Abstract
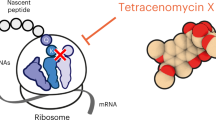
Whereas screening of the small-molecule metabolites produced by most cultivatable microorganisms often results in the rediscovery of known compounds, genome-mining programs allow researchers to harness much greater chemical diversity, and result in the discovery of new molecular scaffolds. Here we report the genome-guided identification of a new antibiotic, klebsazolicin (KLB), from Klebsiella pneumoniae that inhibits the growth of sensitive cells by targeting ribosomes. A ribosomally synthesized post-translationally modified peptide (RiPP), KLB is characterized by the presence of a unique N-terminal amidine ring that is essential for its activity. Biochemical in vitro studies indicate that KLB inhibits ribosomes by interfering with translation elongation. Structural analysis of the ribosome–KLB complex showed that the compound binds in the peptide exit tunnel overlapping with the binding sites of macrolides or streptogramin-B. KLB adopts a compact conformation and largely obstructs the tunnel. Engineered KLB fragments were observed to retain in vitro activity, and thus have the potential to serve as a starting point for the development of new bioactive compounds.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lewis, K. Platforms for antibiotic discovery. Nat. Rev. Drug Discov. 12, 371–387 (2013).
Brown, E.D. & Wright, G.D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Laxminarayan, R. et al. Antibiotic resistance—the need for global solutions. Lancet Infect. Dis. 13, 1057–1098 (2013).
Walsh, C.T. & Wencewicz, T.A. Prospects for new antibiotics: a molecule-centered perspective. J. Antibiot. (Tokyo) 67, 7–22 (2014).
Liu, Y.Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
Doroghazi, J.R. et al. A roadmap for natural product discovery based on large-scale genomics and metabolomics. Nat. Chem. Biol. 10, 963–968 (2014).
Donia, M.S. et al. A systematic analysis of biosynthetic gene clusters in the human microbiome reveals a common family of antibiotics. Cell 158, 1402–1414 (2014).
Zipperer, A. et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 535, 511–516 (2016).
Donia, M.S. & Fischbach, M.A. Small molecules from the human microbiota. Science 349, 1254766 (2015).
Arnison, P.G. et al. Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30, 108–160 (2013).
Oman, T.J. & van der Donk, W.A. Follow the leader: the use of leader peptides to guide natural product biosynthesis. Nat. Chem. Biol. 6, 9–18 (2010).
Koehnke, J. et al. The cyanobactin heterocyclase enzyme: a processive adenylase that operates with a defined order of reaction. Angew. Chem. Int. Edn. Engl. 52, 13991–13996 (2013).
Dunbar, K.L. et al. Discovery of a new ATP-binding motif involved in peptidic azoline biosynthesis. Nat. Chem. Biol. 10, 823–829 (2014).
Burkhart, B.J., Schwalen, C.J., Mann, G., Naismith, J.H. & Mitchell, D.A. YcaO-dependent posttranslational amide activation: biosynthesis, structure, and function. Chem. Rev. 117, 5389–5456 (2017).
Dunbar, K.L., Melby, J.O. & Mitchell, D.A. YcaO domains use ATP to activate amide backbones during peptide cyclodehydrations. Nat. Chem. Biol. 8, 569–575 (2012).
Lee, S.W. et al. Discovery of a widely distributed toxin biosynthetic gene cluster. Proc. Natl. Acad. Sci. USA 105, 5879–5884 (2008).
Cox, C.L., Doroghazi, J.R. & Mitchell, D.A. The genomic landscape of ribosomal peptides containing thiazole and oxazole heterocycles. BMC Genomics 16, 778 (2015).
Crone, W.J. et al. Dissecting bottromycin biosynthesis using comparative untargeted metabolomics. Angew. Chem. Int. Edn. Engl. 55, 9639–9643 (2016).
San Millán, J.L., Kolter, R. & Moreno, F. Plasmid genes required for microcin B17 production. J. Bacteriol. 163, 1016–1020 (1985).
Huo, L., Rachid, S., Stadler, M., Wenzel, S.C. & Müller, R. Synthetic biotechnology to study and engineer ribosomal bottromycin biosynthesis. Chem. Biol. 19, 1278–1287 (2012).
Salomón, R.A. & Farías, R.N. The FhuA protein is involved in microcin 25 uptake. J. Bacteriol. 175, 7741–7742 (1993).
Laviña, M., Pugsley, A.P. & Moreno, F. Identification, mapping, cloning and characterization of a gene (sbmA) required for microcin B17 action on Escherichia coli K12. J. Gen. Microbiol. 132, 1685–1693 (1986).
Patzer, S.I., Baquero, M.R., Bravo, D., Moreno, F. & Hantke, K. The colicin G, H and X determinants encode microcins M and H47, which might utilize the catecholate siderophore receptors FepA, Cir, Fiu and IroN. Microbiology 149, 2557–2570 (2003).
Runti, G. et al. Functional characterization of SbmA, a bacterial inner membrane transporter required for importing the antimicrobial peptide Bac7(1-35). J. Bacteriol. 195, 5343–5351 (2013).
Salomón, R.A. & Farías, R.N. The peptide antibiotic microcin 25 is imported through the TonB pathway and the SbmA protein. J. Bacteriol. 177, 3323–3325 (1995).
Novikova, M. et al. The Escherichia coli Yej transporter is required for the uptake of translation inhibitor microcin C. J. Bacteriol. 189, 8361–8365 (2007).
Mattiuzzo, M. et al. Role of the Escherichia coli SbmA in the antimicrobial activity of proline-rich peptides. Mol. Microbiol. 66, 151–163 (2007).
Maio, A., Brandi, L., Donadio, S. & Gualerzi, C.O. The oligopeptide permease Opp mediates illicit transport of the bacterial P-site decoding inhibitor GE81112. Antibiotics (Basel) 5, 17 (2016).
Vondenhoff, G.H. et al. Characterization of peptide chain length and constituency requirements for YejABEF-mediated uptake of microcin C analogues. J. Bacteriol. 193, 3618–3623 (2011).
Osterman, I.A. et al. Sorting out antibiotics' mechanisms of action: a double fluorescent protein reporter for high-throughput screening of ribosome and DNA biosynthesis inhibitors. Antimicrob. Agents Chemother. 60, 7481–7489 (2016).
Krizsan, A. et al. Insect-derived proline-rich antimicrobial peptides kill bacteria by inhibiting bacterial protein translation at the 70S ribosome. Angew. Chem. Int. Edn. Engl. 53, 12236–12239 (2014).
Seefeldt, A.C. et al. Structure of the mammalian antimicrobial peptide Bac7(1-16) bound within the exit tunnel of a bacterial ribosome. Nucleic Acids Res. 44, 2429–2438 (2016).
Han, M.J., Lee, S.Y. & Hong, S.H. Comparative analysis of envelope proteomes in Escherichia coli B and K-12 strains. J. Microbiol. Biotechnol. 22, 470–478 (2012).
Orelle, C. et al. Tools for characterizing bacterial protein synthesis inhibitors. Antimicrob. Agents Chemother. 57, 5994–6004 (2013).
Ban, N., Nissen, P., Hansen, J., Moore, P.B. & Steitz, T.A. The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution. Science 289, 905–920 (2000).
Polikanov, Y.S., Melnikov, S.V., Söll, D. & Steitz, T.A. Structural insights into the role of rRNA modifications in protein synthesis and ribosome assembly. Nat. Struct. Mol. Biol. 22, 342–344 (2015).
Garza-Ramos, G., Xiong, L., Zhong, P. & Mankin, A. Binding site of macrolide antibiotics on the ribosome: new resistance mutation identifies a specific interaction of ketolides with rRNA. J. Bacteriol. 183, 6898–6907 (2001).
Hartz, D., McPheeters, D.S., Traut, R. & Gold, L. Extension inhibition analysis of translation initiation complexes. Methods Enzymol. 164, 419–425 (1988).
Kannan, K., Vázquez-Laslop, N. & Mankin, A.S. Selective protein synthesis by ribosomes with a drug-obstructed exit tunnel. Cell 151, 508–520 (2012).
Kannan, K. et al. The general mode of translation inhibition by macrolide antibiotics. Proc. Natl. Acad. Sci. USA 111, 15958–15963 (2014).
Davis, A.R., Gohara, D.W. & Yap, M.N. Sequence selectivity of macrolide-induced translational attenuation. Proc. Natl. Acad. Sci. USA 111, 15379–15384 (2014).
Noeske, J. et al. Synergy of streptogramin antibiotics occurs independently of their effects on translation. Antimicrob. Agents Chemother. 58, 5269–5279 (2014).
Roy, R.N., Lomakin, I.B., Gagnon, M.G. & Steitz, T.A. The mechanism of inhibition of protein synthesis by the proline-rich peptide oncocin. Nat. Struct. Mol. Biol. 22, 466–469 (2015).
Gagnon, M.G. et al. Structures of proline-rich peptides bound to the ribosome reveal a common mechanism of protein synthesis inhibition. Nucleic Acids Res. 44, 2439–2450 (2016).
Seefeldt, A.C. et al. The proline-rich antimicrobial peptide Onc112 inhibits translation by blocking and destabilizing the initiation complex. Nat. Struct. Mol. Biol. 22, 470–475 (2015).
Bulkley, D., Innis, C.A., Blaha, G. & Steitz, T.A. Revisiting the structures of several antibiotics bound to the bacterial ribosome. Proc. Natl. Acad. Sci. USA 107, 17158–17163 (2010).
Wiegand, I., Hilpert, K. & Hancock, R.E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3, 163–175 (2008).
Douthwaite, S., Powers, T., Lee, J.Y. & Noller, H.F. Defining the structural requirements for a helix in 23S ribosomal RNA that confers erythromycin resistance. J. Mol. Biol. 209, 655–665 (1989).
Zaporojets, D., French, S. & Squires, C.L. Products transcribed from rearranged rrn genes of Escherichia coli can assemble to form functional ribosomes. J. Bacteriol. 185, 6921–6927 (2003).
Moazed, D. & Noller, H.F. Interaction of tRNA with 23S rRNA in the ribosomal A, P, and E sites. Cell 57, 585–597 (1989).
Moazed, D. & Noller, H.F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989).
Orelle, C. et al. Identifying the targets of aminoacyl-tRNA synthetase inhibitors by primer extension inhibition. Nucleic Acids Res. 41, e144 (2013).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank A. Mankin and T. Florin (University of Illinois at Chicago, Chicago, Illinois, USA) for providing pAM552 plasmids carrying various mutations, as well as for careful reading of the manuscript and valuable suggestions. We are thankful to the members of the K. Severinov, P.V.S., and Y.S.P. laboratories for discussions and critical feedback. We thank R. Roy for critical reading of the manuscript. We also thank I. Kamyshko (MilliporeSigma, Billerica, Massachusetts, USA) for material source consulting and for providing the Supelco C5 HPLC column used for tRNA purification. We thank staff at NE-CAT beamline 24ID-C for help with data collection and freezing of the crystals, especially K. Rajashankar, M. Capel, F. Murphy, I. Kourinov, A. Lynch, S. Banerjee, D. Neau, J. Schuermann, N. Sukumar, J. Withrow, K. Perry, and C. Salbego.
This work is based upon research conducted at the Northeastern Collaborative Access Team beamlines, which are funded by the National Institute of General Medical Sciences, US National Institutes of Health (P41 GM103403). The Pilatus 6M detector on the 24ID-C beamline is funded by an NIH–ORIP HEI grant (S10 RR029205). This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357.
This work was supported by Illinois State startup funds (to Y.S.P.), the US National Institutes of Health (grant R01 AI117210 to K. Severinov), Skoltech Institutional funds (to K. Severinov), the Russian Foundation for Basic Research (grants 16-04-01100 (to P.V.S.) and 15-34-20139 (to I.A.O.)), the Ministry of Education and Science of the Russian Federation (grant 14.B25.31.0004 to K. Severinov), the Russian Science Foundation (grant 15-15-10017 to M.M. (used for the expression, purification, structure determination, and in vivo activity testing of KLB); grant 14-14-00072 to P.V.S. (used for the toe-printing assays, in vitro activity tests and chemical foot-printing)), the Dynasty Foundation (fellowship to M.M.) and FASIE (grant 9186GU/2015 to D.Y.T.).
Author information
Authors and Affiliations
Contributions
M.M. and I.U. assembled constructs for the isolation of KLB and derivatives; M.M., I.U., D.G., and D.Y.T. isolated and purified KLB and its derivatives; M.M., D.G., and I.A.O. evaluated bioactivity; D.G. and D.Y.T. selected resistant mutants; A.L.K., N.F.K., and Y.S.P. designed and performed X-ray crystallography experiments; I.A.O., E.S.K., P.V.S., and A.L.K. designed biochemistry and genetic experiments; A.Y. and K. Shabalin performed and analyzed the NMR experiments; and M.M. performed and analyzed the MS experiments, with critical input from M.K., T.A., and M.S. All authors interpreted the results. M.M., Y.S.P., P.V.S., and K. Severinov wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–4 and Supplementary Figures 1–11 (PDF 5523 kb)
Supplementary Data Set 1
Raw data for the luciferase luminescence assay shown in Figure 2b. (XLSX 15 kb)
Rights and permissions
About this article
Cite this article
Metelev, M., Osterman, I., Ghilarov, D. et al. Klebsazolicin inhibits 70S ribosome by obstructing the peptide exit tunnel. Nat Chem Biol 13, 1129–1136 (2017). https://doi.org/10.1038/nchembio.2462
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2462
This article is cited by
-
Bacteriocin diversity, function, discovery and application as antimicrobials
Nature Reviews Microbiology (2024)
-
Biosynthesis and characterization of fuscimiditide, an aspartimidylated graspetide
Nature Chemistry (2022)
-
Context-specific action of macrolide antibiotics on the eukaryotic ribosome
Nature Communications (2021)
-
Isolation and structure determination of new linear azole-containing peptides spongiicolazolicins A and B from Streptomyces sp. CWH03
Applied Microbiology and Biotechnology (2021)
-
Tetracenomycin X inhibits translation by binding within the ribosomal exit tunnel
Nature Chemical Biology (2020)