nsb994
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb994
This article is cited by
-
NapB Restores cytochrome c biosynthesis in bacterial dsbD-deficient mutants
Communications Biology (2022)
-
Functional mononuclear molybdenum enzymes: challenges and triumphs in molecular cloning, expression, and isolation
JBIC Journal of Biological Inorganic Chemistry (2020)
-
Salt tolerance of nitrate reductase in Halomonas sp. B01
Folia Microbiologica (2020)
-
Multi-omics reveal various potential antimonate reductases from phylogenetically diverse microorganisms
Applied Microbiology and Biotechnology (2019)
-
EXD2 governs germ stem cell homeostasis and lifespan by promoting mitoribosome integrity and translation
Nature Cell Biology (2018)