Abstract
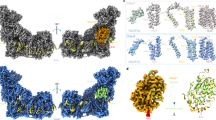
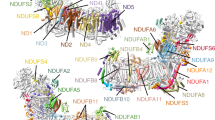
Complex I (NADH:ubiquinone oxidoreductase) is essential for oxidative phosphorylation in mammalian mitochondria. It couples electron transfer from NADH to ubiquinone with proton translocation across the energy-transducing inner membrane, providing electrons for respiration and driving ATP synthesis. Mammalian complex I contains 44 different nuclear- and mitochondrial-encoded subunits, with a combined mass of 1 MDa. The 14 conserved ‘core’ subunits have been structurally defined in the minimal, bacterial complex, but the structures and arrangement of the 30 ‘supernumerary’ subunits are unknown. Here we describe a 5 Å resolution structure of complex I from Bos taurus heart mitochondria, a close relative of the human enzyme, determined by single-particle electron cryo-microscopy. We present the structures of the mammalian core subunits that contain eight iron–sulphur clusters and 60 transmembrane helices, identify 18 supernumerary transmembrane helices, and assign and model 14 supernumerary subunits. Thus, we considerably advance knowledge of the structure of mammalian complex I and the architecture of its supernumerary ensemble around the core domains. Our structure provides insights into the roles of the supernumerary subunits in regulation, assembly and homeostasis, and a basis for understanding the effects of mutations that cause a diverse range of human diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Referenced accessions
Protein Data Bank
Data deposits
The EM map of complex I has been deposited in the Electron Microscopy Data Bank under accession number EMD-2676, and the associated model has been deposited in the Protein Data Bank under accession number 4UQ8.
References
Hirst, J. Mitochondrial complex I. Annu. Rev. Biochem. 82, 551–575 (2013)
Carroll, J., Fearnley, I. M., Shannon, R. J., Hirst, J. & Walker, J. E. Analysis of the subunit composition of complex I from bovine heart mitochondria. Mol. Cell. Proteomics 2, 117–126 (2003)
Hirst, J., Carroll, J., Fearnley, I. M., Shannon, R. J. & Walker, J. E. The nuclear encoded subunits of complex I from bovine heart mitochondria. Biochim. Biophys. Acta 1604, 135–150 (2003)
Baradaran, R., Berrisford, J. M., Minhas, G. S. & Sazanov, L. A. Crystal structure of the entire respiratory complex I. Nature 494, 443–448 (2013)
Efremov, R. G. & Sazanov, L. A. Structure of the membrane domain of respiratory complex I. Nature 476, 414–420 (2011)
Efremov, R. G., Baradaran, R. & Sazanov, L. A. The architecture of respiratory complex I. Nature 465, 441–445 (2010)
Sazanov, L. A. & Hinchliffe, P. Structure of the hydrophilic domain of respiratory complex I from Thermus thermophilus. Science 311, 1430–1436 (2006)
Hunte, C., Zickermann, V. & Brandt, U. Functional modules and structural basis of conformational coupling in mitochondrial complex I. Science 329, 448–451 (2010)
Leonard, K., Haiker, H. & Weiss, H. Three-dimensional structure of NADH:ubiquinone reductase (complex I) from Neurospora mitochondria determined by electron microscopy of membrane crystals. J. Mol. Biol. 194, 277–286 (1987)
Grigorieff, N. Three-dimensional structure of bovine NADH:ubiquinone oxidoreductase (complex I) at 22 Å in ice. J. Mol. Biol. 277, 1033–1046 (1998)
Clason, T. et al. The structure of eukaryotic and prokaryotic complex I. J. Struct. Biol. 169, 81–88 (2010)
Fassone, E. & Rahman, S. Complex I deficiency: clinical features, biochemistry and molecular genetics. J. Med. Genet. 49, 578–590 (2012)
Sharpley, M. S., Shannon, R. J., Draghi, F. & Hirst, J. Interactions between phospholipids and NADH:ubiquinone oxidoreductase (complex I) from bovine mitochondria. Biochemistry 45, 241–248 (2006)
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Bai, X.-C., Fernandez, I. S., McMullan, G. & Scheres, S. H. W. Ribosome structures to near-atomic resolution from thirty thousand cryo-EM particles. eLife 2, e00461 (2013)
Birrell, J. A. & Hirst, J. Truncation of subunit ND2 disrupts the threefold symmetry of the antiporter-like subunits in complex I from higher metazoans. FEBS Lett. 584, 4247–4252 (2010)
Kussmaul, L. & Hirst, J. The mechanism of superoxide production by NADH:ubiquinone oxidoreductase (complex I) from bovine heart mitochondria. Proc. Natl Acad. Sci. USA 103, 7607–7612 (2006)
Mimaki, M., Wang, X., McKenzie, M., Thorburn, D. R. & Ryan, M. T. Understanding mitochondrial complex I assembly in health and disease. Biochim. Biophys. Acta 1817, 851–862 (2012)
Dieteren, C. E. J. et al. Subunit-specific incorporation efficiency and kinetics in mitochondrial complex I homeostasis. J. Biol. Chem. 287, 41851–41860 (2012)
Brockmann, C. et al. The oxidised subunit B8 from human complex I adopts a thioredoxin fold. Structure 12, 1645–1654 (2004)
Keeney, P. M., Xie, J., Capaldi, R. A. & Bennett, J. P. Parkinson’s disease brain mitochondrial complex I has oxidatively damaged subunits and is functionally impaired and misassembled. J. Neurosci. 26, 5256–5264 (2006)
Leshinsky-Silver, E. et al. NDUFS4 mutations cause Leigh syndrome with predominant brainstem involvement. Mol. Genet. Metab. 97, 185–189 (2009)
Kirby, D. M. et al. NDUFS6 mutations are a novel cause of lethal neonatal mitochondrial complex I deficiency. J. Clin. Invest. 114, 837–845 (2004)
Sharpley, M. S. Studies of the Catalytic Activity of NADH:Ubiquinone Oxidoreductase (Complex I) from Bovine Mitochondria. PhD thesis, Cambridge Univ. (2005)
Morais, V. A. et al. PINK1 loss of function mutations affect mitochondrial complex I activity via NdufA10 ubiquinone uncoupling. Science 344, 203–207 (2014)
Fearnley, I. M. & Walker, J. E. Conservation of sequences of subunits of mitochondiral complex I and their relationships with other proteins. Biochim. Biophys. Acta 1140, 105–134 (1992)
Abdrakhmanova, A., Zwicker, K., Kerscher, S., Zickermann, V. & Brandt, U. Tight binding of NADPH to the 39-kDa subunit of complex I is not required for catalytic activity but stabilizes the multiprotein complex. Biochim. Biophys. Acta 1757, 1676–1682 (2006)
Babot, M. et al. ND3, ND1 and 39 kDa subunits are more exposed in the de-active form of bovine mitochondrial complex I. Biochim. Biophys. Acta 1837, 929–939 (2014)
Runswick, M. J., Fearnley, I. M., Skehel, J. M. & Walker, J. E. Presence of an acyl carrier protein in NADH:ubiquinone oxidoreductase from bovine heart mitochondria. FEBS Lett. 286, 121–124 (1991)
Cronan, J. E., Fearnley, I. M. & Walker, J. E. Mammalian mitochondria contain a soluble acyl carrier protein. FEBS Lett. 579, 4892–4896 (2005)
Dobrynin, K. et al. Characterization of two different acyl carrier proteins in complex I from Yarrowia lipolytica. Biochim. Biophys. Acta 1797, 152–159 (2010)
Angerer, H. et al. The LYR protein subunit NB4M/NDUFA6 of mitochondrial complex I anchors an acyl carrier protein and is essential for catalytic activity. Proc. Natl Acad. Sci. USA 111, 5207–5212 (2014)
Andrews, B., Carroll, J., Ding, S., Fearnley, I. M. & Walker, J. E. Assembly factors for the membrane arm of human complex I. Proc. Natl Acad. Sci. USA 110, 18934–18939 (2013)
Angerer, H. et al. A scaffold of accessory subunits links the peripheral arm and the distal proton-pumping module of mitochondrial complex I. Biochem. J. 437, 279–288 (2011)
Fearnley, I. M. et al. GRIM-19, a cell death regulatory gene product, is a subunit of bovine mitochondrial NADH:ubiquinone oxidoreductase (complex I). J. Biol. Chem. 276, 38345–38348 (2001)
Banci, L. et al. Structural characterization of CHCHD5 and CHCHD7: two atypical human twin CX9C proteins. J. Struct. Biol. 180, 190–200 (2012)
Szklarczyk, R. et al. NDUFB7 and NDUFA8 are located at the intermembrane surface of complex I. FEBS Lett. 585, 737–743 (2011)
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013)
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014)
Allegretti, M., Mills, D. J., McMullan, G., Kühlbrandt, W. & Vonck, J. Atomic model of the F420-reducing [NiFe] hydrogenase by electron cryo-microscopy using a direct electron detector. eLife 3, e01963 (2014)
Bellare, J. R., Davis, H. T., Scriven, L. E. & Talmon, Y. Controlled environment vitrification system: an improved sample preparation technique. J. Electron Microsc. Tech. 10, 87–111 (1988)
Grigorieff, N. FREALIGN: high-resolution refinement of single particle structures. J. Struct. Biol. 157, 117–125 (2007)
Smith, J. M. XIMDISP—a visualization tool to aid structure determination from electron microscope images. J. Struct. Biol. 125, 223–228 (1999)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Henderson, R. et al. Tilt-pair analysis of images from a range of different specimens in single-particle electron cryomicroscopy. J. Mol. Biol. 413, 1028–1046 (2011)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Jones, D. T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 292, 195–202 (1999)
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001)
Tusnády, G. E. & Simon, I. Principles governing amino acid composition of integral membrane proteins: applications to topology prediction. J. Mol. Biol. 283, 489–506 (1998)
Bernsel, A., Viklund, H., Hennerdal, A. & Elofsson, A. TOPCONS: consensus prediction of membrane protein topology. Nucleic Acids Res. 37, W465–W468 (2009)
Söding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005)
Eswar, N. et al. Comparative protein structure modeling using Modeller. Curr. Protoc. Bioinform. Chapter 5, Unit 5.6. (2006)
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006)
Efremov, R. G. & Sazanov, L. A. Respiratory complex I: ‘steam engine’ of the cell? Curr. Opin. Struct. Biol. 21, 532–540 (2011)
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D 60, 2256–2268 (2004)
Balsa, E. et al. NDUFA4 is a subunit of complex IV of the mammalian electron transport chain. Cell Metab. 16, 378–386 (2012)
Johansson, K. et al. Structural basis for substrate specificities of cellular deoxyribonucleoside kinases. Nature Struct. Biol. 8, 616–620 (2001).
King, J. D. et al. Predicting protein function from structure - the roles of short-chain dehydrogenase/reductase enzymes in Bordetella O-antigen biosynthesis. J. Mol. Biol. 374, 749–763 (2007)
Parris, K. D. et al. Crystal structures of substrate binding to Bacillus subtilis holo-(acyl carrier protein) synthase reveal a novel trimeric arrangement of molecules resulting in three active sites. Structure 8, 883–895 (2000)
Acknowledgements
We thank R. Henderson, S. H. W. Scheres, G. McMullan, G. Murshudov, P. Emsley and J. E. Walker for helpful advice, the FEI fellows for educating us on use of the Titan Krios, J. Grimmett and T. Darling for computational help, and S. Chen and C. Savva for EM help. This work was supported by the Medical Research Council, grant numbers U105184322 (K.R.V., in R. Henderson’s group) and U105663141 (J.H.).
Author information
Authors and Affiliations
Contributions
K.R.V. carried out EM experiments and analysis; J.Z. prepared protein; K.R.V., J.Z. and J.H. modelled and analysed data; J.H. designed the project; K.R.V., J.Z. and J.H. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Single-particle cryo-EM analysis of B. taurus complex I.
a, Typical micrograph of complex I particles imaged after freezing in vitreous ice on a holey-carbon grid. Some of the selected particles are marked with red boxes. Scale bar, 50 nm. b, Two-dimensional reference classification showing particles lying in different orientations in the ice. The size of each box is 280 pixels and the two-dimensional classification was made in RELION14.
Extended Data Figure 2 Validation of the map and resolution.
a, Tilt-pair analysis45 of complex I in Cymal-7. One-hundred complex I particles from eight image pairs, recorded with a relative tilt angle of 10°, were extracted and subjected to tilt-pair analysis with FREALIGN42. The outer radius of the plot is 40° and the orange circle centred at the expected tilt angle has a radius of 6°. b, Phase randomization to check for overfitting. Phases that are beyond 10 Å in each of the micrographs used in the final data set (frames 1–32) were randomized, and then refinement was performed as for a normal data set (FSC summed image corresponding to frames 1–32). As expected, the graph shows a drop in the Fourier shell correlation (FSC) curve at 10 Å, validating the presence of information beyond 10 Å in the images. Note that the use of gold-standard refinement procedures in RELION14 prevents any overfitting, and this test was done only as an additional control. c, An overview of the final map and the model built into it. d, FSC curves of the final map and of the model versus the map. The curve in red is the gold-standard FSC of the final map (after classification) and the resolution at FSC = 0.143 is ∼4.95 Å. The curve in cyan is the FSC between the final map and the model, and at FSC = 0.5 the resolution is 6.7 Å. Note that the present model is not complete since it is only a polyalanine model without any side chains, and loop regions in a number of subunits have not been modelled. e, The final map of mammalian complex I was analysed with ResMap49. The left-hand panel (with lower density threshold) shows that the detergent–phospholipid belt is of lower resolution, and most of the protein regions of the map show resolution distributed from 5 to 6 Å. In the right-hand panel the map is shown at a higher density threshold, so the detergent–phospholipid belt is not visualized. Some of the interior parts of the map have resolution of 4.8–5 Å.
Extended Data Figure 3 Example regions of the density map with the model fitted to the map.
a, ND2 is shown from the membrane plane, highlighting the densities for three aromatic side chains and one of the helix-breaking loops. b, Subunit ND4 viewed from the matrix. c, The density for a [4Fe–4S] cluster and surrounding protein is shown in the PSST subunit. d, A region of the 49 kDa subunit shows a well resolved α-helical stretch and aromatic side chains, and the β-strands are beginning to be resolved. e, Subunit B8 is an example of a supernumerary subunit in a peripheral region of the molecule. f, Density consistent with a bound nucleotide is observed in the 39 kDa subunit, in a similar position to in homologous structures and as expected from analysis of Y. lipolytica complex I (ref. 27). However, the present resolution of the map precludes the inclusion of this nucleotide in the final model.
Extended Data Figure 4 Global comparison of the core subunit structures of bacterial and mammalian complex I.
The core subunits from B. taurus are in blue, and from T. thermophilus (PDB accession 4HEA4) in orange. The structures have been superimposed using ND1 (the heel subunit). Top: the ND2, ND4 and ND5 domain is rotated in B. taurus relative to in T. thermophilus, increasing the curvature in the B. taurus membrane domain. The complex is viewed along the 11° rotation vector (orange) that maps the T. thermophilus ND2, ND4 and ND5 domain to the B. taurus domain, along with a small 5 Å translation to superimpose the domain centres. Correspondingly, the ND3, ND4L and ND6 domains are superimposed by a 4° rotation and a 1 Å translation. Rotation of ND2, 4 and 5 about the long axis of the domain, as noted for Y. lipolytica58, is not observed. Bottom: the NADH dehydrogenase domain containing the 51 and 24 kDa subunits is rotated by 23° and translated by 14 Å in B. taurus, relative to in T. thermophilus, causing the FeS chains to diverge as the distance from ND1 increases. A similar rotation was observed in Y. lipolytica58. The complex is viewed from behind ND1. Correspondingly, the 49 kDa, PSST and TYKY subunits are superimposed by a 6° rotation and a 2 Å translation. The structures were analysed using Superpose from the CCP4 suite59 and the 75 kDa and 30 kDa subunits were not included due to their lower structural conservation.
Extended Data Figure 5 Comparison of the individual structures of the core subunits of bacterial and mammalian complex I.
a, The structure of each subunit from T. thermophilus (wheat) (PDB accession 4HEA4) has been superimposed separately on its corresponding subunit from B. taurus (coloured as labelled) with the transverse helix plus TMH16 of ND5 also aligned separately. The complexes are viewed from behind ND1 (top), from the side (middle) and from the matrix (bottom, ND subunits only). b, Observed differences in the structures of the core subunits of B. taurus and T. thermophilus complexes I. Grey, conserved structure from B. taurus and T. thermophilus (PDB accession 4HEA4); red, structural elements present only in T. thermophilus; blue, structural elements present only in B. taurus. The C-terminal domain of the 75 kDa subunit is not resolved in B. taurus, but its structure is clearly different to in T. thermophilus.
Rights and permissions
About this article
Cite this article
Vinothkumar, K., Zhu, J. & Hirst, J. Architecture of mammalian respiratory complex I. Nature 515, 80–84 (2014). https://doi.org/10.1038/nature13686
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13686
This article is cited by
-
NDUFS4 regulates cristae remodeling in diabetic kidney disease
Nature Communications (2024)
-
The long non-coding RNA OTX2-AS1 promotes tumor growth and predicts response to BCL-2 inhibition in medulloblastoma
Journal of Neuro-Oncology (2023)
-
The coupling mechanism of mammalian mitochondrial complex I
Nature Structural & Molecular Biology (2022)
-
Down regulation of the expression of mitochondrial phosphopantetheinyl-proteins in pantothenate kinase-associated neurodegeneration: pathophysiological consequences and therapeutic perspectives
Orphanet Journal of Rare Diseases (2021)
-
Positive selection on two mitochondrial coding genes and adaptation signals in hares (genus Lepus) from China
BMC Ecology and Evolution (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.