Abstract
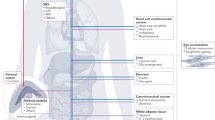
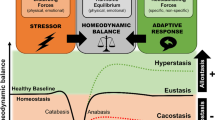
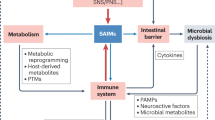
Accumulating evidence points to a major role for chronic stress of cell renewal systems in the pathogenesis of important human diseases, including cancer, atherosclerosis and diabetes. Here we discuss emerging evidence that epigenetic abnormalities may make substantial contributions to these stress-induced pathologies. Although the mechanisms remain to be fully elucidated, we suggest that chronic stress can elicit heritable changes in the chromatin landscape that 'lock' cells in abnormal states, which then lead to disease. We emphasize the need to investigate epigenetic states in disease and links to stress and to consider how the knowledge gained through these studies may foster new means of disease prevention and management.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Beckman, J. A., Creager, M. A. & Libby, P. Diabetes and atherosclerosis: epidemiology, pathophysiology, and management. JAMA 287, 2570–2581 (2002).
Hotamisligil, G. S. Endoplasmic reticulum stress and the inflammatory basis of metabolic disease. Cell 140, 900–917 (2010).
Visconti, R. & Grieco, D. New insights on oxidative stress in cancer. Curr. Opin. Drug Discov. Devel. 12, 240–245 (2009).
Zawia, N. H., Lahiri, D. K. & Cardozo-Pelaez, F. Epigenetics, oxidative stress, and Alzheimer disease. Free Radic. Biol. Med. 46, 1241–1249 (2009).
Rossi, D. J., Jamieson, C. H. & Weissman, I. L. Stems cells and the pathways to aging and cancer. Cell 132, 681–696 (2008).
Silva, H. & Conboy, I. Aging and stem cell renewal. StemBook 1–13 (2008).
Chambers, S. M. et al. Aging hematopoietic stem cells decline in function and exhibit epigenetic dysregulation. PLoS Biol. 5, e201 (2007).
Coussens, L. M. & Werb, Z. Inflammation and cancer. Nature 420, 860–867 (2002).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
Misteli, T. Beyond the sequence: cellular organization of genome function. Cell 128, 787–800 (2007).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Hawkins, R. D. et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6, 479–491 (2010).
Esteller, M. Epigenetics in evolution and disease. Lancet 372, S90–S96 (2008).
Nelson, W. G., De Marzo, A. M., DeWeese, T. L. & Isaacs, W. B. The role of inflammation in the pathogenesis of prostate cancer. J. Urol. 172, S6–S12 (2004).
Pani, G., Galeotti, T. & Chiarugi, P. Metastasis: cancer cell's escape from oxidative stress. Cancer Metastasis Rev. 29, 351–378 (2010).
Meng, X. & Riordan, N. H. Cancer is a functional repair tissue. Med. Hypotheses 66, 486–490 (2006).
Dvorak, H. F. Tumors: wounds that do not heal. Similarities between tumor stroma generation and wound healing. N. Engl. J. Med. 315, 1650–1659 (1986).
Iliopoulos, D., Hirsch, H. A. & Struhl, K. An epigenetic switch involving NF-κB, Lin28, Let-7 microRNA, and IL6 links inflammation to cell transformation. Cell 139, 693–706 (2009).
Tlsty, T. D. & Coussens, L. M. Tumor stroma and regulation of cancer development. Annu. Rev. Pathol. 1, 119–150 (2006).
Orimo, A. & Weinberg, R. A. Stromal fibroblasts in cancer: a novel tumor-promoting cell type. Cell Cycle 5, 1597–1601 (2006).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Gidekel Friedlander, S. Y. et al. Context-dependent transformation of adult pancreatic cells by oncogenic K-Ras. Cancer Cell 16, 379–389 (2009).
Baylin, S. B. & Ohm, J. E. Epigenetic gene silencing in cancer — a mechanism for early oncogenic pathway addiction? Nature Rev. Cancer 6, 107–116 (2006).
Feinberg, A. P., Ohlsson, R. & Henikoff, S. The epigenetic progenitor origin of human cancer. Nature Rev. Genet. 7, 21–33 (2006).
Lowe, S. W., Cepero, E. & Evan, G. Intrinsic tumour suppression. Nature 432, 307–315 (2004).
Barker, N. et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 457, 608–611 (2009).
Baylin, S. B. Stem cells, cancer, and epigenetics. StemBook 1–14 (2009).
Issa, J. P. CpG-island methylation in aging and cancer. Curr. Top. Microbiol. Immunol. 249, 101–118 (2000).
Issa, J. P. et al. Methylation of the oestrogen receptor CpG island links ageing and neoplasia in human colon. Nature Genet. 7, 536–540 (1994).
Gazin, C., Wajapeyee, N., Gobeil, S., Virbasius, C. M. & Green, M. R. An elaborate pathway required for Ras-mediated epigenetic silencing. Nature 449, 1073–1077 (2007).
Opavsky, R. et al. CpG island methylation in a mouse model of lymphoma is driven by the genetic configuration of tumor cells. PLoS Genet. 3, 1757–1769 (2007).
Weinstein, I. B. Cancer. Addiction to oncogenes — the Achilles heal of cancer. Science 297, 63–64 (2002).
Widschwendter, M. et al. Epigenetic stem cell signature in cancer. Nature Genet. 39, 157–158 (2007).
Schlesinger, Y. et al. Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nature Genet. 39, 232–236 (2007).
Ohm, J. E. & Baylin, S. B. Stem cell chromatin patterns: an instructive mechanism for DNA hypermethylation? Cell Cycle 6, 1040–1043 (2007).
Cedar, H. & Bergman, Y. Linking DNA methylation and histone modification: patterns and paradigms. Nature Rev. Genet. 10, 295–304 (2009).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Chi, A. S. & Bernstein, B. E. Developmental biology. Pluripotent chromatin state. Science 323, 220–221 (2009).
Vire, E. et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439, 871–874 (2006).
Tiwari, V. K. et al. PcG proteins, DNA methylation, and gene repression by chromatin looping. PLoS Biol. 6, 2911–2927 (2008).
O'Hagan, H. M., Mohammad, H. P. & Baylin, S. B. Double strand breaks can initiate gene silencing and SIRT1-dependent onset of DNA methylation in an exogenous promoter CpG island. PLoS Genet. 4, e1000155 (2008).
Ben-Porath, I. et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nature Genet. 40, 499–507 (2008).
Wierda, R. J., Geutskens, S. B., Jukema, J. W., Quax, P. H. & van den Elsen, P. J. Epigenetics in atherosclerosis and inflammation. J. Cell. Mol. Med. 14, 1225–1240 (2010).
Hang, C. T. et al. Chromatin regulation by Brg1 underlies heart muscle development and disease. Nature 466, 62–67 (2010).
Victor, V. M. et al. Oxidative stress, endothelial dysfunction and atherosclerosis. Curr. Pharm. Des 15, 2988–3002 (2009).
Hansson, G. K. Inflammation, atherosclerosis, and coronary artery disease. N. Engl. J. Med. 352, 1685–1695 (2005).
Turunen, M. P., Aavik, E. & Yla-Herttuala, S. Epigenetics and atherosclerosis. Biochim. Biophys. Acta 1790, 886–891 (2009).
Kim, J. et al. Epigenetic changes in estrogen receptor β gene in atherosclerotic cardiovascular tissues and in vitro vascular senescence. Biochim. Biophys. Acta 1772, 72–80 (2007).
Yoshida, T., Gan, Q. & Owens, G. K. Kruppel-like factor 4, Elk-1, and histone deacetylases cooperatively suppress smooth muscle cell differentiation markers in response to oxidized phospholipids. Am. J. Physiol. Cell Physiol. 295, C1175–C1182 (2008).
Owens, G. K., Kumar, M. S. & Wamhoff, B. R. Molecular regulation of vascular smooth muscle cell differentiation in development and disease. Physiol. Rev. 84, 767–801 (2004).
Ying, A. K. et al. Methylation of the estrogen receptor-α gene promoter is selectively increased in proliferating human aortic smooth muscle cells. Cardiovasc. Res. 46, 172–179 (2000).
Kim, J. et al. Epigenetic changes in estrogen receptor β gene in atherosclerotic cardiovascular tissues and in-vitro vascular senescence. Biochim. Biophys. Acta 1772, 72–80 (2007).
Villeneuve, L. M. & Natarajan, R. The role of epigenetics in the pathology of diabetic complications. Am. J. Physiol. Renal Physiol. 299, F14–F25 (2010).
Chauhan, V. & Chauhan, A. Oxidative stress in Alzheimer's disease. Pathophysiology 13, 195–208 (2006).
Chauhan, A. & Chauhan, V. Oxidative stress in autism. Pathophysiology 13, 171–181 (2006).
Morris, L. G., Veeriah, S. & Chan, T. A. Genetic determinants at the interface of cancer and neurodegenerative disease. Oncogene 29, 3453–3464 (2010).
Dulac, C. Brain function and chromatin plasticity. Nature 465, 728–735 (2010).
Abel, T. & Zukin, R. S. Epigenetic targets of HDAC inhibition in neurodegenerative and psychiatric disorders. Curr. Opin. Pharmacol. 8, 57–64 (2008).
Urdinguio, R. G., Sanchez-Mut, J. V. & Esteller, M. Epigenetic mechanisms in neurological diseases: genes, syndromes, and therapies. Lancet Neurol. 8, 1056–1072 (2009).
Wang, L. W., Berry-Kravis, E. & Hagerman, R. J. Fragile X: leading the way for targeted treatments in autism. Neurotherapeutics 7, 264–274 (2010).
Amir, R. E. et al. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nature Genet. 23, 185–188 (1999).
Yasui, D. H. et al. Integrated epigenomic analyses of neuronal MeCP2 reveal a role for long-range interaction with active genes. Proc. Natl Acad. Sci. USA 104, 19416–19421 (2007).
Chahrour, M. et al. MeCP2, a key contributor to neurological disease, activates and represses transcription. Science 320, 1224–1229 (2008).
Schanen, N. C. Epigenetics of autism spectrum disorders. Hum. Mol. Genet. 15, R138–R150 (2006).
Nguyen, A., Rauch, T. A., Pfeifer, G. P. & Hu, V. W. Global methylation profiling of lymphoblastoid cell lines reveals epigenetic contributions to autism spectrum disorders and a novel autism candidate gene, RORA, whose protein product is reduced in autistic brain. FASEB J. 24, 3036–3051 (2010).
Bonda, D. J. et al. Oxidative stress in Alzheimer disease: a possibility for prevention. Neuropharmacology 59, 290–294 (2010).
Guglielmotto, M., Giliberto, L., Tamagno, E. & Tabaton, M. Oxidative stress mediates the pathogenic effect of different Alzheimer's disease risk factors. Front. Aging Neurosci. 2, 3 (2010).
Marques, S. C., Oliveira, C. R., Outeiro, T. F. & Pereira, C. M. Alzheimer's disease: the quest to understand complexity. J. Alzheimers Dis. 21, 373–383 (2010).
Donmez, G., Wang, D., Cohen, D. E. & Guarente, L. SIRT1 suppresses β-amyloid production by activating the α-secretase gene ADAM10. Cell 142, 320–332 (2010).
Miao, F. et al. Lymphocytes from patients with type 1 diabetes display a distinct profile of chromatin histone H3 lysine 9 dimethylation: an epigenetic study in diabetes. Diabetes 57, 3189–3198 (2008).
Broske, A. M. et al. DNA methylation protects hematopoietic stem cell multipotency from myeloerythroid restriction. Nature Genet. 41, 1207–1215 (2009).
Trowbridge, J. J., Snow, J. W., Kim, J. & Orkin, S. H. DNA methyltransferase 1 is essential for and uniquely regulates hematopoietic stem and progenitor cells. Cell Stem Cell 5, 442–449 (2009).
Sen, G. L., Reuter, J. A., Webster, D. E., Zhu, L. & Khavari, P. A. DNMT1 maintains progenitor function in self-renewing somatic tissue. Nature 463, 563–567.
Jankowski, J. A. et al. Molecular evolution of the metaplasia–dysplasia–adenocarcinoma sequence in the esophagus. Am. J. Pathol. 154, 965–973 (1999).
Bian, Y. S., Osterheld, M. C., Fontolliet, C., Bosman, F. T. & Benhattar, J. p16 inactivation by methylation of the CDKN2A promoter occurs early during neoplastic progression in Barrett's esophagus. Gastroenterology 122, 1113–11121 (2002).
Eads, C. A. et al. Epigenetic patterns in the progression of esophageal adenocarcinoma. Cancer Res. 61, 3410–3418 (2001).
Belinsky, S. A. Gene-promoter hypermethylation as a biomarker in lung cancer. Nature Rev. Cancer 4, 707–717 (2004).
Ling, C. & Groop, L. Epigenetics: a molecular link between environmental factors and type 2 diabetes. Diabetes 58, 2718–2725 (2009).
Woo, M. & Patti, M. E. Diabetes risk begins in utero. Cell Metab. 8, 5–7 (2008).
Park, J. H., Stoffers, D. A., Nicholls, R. D. & Simmons, R. A. Development of type 2 diabetes following intrauterine growth retardation in rats is associated with progressive epigenetic silencing of Pdx1. J. Clin. Invest. 118, 2316–2324 (2008).
Steger, D. J. et al. Propagation of adipogenic signals through an epigenomic transition state. Genes Dev. 24, 1035–1044 (2010).
Lefterova, M. I. et al. PPARγ and C/EBP factors orchestrate adipocyte biology via adjacent binding on a genome-wide scale. Genes Dev. 22, 2941–2952 (2008).
Fujiki, K., Kano, F., Shiota, K. & Murata, M. Expression of the peroxisome proliferator activated receptor γ gene is repressed by DNA methylation in visceral adipose tissue of mouse models of diabetes. BMC Biol. 7, 38 (2009).
Thorel, F. et al. Conversion of adult pancreatic α-cells to β-cells after extreme β-cell loss. Nature 464, 1149–1154 (2010).
Jones, P. A. et al. Moving AHEAD with an international human epigenome project. Nature 454, 711–715 (2008).
Feinberg, A. P. Genome-scale approaches to the epigenetics of common human disease. Virchows Arch. 456, 13–21 (2010).
O'Neill, L. P., VerMilyea, M. D. & Turner, B. M. Epigenetic characterization of the early embryo with a chromatin immunoprecipitation protocol applicable to small cell populations. Nature Genet. 38, 835–841 (2006).
Jones, P. A. & Baylin, S. B. The epigenomics of cancer. Cell 128, 683–692 (2007).
Egger, G., Liang, G., Aparicio, A. & Jones, P. A. Epigenetics in human disease and prospects for epigenetic therapy. Nature 429, 457–463 (2004).
Guy, J., Gan, J., Selfridge, J., Cobb, S. & Bird, A. Reversal of neurological defects in a mouse model of Rett syndrome. Science 315, 1143–1147 (2007).
Qin, W. et al. Neuronal SIRT1 activation as a novel mechanism underlying the prevention of Alzheimer disease amyloid neuropathology by calorie restriction. J. Biol. Chem. 281, 21745–21754 (2006).
Guarente, L. Cell biology. Hypoxic hookup. Science 324, 1281–1282 (2009).
Peleg, S. et al. Altered histone acetylation is associated with age-dependent memory impairment in mice. Science 328, 753–756 (2010).
Acknowledgements
The authors thank members of the Baylin laboratory for helpful suggestions and reading of the manuscript and K. Bender for help with manuscript preparation. Portions of the authors' work cited have been supported by the National Cancer Institute grant CA043318, the National Institute of Environmental Health Sciences grant ES011858 and the National Institutes of Health grant CA116160.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Amyloid plaque
-
A focal extracellular collection of amyloid protein that surrounds dystrophic neurites in the hippocampus, amygdala and neocortex.
- Committed progenitor cell
-
A multipotent cell that is committed to a particular lineage and incapable of self-renewal.
- Histone post-translational modification
-
A covalent alteration of a histone tail residue that alters chromatin structure. Modifications include phosphorylation, methylation, acetylation, sumoylation and ubiquitylation.
- Histone variant
-
A histone that differs from canonical histones in terms of its structure and regulation.
- Neoplasia
-
The abnormal proliferation of cells. A neoplasm can be further characterized as benign or malignant based on its ability to invade other tissues.
- Neurofibrillary tangle
-
A filamentous bundle of tau protein that aggregates in the cytoplasm of neurons and is a histological hallmark of Alzheimer's disease.
- Nucleosome remodelling
-
An enzymatic process that alters the position of nucleosomes and can influence chromatin condensation.
- Peripheral resistance
-
The resistance of the peripheral vasculature to blood flow.
- Polycomb group
-
A family of proteins that are involved in gene silencing during development through methylation of histone H3 lysine 27.
- Pre-malignant
-
A state in which a cell is benign but poised to become malignant.
Rights and permissions
About this article
Cite this article
Johnstone, S., Baylin, S. Stress and the epigenetic landscape: a link to the pathobiology of human diseases?. Nat Rev Genet 11, 806–812 (2010). https://doi.org/10.1038/nrg2881
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg2881
This article is cited by
-
La vida en la frontera: protocol for a prospective study exploring stress and health resiliencies among Mexican-origin individuals living in a US-Mexico border community
BMC Public Health (2022)
-
Achieving equity through science and integrity: dismantling race-based medicine
Pediatric Research (2022)
-
“Racism as a public health issue” APS racism series: at the intersection of equity, science, and social justice
Pediatric Research (2020)
-
Targeting metabolic disorders by natural products
Journal of Diabetes & Metabolic Disorders (2015)
-
Stress und optimales Altern
Zeitschrift für Gerontologie und Geriatrie (2015)