Abstract
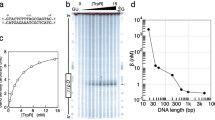
Experimental melting transitions of several natural DNAs of known nucleotide sequences have recently been obtained. The differential melting curves of these DNAs—φX174 DNA1–3, fd DNA4 and SV40 DNA5—all show distinctive sets of peaks or fine structure. Theoretical melting curves calculated from the sequences and a few a priori parameters have not accurately predicted the experimental transitions2,6,7. Although calculated fine structure resembled experimental curves in some cases, the characteristic features of a DNA's differential melting curve could not generally be produced. Azbel8,9 and Gabbarro-Arpa et al.5 have recently obtained good agreement between calculated and experimental curves using a different theoretical approach—only ground-state configurations of DNA were considered for temperatures inside the transition region. Their results suggest that the basic model of DNA melting, common to all theoretical approaches, is accurate. We have used here an exact theoretical approach to calculate melting curves of four DNA restriction fragments of 95–301 base pairs containing the lactose promoter region (Fig. 1). Theoretical curves agree very well with the experimental transitions published by Hardies et al.10 and obtained in this laboratory.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Lyubchenko, Y. L., Vologodskii, A. V. & Frank-Kamenetskii, M. D. Nature 271, 28–31 (1978).
Vizard, D. L., White, R. A. & Ansevin, A. T. Nature 275, 250–251 (1978).
Wada, A., Tachibana, H., Ueno, A., Husimi, V. & Machida, Y. Nature 269, 352–353 (1977).
Tachibana, H., Wada, A., Gotoh, O. & Takanami, M. Biochim. biophys. Acta 517, 319–328 (1978).
Gabbarro-Arpa, J., Tougard, P. & Reiss, C. Nature 280, 515–517 (1979).
Tong, B. T. & Battersby, S. J. Biopolymers 18, 1917–1936 (1979).
Ueno, S., Tachibana, H., Husimi, Y. & Wada, A. J. Biochem. 84, 917–924 (1978).
Azbel, M. Y. Proc. natn. Acad. Sci. U.S.A. 76, 105 (1979).
Azbel, M. Y. Biopolymers 19, 61–109 (1980).
Hardies, S. C., Hillen, W., Goodman, T. C. & Wells, R. D. J. biol. Chem. 254, 10128–10134 (1979).
Dickson, R. C., Abelson, J., Barnes, W. M. & Reznikoff, W. S. Science 187, 27–35 (1975).
Lyubchenko, Y. L., Frank-Kamenetskii, M. D., Vologodskii, A. V., Luzurkin, Y. & Gause, G. G. Jr Biopolymers 15, 1019–1036 (1976).
Poland, D. & Scheraga, H. A. Theory of Helix–Coil Transitions in Biopolymers (Academic, New York, 1970).
Wartell, R. M. & Montroll, E. W. Adv. chem. Phys. 22, 129–203 (1972).
Poland, D. Biopolymers 13, 1859–1871 (1974).
Applequist, J. & Damle, V. J. chem. Phys. 39, 10 (1963).
Landau, L. D. & Lifshitz, E. M. Statistical Physics (Addison-Wesley, Reading, Massachusetts, 1958).
Botchan, P. J. molec. Biol. 105, 161 (1976).
Jones, B. B., Chan, H., Rothstein, S., Wells, R. D. & Reznikoff, W. S. Proc. natn. Acad. Sci. U.S.A. 74, 4914 (1977).
Vollenweider, H. J., Fiandt, M. & Szybalski, W. Science 205, 508 (1979).
Hardies, S. C. & Wells, R. D. Gene 7, 1–14 (1979).
Wartell, R. M. & Reznikoff, W. S. Gene (in the press).
Wartell, R. M. Nucleic Acids Res. 4, 2719 (1977).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Benight, A., Wartell, R. & Howell, D. Theory agrees with experimental thermal denaturation of short DNA restriction fragments. Nature 289, 203–205 (1981). https://doi.org/10.1038/289203a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/289203a0
This article is cited by
-
Statistical thermodynamic analysis and designof DNA-based computers
Natural Computing (2004)
-
Physical modeling of biomolecular computers: Models, limitations, and experimental validation
Natural Computing (2004)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.