Abstract
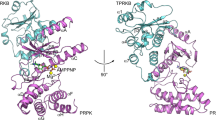
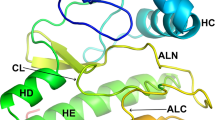
We have determined X-ray crystal structures with up to 1.5 Å resolution of the catalytic domain of death-associated protein kinase (DAPK), the first described member of a novel family of pro-apoptotic and tumor-suppressive serine/threonine kinases. The geometry of the active site was studied in the apo form, in a complex with nonhydrolyzable AMPPnP and in a ternary complex consisting of kinase, AMPPnP and either Mg2+ or Mn2+. The structures revealed a previously undescribed water-mediated stabilization of the interaction between the lysine that is conserved in protein kinases and the β- and γ-phosphates of ATP, as well as conformational changes at the active site upon ion binding. Comparison between these structures and nucleotide triphosphate complexes of several other kinases disclosed a number of unique features of the DAPK catalytic domain, among which is a highly ordered basic loop in the N-terminal domain that may participate in enzyme regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Laster, S.M., Wood, J.G. & Gooding, L.R. Tumor necrosis factor can induce both apoptotic and necrotic forms of cell lysis. J. Imunnol. 141, 2629–2634 (1988).
Trauth, B.C. et al. Monoclonal antibody-mediated tumor regression by induction of apoptosis. Science 245, 301–305 (1989).
Itoh, N. et al. The polypeptide encoded by the cDNA for human cell surface antigen Fas can mediate apoptosis. Cell 66, 233–243 (1991).
Lin, J.K. & Chou, C.K. In vitro apoptosis in the human hepatoma cell line induced by transforming growth factor β1. Cancer Res. 52, 385–388 (1992).
Novelli, F. et al. Environmental signals influencing expression of the IFN-γ receptor on human T cells control whether IFN-γ promotes proliferation of apoptosis. J. Immunol. 152, 496–504 (1994).
Gjertsen, B. & Doskeland, S. Protein phosphorylation in apoptosis. Biochim. Biophys. Acta 1269, 187–189 (1995).
Anderson, P. Kinase cascades regulating entry into apoptosis. Microbiol. Molec. Biol. Rev. 61, 33–46 (1997).
Inbal, B. et al. DAP kinase links the control of apoptosis to metastasis. Nature 390, 180–184 (1997).
Raveh, T., Droguett, G., Horvitz, M.S., DePinho, R.A. & Kimchi, A. DAP kinase activates a p19ARF/p53-mediated apoptotic checkpoint to suppress oncogenic transformation. Nature Cell Biol. 3, 1–7 (2001).
Cohen, O., Feinstein, E. & Kimchi, A. DAP-kinase is a Ca2+/calmodulin-dependent kinase, with cell death-inducing functions that depend on its catalytic activity. EMBO J. 16, 998–1008 (1997).
Kimchi, A. DAP genes: novel apoptotic genes isolated by a functional approach to gene cloning. Biochim. Biophys. Acta 1377, 13–33 (1998).
Cohen, O. et al. DAP-kinase participates in TNFα- and Fas-induced apoptosis and its function requires the death domain. J. Cell. Biol. 146, 141–148 (1999).
Raveh, T., Berissi, H., Eisenstein, M., Spivak, T. & Kimchi, A. A functional genetic screen identifies regions at the C-terminal tail and death-domain of death-associated protein kinase that are critical for its proapoptotic activity. Proc. Natl. Acad. Sci. USA 97, 1572–1577 (2000).
Kawai, T. et al. Death-asociated protein kinase 2 is a new calcium/calmodulin-dependent protein kinase that signals apoptosis through its catalytic activity. Oncogene 18, 3471–3480 (1999).
Inbal, B., Shani, G., Cohen, O., Kissil, J.L. & Kimchi, A. Death-associated protein kinase-related protein 1, a novel serine/threonine kinase involved in apoptosis. Mol. Cell Biol. 18, 1642–1651 (2000).
Kawai, T., Matusmoto, M., Takeda, K., Sanjo, H. & Akira, S. ZIP-kinase, a novel serine/threonine kinase which mediates apoptosis. Mol. Cell Biol. 18, 1642–1651 (1998).
Kogel, D., Plottner, O., Landsberg, G., Christian, S. & Scheidtmann, K.H. Cloning and characterization of Dlk, a novel serine/threonine kinase that is tightly associated with chromatin and phosphorylates core histones. Oncogene 17, 2645–2654 (1998).
Sanjo, H., Kawai, T. & Akira, S. DRAKs, novel serine/threonine kinases related to death-associated protein kinase that trigger apoptosis. J. Biol. Chem. 273, 29066–29071 (1998).
Shani, G. et al. Autophosphorylation restrains the apoptotic activity of DRP-1 kinase by controlling dimerization and calmodulin binding. EMBO J. 20, 1099–1113 (2001).
Velentza, A.V., Schumacher, A.M., Weiss, C., Egli, M. & Watterson, D.M. A protein kinase associated with apoptosis and tumor suppression: structure, activity and discovery of peptide substrates. J. Biol. Chem. In the press (2001).
Kobe, B. et al. Giant protein kinases: domain interactions and structural basis for autoregulation. EMBO J. 15, 6810–6821 (1996).
Mayans, O. et al. Structural basis for activation of the titin kinase domain during myofibrillogenesis. Nature 395, 863–868 (1998).
Goldberg, J. Nairn, A.C. & Kuriyan, J. Structural basis for the autoinhibition of calcium/calmodulin-dependent protein kinase I. Cell 84, 875–887 (1996).
Lowe, E.D. et al. The crystal structure of a phosphorylase kinase peptide substrate complex: kinase substrate recognition. EMBO J. 16, 6646–6658 (1997).
Zheng, J. et al. 2.2 Å refined crystal structure of the catalytic subunit of cAMP-dependent protein kinase complexed with MNATP and a peptide inhibitor. Acta Crystallogr. D 49, 362–365 (1993).
Knighton, D.R. et al. Crystal structure of the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 407–414 (1991).
Knighton, D.R. et al. Structure of a peptide inhibitor bound to the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 414–420 (1991).
Dobler, M. Ionophores and their structures (Wiley, New York; 1981).
Lukas, T.J., Mirzoeva, S. & Watterson, D.M. In Calmodulin and signal transduction (eds. Van Eldik, L. & Watterson, D.M.) 65–168 (Academic Press, San Diego; 1998).
Shoemaker, M. O. et al. Use of DNA sequence and mutant analyses and antisense oligodeoxynucleotides to examine the molecular basis of nonmuscle myosin light chain kinase autoinhibition, calmodulin recognition, and activity. J. Cell. Biol. 111, 1107–1125 (1990).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Brünger, A.T. Crystallography & NMR system (CNS), Version 0.9 (Yale University, New Haven; 1998).
Perrakis, A., Morris. R. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nature Struct. Biol. 6, 458–463 (1999).
Cambillau, C. & Roussel, A. Turbo Frodo, Version OpenGL.1 (Université Aix-Marseille II, Marseille; 1997).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Brünger, A.T. Free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–475 (1992).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Vagin, A. A., Murshudov, G. N. & Strokopytov, B. V. BLANC: the program suite for protein crystallography. J. Appl. Crystallogr. 31, 98–02 (1998).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Protein Struc. Func. Genet. 11, 281–296 (1991).
Koradi, R., Billeter, M. & Wüthrich, K. MOLMOL, a program for display and analysis of macromolecular structures. J. Mol. Graphics 14, 51–55 (1996).
Acknowledgements
This work was supported in part by a NIH grant to M.E. and in part by grants from the Alzheimer's Association, the Institute for the Study of Aging and the NIH to D.M.W. We are grateful to W.F. Anderson, T J. Lukas, G. Minasov and A. Velentza for expert assistance during the various stages of the project. Use of the Advanced Photon Source was supported by the U.S. Department of Energy, Basic Energy Sciences, Office of Science. The DuPont-Northwestern-Dow Collaborative Access Team (DND-CAT) Synchrotron Research Center at the Advanced Photon Source (Sector 5) is supported by E. I. DuPont de Nemours & Co., The Dow Chemical Company, the National Science Foundation and the State of Illinois. The Industrial Macromolecular Crystallography Association Collaborative Access Team (IMCA-CAT) at the Advanced Photon Source (Sector 17) is supported by the companies of the Industrial Macromolecular Crystallography Association through a contract with the Illinois Institute of Technology (IIT), executed through the IIT's Center for Synchrotron Radiation Research and Instrumentation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tereshko, V., Teplova, M., Brunzelle, J. et al. Crystal structures of the catalytic domain of human protein kinase associated with apoptosis and tumor suppression. Nat Struct Mol Biol 8, 899–907 (2001). https://doi.org/10.1038/nsb1001-899
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1001-899
This article is cited by
-
Death-associated protein kinase 1 as a therapeutic target for Alzheimer's disease
Translational Neurodegeneration (2024)
-
Role and mechanism of action of leucine-rich repeat kinase 1 in bone
Bone Research (2017)
-
DAPK1 Signaling Pathways in Stroke: from Mechanisms to Therapies
Molecular Neurobiology (2017)
-
The DAPK family: a structure–function analysis
Apoptosis (2014)
-
GTP binding and intramolecular regulation by the ROC domain of Death Associated Protein Kinase 1
Scientific Reports (2012)