Abstract
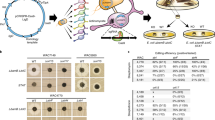
We show that selection of drug-resistant bacterial mutants allows the discovery of antibacterial compounds. Mutant strains of a soil-isolated Streptomyces species that does not produce antibacterials synthesize a previously unknown class of antibacterial, which we name piperidamycin. Overall, 6% of non-Streptomyces actinomycetes species and 43% of Streptomyces species that do not produce antibacterials are activated to produce them. The antibacterial-producing mutants all carried mutations in RNA polymerase and/or the ribosomal protein S12.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Bentley, S.D. et al. Nature 417, 141–147 (2002).
Ikeda, H. et al. Nat. Biotechnol. 21, 526–531 (2003).
Oliynyk, M. et al. Nat. Biotechnol. 25, 447–453 (2007).
Ohnishi, Y. et al. J. Bacteriol. 190, 4050–4060 (2008).
Clardy, J., Fischbach, M.A. & Walsh, C.T. Nat. Biotechnol. 24, 1541–1550 (2006).
Bode, H.B. & Muller, R. Angew. Chem. Int. Edn Engl. 44, 6828–6846 (2005).
Bode, H.B., Bethe, B., Höfs, R. & Zeeck, A. ChemBioChem 3, 619–627 (2002).
Ochi, K. et al. Adv. Appl. Microbiol. 56, 155–184 (2004).
Shima, J., Hesketh, A., Okamoto, S., Kawamoto, S. & Ochi, K. J. Bacteriol. 178, 7276–7284 (1996).
Hu, H., Zhang, Q. & Ochi, K. J. Bacteriol. 184, 3984–3991 (2002).
Hosaka, T., Xu, J. & Ochi, K. Mol. Microbiol. 61, 883–897 (2006).
Wang, G., Hosaka, T. & Ochi, K. Appl. Environ. Microbiol. 74, 2834–2840 (2008).
Tamehiro, N. et al. Appl. Environ. Microbiol. 69, 6412–6417 (2003).
Rigali, S. et al. EMBO Rep. 9, 670–675 (2008).
Tala, A. et al. J. Bacteriol. 191, 805–814 (2009).
Acknowledgements
This work was supported by a grant (to K.O.) from the Effective Promotion of Joint Research of Special Coordination Funds (Ministry of Education, Culture, Sports, Science, and Technology of the Japanese Government) and by an award (to T.H.) from the Bioscience Biotechnology and Agrochemistry Foundation. We are grateful to Haifeng Hu, Kazuhiko Kurosawa, Jun Xu, Hiroyuki Aoki and Dang T. Le for experimental contributions, and Tomoko Sato, Ikuko Maeda, Keiko Kubota, and Minako Ishihara for technical assistance.
Author information
Authors and Affiliations
Contributions
K.O. designed the project, analyzed the data and wrote the paper; T.H. performed all analyses of the strain 631689, isolated antibacterial compounds and wrote the paper; M.O.-K., S.K. and M.Y. determined chemical structure of piperidamycins and wrote the paper; H.M., K.M., Y.T. and A.F. isolated actinomycetes from soil and determined antibacterial productivity.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–5 and Supplementary Methods (PDF 3489 kb)
Rights and permissions
About this article
Cite this article
Hosaka, T., Ohnishi-Kameyama, M., Muramatsu, H. et al. Antibacterial discovery in actinomycetes strains with mutations in RNA polymerase or ribosomal protein S12. Nat Biotechnol 27, 462–464 (2009). https://doi.org/10.1038/nbt.1538
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1538
This article is cited by
-
Mining and unearthing hidden biosynthetic potential
Nature Communications (2021)
-
Eliciting the silent lucensomycin biosynthetic pathway in Streptomyces cyanogenus S136 via manipulation of the global regulatory gene adpA
Scientific Reports (2021)
-
Genetically engineered rpsL merodiploidy impacts secondary metabolism and antibiotic resistance in Streptomyces
World Journal of Microbiology and Biotechnology (2021)
-
Molecular engineering of antimicrobial peptides: microbial targets, peptide motifs and translation opportunities
Biophysical Reviews (2021)
-
Activation and enhancement of caerulomycin A biosynthesis in marine-derived Actinoalloteichus sp. AHMU CJ021 by combinatorial genome mining strategies
Microbial Cell Factories (2020)