Key Points
-
Fanconi anaemia (FA) is a rare, autosomal, recessive disease that causes life-threatening bone marrow failure, diverse somatic abnormalities and cancer susceptibility. It is characterized by chromosomal instability and hypersensitivity to crosslinking agents, and is genetically heterogeneous.
-
The DNA-damage-response pathway that is altered in FA remains to be elucidated, but the cloning and recent characterization of several FA genes have provided clues as to their function.
-
FA patients are assigned to one of eight FA complementation groups. For six of these, disease genes have been found: FANCA, FANCC, FANCD2, FANCE, FANCF and FANCG.
-
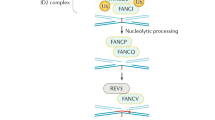
The predicted proteins of these genes lack homologues in non-vertebrates, except for FANCD2. All of them, except for FANCD2, are essential for the assembly of a multiprotein nuclear complex, which is required for FANCD2 to be monoubiquitylated to its active form.
-
Monoubiquitylated FANCD2 co-localizes with BRCA1 to subnuclear foci, the number of which increases upon DNA damage. There is also evidence that the FA pathway has a cytoplasmic component(s).
-
A proportion of FA patients show phenotypic reversion to wild type in lymphocytes. This is due to a correction of the gene defect in haematopoietic stem cells by mitotic recombination, fuelling hopes that gene therapy could be used to treat the life-threatening bone marrow failure that develops in FA patients.
Abstract
The past few years have witnessed a considerable expansion in our understanding of the pathways that maintain chromosome stability in dividing cells through the identification of genes that are mutated in certain human chromosome instability disorders. Cells that are derived from patients with Fanconi anaemia (FA) show spontaneous chromosomal instability and mutagen hypersensitivity, but FA poses a unique challenge as the nature of the DNA-damage-response pathway thought to be affected by the disease has long been a mystery. However, the recent cloning of most of the FA-associated genes, and the characterization of their protein products, has provided tantalizing clues as to the molecular basis of this disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fanconi, G. Familiäre infantile perniziosartige Anämie (perniziöses Blutbild und Konstitution). Jahrb. Kinderh. 117, 257–280 (1927).
Auerbach, A. D., Buchwald, M. & Joenje, H. in The Metabolic and Molecular Bases of Inherited Disease (ed. Scriver, C. R. et al.) 753–768 (McGraw–Hill, New York, 2001).
Verlander, P. C. et al. Carrier frequency of the IVS4 + 4 A→T mutation of the Fanconi anemia gene FAC in the Ashkenazi Jewish population. Blood 86, 4034–4038 (1995).
Auerbach, A. D. & Allen, R. G. Leukemia and preleukemia in Fanconi anemia patients: a review of the literature and report of the International Fanconi Anemia Registry. Cancer Genet. Cytogenet. 51, 1–12 (1991).
Alter, B. P. Fanconi's anemia and malignancies. Am. J. Hematol. 53, 99–110 (1996).
Schroeder, T. M., Anschütz, F. & Knopp, A. Spontane Chromosomenaberrationen bei familiärer Panmyelopathie. Humangenetik 1, 194–196 (1964).
Sasaki, M. S. & Tonomura, A. A high susceptibility of Fanconi's anemia to chromosome breakage by DNA cross-linking agents. Cancer Res. 33, 1829–1836 (1973).A thorough account of the hypersensitivity of Fanconi anaemia lymphocytes to the chromosome-breaking effect of crosslinking agents.
Auerbach, A. D. & Wolman, S. R. Susceptibility of Fanconi's anaemia fibroblasts to chromosome damage by carcinogens. Nature 261, 494–496 (1976).
Ishida, R. & Buchwald, M. Susceptibility of Fanconi's anemia lymphoblasts to DNA cross-linking and alkylating agents. Cancer Res. 42, 4000–4006 (1982).
Duckworth-Rysiecki, G. & Taylor, A. M. Effects of ionizing radiation on cells from Fanconi's anemia patients. Cancer Res. 45, 416–420 (1985).
Burnet, N. G., Wurm, R., Tait, D. M. & Peacock, J. H. Cellular sensitivity and low dose-rate recovery in Fanconi anaemia fibroblasts. Br. J. Radiol. 67, 579–583 (1994).
Strathdee, C. A., Duncan, A. M. V. & Buchwald, M. Evidence for at least four Fanconi anaemia genes including FACC on chromosome 9. Nature Genet. 1, 196–198 (1992).
Joenje, H. et al. Evidence for at least eight Fanconi anemia genes. Am. J. Hum. Genet. 61, 940–944 (1997).
Joenje, H. et al. Complementation analysis in Fanconi anemia: assignment of the reference FA-H patient to group A. Am. J. Hum. Genet. 67, 759–762 (2000).References 12–14 describe complementation group assignments in Fanconi anaemia. Reference 14 describes a caveat in the procedure due to the occasional reversion of the phenotype in fusion hybrids.
Strathdee, C. A., Gavish, H., Shannon, W. & Buchwald, M. Cloning of cDNAs for Fanconi's anaemia by functional complementation. Nature 356, 763–767 (1992).Identification of the first Fanconi anaemia gene ( FANCC ) by complementation cloning.
Pronk, J. C. et al. Localisation of the Fanconi anaemia complementation group A gene to chromosome 16q24.3. Nature Genet. 11, 338–340 (1995).
Lo Ten Foe, J. R. et al. Expression cloning of a cDNA for the major Fanconi anaemia gene, FAA. Nature Genet. 14, 320–323 (1996).
De Winter, J. P. et al. The Fanconi anaemia group G gene FANCG is identical with XRCC9. Nature Genet. 20, 281–283 (1998).
De Winter, J. P. et al. The Fanconi anaemia gene FANCF encodes a novel protein with homology to ROM. Nature Genet. 24, 15–16 (2000).
De Winter, J. P. et al. Isolation of a cDNA representing the Fanconi anemia complementation group E gene. Am. J. Hum. Genet. 67, 1306–1308 (2000).
Fanconi Anemia/Breast Cancer Consortium. Positional cloning of the Fanconi anaemia group A gene. Nature Genet. 14, 324–328 (1996).Papers 17–21 describe the identification of the genes that are defective in Fanconi anaemia complementation groups A, E, F and G.
Whitney, M. A. et al. Microcell-mediated chromosome transfer maps the Fanconi anemia group D gene to chromosome 3p. Nature Genet. 11, 341–343 (1995).
Hejna, J. A. et al. Localization of the Fanconi anemia complementation group D gene to a 200-kb region on chromosome 3p25.3. Am. J. Hum. Genet. 66, 1540–1551 (2000).
Timmers, C. et al. Positional cloning of a novel Fanconi anemia gene, FANCD2. Mol. Cell 7, 241–248 (2001).This study describes the positional cloning of FANCD2 , the first Fanconi anaemia gene for which homologues in non-vertebrate species have been identified.
Wevrick, R., Clarke, C. A. & Buchwald, M. Cloning and analysis of the murine Fanconi anemia group C cDNA. Hum. Mol. Genet. 2, 655–662 (1993).
Krasnoshtein, F. & Buchwald, M. Developmental expression of the Fac gene correlates with congenital defects in Fanconi anemia patients. Hum. Mol. Genet. 5, 85–93 (1996).
Liu, J. M. et al. Retroviral mediated gene transfer of the Fanconi anemia complementation group C gene to hematopoietic progenitors of group C patients. Hum. Gene Ther. 20, 1715–1730 (1997).
Abu-Issa, R., Eichele, G. & Youssoufian, H. Expression of the Fanconi anemia group A gene (Fanca) during mouse embryogenesis. Blood 94, 818–824 (1999).
Van de Vrugt, H. J. et al. Cloning and characterization of murine Fanconi anemia group A gene: Fanca protein is expressed in lymphoid tissues, testis, and ovary. Mamm. Genome 11, 326–331 (2000).
De Winter, J. P. et al. The Fanconi anemia protein FANCF forms a nuclear complex with FANCA, FANCC and FANCG. Hum. Mol. Genet. 9, 2665–2674 (2000).This work showed the nuclear localization of FANCF and provided evidence for a sequential interaction model for the Fanconi anaemia proteins that are defective in complementation groups A, C, E, F and G.
Xie, Y. et al. Aberrant Fanconi anemia protein profiles in acute myeloid leukemia cells. Br. J. Haematol. 111, 1057–1064 (2000).
Heinrich, M. C. et al. Posttranscriptional cell cycle-dependent regulation of human FANCC expression. Blood 95, 3970–3977 (2000).
Hoatlin, M. E. et al. A novel BTB/POZ transcriptional repressor protein interacts with the Fanconi anemia group C protein and PLZF. Blood 94, 3737–3747 (1999).A two-hybrid study connected FANCC to the chromatin-remodelling system.
Wijker, M. et al. Heterogeneous spectrum of mutations in the Fanconi anemia group A gene. Eur. J. Hum. Genet. 7, 52–59 (1999).
Morgan, N. V., Tipping, A. J., Joenje, H. & Mathew, C. G. High frequency of large intragenic deletions in the Fanconi anemia group A gene. Am. J. Hum. Genet. 65, 1330–1341 (1999).
Levran, O., Doggett, N. A. & Auerbach, A. D. Identification of Alu-mediated deletions in the Fanconi anemia gene FAA. Hum. Mutat. 12, 145–152 (1998).
Levran, O. et al. Sequence variation in the Fanconi anemia gene FAA. Proc. Natl Acad. Sci. USA 94, 13051–13056 (1997).
Kupfer, G. et al. A patient-derived mutant form of the Fanconi anemia protein, FANCA, is defective in nuclear accumulation. Exp. Hematol. 27, 587–593 (1999).
Hoatlin, M. E. et al. The Fanconi anemia group C gene product is located in both the nucleus and cytoplasm of human cells. Blood 91, 1418–1425 (1998).
Savoia, A., Garcia-Higuera, I. & D'Andrea, A. D. Nuclear localization of the Fanconi anemia protein FANCC is required for functional activity. Blood 93, 4025–4026 (1999).
Gavish, H., Dos Santos, C. C. & Buchwald, M. Leu554-Pro substitution completely abolishes complementing activity of the Fanconi anemia (FACC) protein. Hum. Mol. Genet. 2, 123–126 (1993).
Faivre, L. et al. Association of complementation group and mutation type with clinical outcome in Fanconi anemia. Blood 96, 4064–4070 (2000).
Verlander, P. C. et al. Mutation analysis of the Fanconi anemia gene FACC. Am. J. Hum. Genet. 54, 595–601 (1994).
Gillio, A. P., Verlander, P. C., Batish, S. D., Giampietro, P. F. & Auerbach, A. D. Phenotypic consequences of mutations in the Fanconi anemia FANCC gene: an International Fanconi Anemia Registry study. Blood 90, 105–110 (1997).
Yamashita, T. et al. The clinical variability of Fanconi anemia (type C) results from expression of an amino terminal truncated Fanconi anemia complementation group C polypeptide with partial activity. Blood 87, 4424–4432 (1996).
Chen, M. et al. Inactivation of Fac in mice produces inducible chromosomal instability and reduced fertility reminiscent of Fanconi anaemia. Nature Genet. 12, 448–451 (1996).
Whitney, M. A. et al. Germ cell defects and hematopoietic hypersensitivity to gamma-interferon in mice with a targeted disruption of the Fanconi anemia C gene. Blood 88, 49–58 (1996).
Cheng, N. C. et al. Mice with a targeted disruption of the Fanconi anemia homolog Fanca. Hum. Mol. Genet. 9, 1805–1811 (2000).
Nadler, J. J. & Braun, R. E. Fanconi anemia complementation group C is required for proliferation of murine primordial germ cells. Genesis 27, 117–123 (2000).
Haneline, L. S. et al. Loss of FancC function results in decreased hematopoietic stem cell repopulating ability. Blood 94, 1–8 (1999).
Luo, G. et al. Cancer predisposition caused by elevated mitotic recombination in Bloom mice. Nature Genet. 26, 424–429 (2000).
Carreau, M. et al. Bone marrow failure in the Fanconi anemia group C mouse model after DNA damage. Blood 91, 2737–2744 (1998).
Gush, K. A., Fu, K. L., Grompe, M. & Walsh, C. E. Phenotypic correction of Fanconi anemia group C knockout mice. Blood 95, 700–704 (2000).
Noll, M., Bateman, R. L., D' Andrea, A. D. & Grompe, M. Preclinical protocol for in vivo selection of hematopoietic stem cells corrected by gene therapy in Fanconi anemia group C. Mol. Ther. 3, 14–23 (2001).
Kwee, M. L. et al. Unusual response to bifunctional alkylating agents in a case of Fanconi anaemia. Hum. Genet. 64, 384–387 (1983).
Auerbach, A. D., Rogatko, A. & Schroeder-Kurth, T. M. International Fanconi Anemia Registry: relation of clinical symptoms to diepoxybutane sensitivity. Blood 73, 391–396 (1989).
Lo Ten Foe, J. R. et al. Somatic mosaicism in Fanconi anemia: molecular basis and clinical significance. Eur. J. Hum. Genet. 5, 137–148 (1997).
Pulsipher, M. et al. Subtyping analysis of Fanconi anemia by immunoblotting and retroviral gene transfer. Mol. Med. 4, 468–479 (1998).
Gregory, J. J. et al. Somatic mosaicism in Fanconi anemia: evidence of genotypic reversion in lymphohematopoietic stem cells. Proc. Natl Acad. Sci. USA 98, 2532–2537 (2001).
Waisfisz, Q. et al. Spontaneous functional correction of homozygous Fanconi anaemia alleles reveals novel mechanistic basis for reverse mosaicism. Nature Genet. 22, 379–383 (1999).
Fu, K. L. et al. Functional correction of Fanconi anemia group A hematopoietic cells by retroviral gene transfer. Blood 90, 3296–3303 (1997).
Kupfer, G. M., Naf, D., Suliman, A., Pulsipher, M. & D'Andrea, A. D. The Fanconi anaemia proteins, FAA and FAC, interact to form a nuclear complex. Nature Genet. 17, 487–490 (1997).
Waisfisz, Q. et al. A physical complex of the Fanconi anemia proteins FANCG/XRCC9 and FANCA. Proc. Natl Acad. Sci. USA 96, 10320–10325 (1999).
Garcia-Higuera, I., Kuang, Y., Naf, D., Wasik, J. & D'Andrea, A. D. Fanconi anemia proteins FANCA, FANCC, and FANCG/XRCC9 interact in a functional nuclear complex. Mol. Cell. Biol. 19, 4866–4873 (1999).
Kruyt, F. A., Abou-Zahr, F., Mok, H. & Youssoufian, H. Resistance to mitomycin C requires direct interaction between the Fanconi anemia proteins FANCA and FANCG in the nucleus through an arginine-rich domain. J. Biol. Chem. 274, 34212–34218 (1999).
Medhurst, A. L., Huber, P. A., Waisfisz, Q., De Winter, J. P. & Mathew, C. G. Direct interactions of the five known Fanconi anaemia proteins suggest a common functional pathway. Hum. Mol. Genet. 10, 423–429 (2001).
Youssoufian, H. Localization of Fanconi anemia C protein to the cytoplasm of mammalian cells. Proc. Natl Acad. Sci. USA 91, 7975–7979 (1994).
Kruyt, F. A. E. et al. Cytoplasmic localization of a functionally active Fanconi anemia group A — green fluorescent protein chimera in human 293 cells. Blood 90, 3288–3295 (1997).
Lightfoot, J., Alon, N., Bosnoyan-Collins, L. & Buchwald, M. Characterization of regions functional in the nuclear localization of the Fanconi anemia group A protein. Hum. Mol. Genet. 6, 1007–1015 (1999).
Yamashita, T. et al. The Fanconi anemia pathway requires FAA phosphorylation and FAA/FAC nuclear accumulation. Proc. Natl Acad. Sci. USA 95, 13085–13090 (1998).This study showed that FANCA needs to be phosphorylated to function; this happens in all complementation groups except FA-B.
Garcia-Higuera, I. et al. Interaction of the Fanconi anemia proteins and BRCA1 in a common pathway. Mol. Cell 7, 249–262 (2001).This important study characterized FANCD2, defined its monoubiquitylation as an activating mechanism and implicated BRCA1 as an important element of the Fanconi anaemia pathway.
Hicke, L. Protein regulation by monoubiquitin. Nature Rev. Mol. Cell Biol. 2, 195–201 (2001).
Scully, R. et al. Dynamic changes of BRCA1 subnuclear location and phosphorylation state are initiated by DNA damage. Cell 90, 425–435 (1997).
Paul, T. P. et al. A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr. Biol. 10, 886–895 (2000).
Lorick, K. L. et al. RING fingers mediate ubiquitin-conjugating enzyme (E2)-dependent ubiquitination. Proc. Natl Acad. Sci. USA 96, 11364–11369 (1999).
Joazeiro, C. A. P. & Weissman, A. M. RING finger proteins: mediators of ubiquitin ligase activity. Cell 102, 549–552 (2000).
Friedberg, E. C., Walker, G. C. & Siede, W. DNA Repair and Mutagenesis (ASM, Washington, DC, 1995).
Pierce, A. J., Johnson, R. D., Thompson, L. H. & Jasin, M. XRCC3 promotes homology-directed repair of DNA damage in mammalian cells. Genes Dev. 13, 2633–2638 (1999).
Johnson, R. D., Liu, N. & Jasin, M. Mammalian XRCC2 promotes the repair of DNA double-strand breaks by homologous recombination. Nature 401, 397–399 (1999).
Busch, D. B. et al. A CHO mutant, UV40, that is sensitive to diverse mutagens and represents a new complementation group of mitomycin C sensitivity. Mutat. Res. 363, 209–221 (1996).
Thyagarajan, B. & Campbell, C. Elevated homologous recombination activity in Fanconi anemia fibroblasts. J. Biol. Chem. 272, 23328–23333 (1997).
Smith, J. et al. Abnormal rearrangements associated with V(D)J recombination in Fanconi anemia. J. Mol. Biol. 281, 815–825 (1998).
Escarceller, M. et al. The Fanconi anemia C gene product plays a role in the fidelity of blunt DNA end-joining. J. Mol. Biol. 279, 375–385 (1998).
Papadopoulo, D., Guillouf, C., Mohrenweiser, H. & Moustacchi, E. Hypomutability in Fanconi anemia cells is associated with increased deletion frequency at the HPRT locus. Proc. Natl Acad. Sci. USA 87, 8383–8387 (1990).
Lundberg, R., Mavinakere, M. & Campbell, C. Deficient DNA end joining activity in extracts from Fanconi anemia fibroblasts. J. Biol. Chem. 279, 9543–9549 (2001).
Myung, K., Datta, A. & Kolodner, R. D. Suppression of spontaneous chromosomal rearrangements by S phase checkpoint functions in Saccharomyces cerevisiae. Cell 104, 397–408 (2001).
Salla-Trepat, M. et al. Arrest of S phase progression is impaired in Fanconi anemia cells. Exp. Cell Res. 260, 208–215 (2000).
Centurion, S. A., Kou, H. R. & Lambert, W. C. Damage resistant DNA synthesis in Fanconi anemia cells treated with a DNA crosslinking agent. Exp. Cell Res. 260, 216–221 (2000).
Scully, R. & Livingston, D. M. In search of the tumour-suppressor functions of BRCA1 and BRCA2. Nature 408, 429–432 (2000).
Wang, Y. et al. BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures. Genes Dev. 14, 927–939 (2000).
Bochar, D. A. et al. BRCA1 is associated with a human SWI/SNF-related complex: linking chromatin remodelling to breast cancer. Cell 102, 257–265 (2000).
Ikura, T. et al. Involvement of the TIP60 histone acetylase complex in DNA repair and apoptosis. Cell 102, 463–467 (2000).
Gowen, L. C., Avrutskaya, A. V., Latour, A. M., Koller, B. H. & Leadon, S. A. BRCA1 required for transcription-coupled repair of oxidative DNA damage. Science 281, 1009–1012 (1998).
Moynahan, M. E., Chiu, J. W., Koller, B. H. & Jasin, M. Brca1 controls homology-directed DNA repair. Mol. Cell 4, 511–518 (1999).
Xu, X. et al. Centrosome amplification and a defective G2-M cell cycle checkpoint induce genetic instability in BRCA1 exon 11 isoform-deficient cells. Mol. Cell 3, 389–395 (1999).
Hakem, R. et al. The tumor suppressor gene Brca1 is required for embryonic cellular proliferation in the mouse. Cell 85, 1009–1023 (1996).
Xu, X. et al. Conditional mutation of Brca1 in mammary epithelial cells results in blunted ductal morphogenesis and tumour formation. Nature Genet. 22, 37–43 (1999).
Joenje, H., Arwert, F., Eriksson, A. W., de Koning, H. & Oostra, A. B. Oxygen-dependence of chromosomal aberrations in Fanconi's anaemia. Nature 290, 143–144 (1981).
Kubbies, M., Schindler, D., Hoehn, H., Schinzel, A. & Rabinovitch, P. S. Endogenous blockage and delay of the chromosome cycle despite normal recruitment and growth phase explain poor proliferation and frequent endomitosis in Fanconi anemia cells. Am. J. Hum. Genet. 37, 1022–1030 (1985).
Schindler, D. & Hoehn, H. Fanconi anemia mutation causes cellular susceptibility to ambient oxygen. Am. J. Hum. Genet. 43, 429–435 (1988).
Joenje, H. Genetic toxicology of oxygen. Mutat. Res. 219, 193–208 (1989).
Gille, J. J. P., Van Berkel, C. & Joenje, H. Mutagenicity of metabolic oxygen radicals in mammalian cell cultures. Carcinogenesis 15, 2695–2699 (1994).
Kruyt, F. A. et al. Abnormal microsomal detoxification implicated in Fanconi anemia group C by interaction of the FAC protein with NADPH cytochrome P450 reductase. Blood 92, 3050–3056 (1998).
Korkina, L. G. et al. Release of active oxygen radicals by leukocytes of Fanconi anemia patients. J. Leukoc. Biol. 52, 357–362 (1992).
Rosselli, F., Sanceau, J., Gluckman, E., Wietzerbin, J. & Moustacchi, E. Abnormal lymphokine production: a novel feature of the genetic disease Fanconi anemia. II. In vitro and in vivo spontaneous production of tumor necrosis factor α. Blood 84, 1216–1225 (1994).
Cumming, R. C., Liu, J. M., Youssoufian, H. & Buchwald, M. Suppression of apoptosis in hematopoietic factor-dependent progenitor cell lines by expression of the FAC gene. Blood 88, 4558–4567 (1996).
Clarke, A. A., Philpott, N. J., Gordon-Smith, E. C. & Rutherford, T. R. The sensitivity of Fanconi anaemia group C cells to apoptosis induced by mitomycin C is due to oxygen radical generation, not DNA crosslinking. Br. J. Haematol. 96, 240–247 (1997).
Rathbun, R. K. et al. Inactivation of the Fanconi anemia group C gene augments interferon-γ-induced apoptotic responses in hematopoietic cells. Blood 90, 974–985 (1997).
Haneline, L. S. et al. Multiple inhibitory cytokines induce deregulated progenitor growth and apoptosis in hematopoietic cells from Fac-/- mice. Blood 91, 4092–4098 (1998).
Wang, J. et al. Overexpression of the Fanconi anemia group C gene (FAC) protects hematopoietic progenitors from death induced by Fas-mediated apoptosis. Cancer Res. 58, 3538–3541 (1998).
Pang, Q. et al. The Fanconi anemia protein FANCC binds to and facilitates the activation of STAT1 by γ interferon and hematopoietic growth factors. Mol. Cell. Biol. 20, 4724–4735 (2000).
Pang, Q. et al. Role of double-stranded RNA-dependent protein kinase in mediating hypersensitivity of Fanconi anemia complementation group C cells to interferon γ, tumor necrosis factor α, and double-stranded RNA. Blood 97, 1644–1652 (2001).
Cassinat, B. et al. Constitutive elevation of serum α-fetoprotein in Fanconi anemia. Blood 96, 859–863 (2000).
Tipping, A. et al. Multiple founder mutations for Fanconi anemia in the Afrikaner population of South Africa. Proc. Natl Acad. Sci. USA 98, 5734–5739 (2001).
Whitney, M. A. et al. A common mutation in the FACC gene causes Fanconi anaemia in Ashkenazi-Jewish individuals. Nature Genet. 4, 202–205 (1993).
Demuth, I. et al. Spectrum of mutations in the Fanconi anaemia group G gene, FANCG/XRCC9. Eur. J. Hum. Genet. 8, 861–868 (2000).
Gatti, R. in The Metabolic and Molecular Bases of Inherited Disease (ed. Scriver, C. R. et al.) 705–732 (McGraw–Hill, New York, 2001).
German, J. & Ellis, N. A. in The Metabolic and Molecular Bases of Inherited Disease (ed. Scriver, C. R. et al.) 733–751 (McGraw–Hill, New York, 2001).
Schellenberg, G. D., Miki, T., Yu, C.-E. & Nakura, J. in The Metabolic and Molecular Bases of Inherited Disease (ed. Scriver, C. R. et al.) 785–797 (McGraw–Hill, New York, 2001).
Bootsma, D., Kraemer, K. H., Cleaver, J. E. & Hoeijmakers, J. H. J. in The Metabolic and Molecular Bases of Inherited Disease (ed. Scriver, C. R. et al.) 677–703 (McGraw–Hill, New York, 2001).
Boland, C. R. in The Metabolic and Molecular Bases of Inherited Disease (ed. Scriver, C. R. et al.) 769–783 (McGraw–Hill, New York, 2001).
Couch, F. J. & Weber, B. L. in The Metabolic and Molecular Bases of Inherited Disease (ed. Scriver, C. R. et al.) 999–1031 (McGraw–Hill, New York, 2001).
Kinzler, K. W. & Vogelstein, B. in The Metabolic and Molecular Bases of Inherited Disease (ed. Scriver, C. R. et al.) 675–676 (McGraw–Hill, New York, 2001).
Patel, K. J. et al. Involvement of Brca2 in DNA repair. Mol. Cell 1, 347–357 (1998).
Moyanan, M. E., Pierce, A. J. & Jasin, M. BRCA2 is required for homology-directed repair of chromosomal breaks. Mol. Cell 7, 263–272 (2001).
Liu, N. et al. The human XRCC9 gene corrects chromosomal instability and mutagen sensitivities in CHO UV40 cells. Proc. Natl Acad. Sci. USA 94, 9232–9237 (1997).
Qiao, F., Moss, A. & Kupfer, G. M. Fanconi anemia proteins localize to chromatin and the nuclear matrix in a DNA damage and cell cycle-regulated manner. J. Biol. Chem. (in the press).
Hadjur, S. et al. Defective hematopoiesis and hepatic steatosis in mice with combined deficiencies of the genes encoding Fancc and Cu/Zn superoxide dismutase. Blood (in the press).
Acknowledgements
We thank our lab members for stimulating discussions and valuable comments on the manuscript. Work in our laboratories is supported by grants from the Dutch Cancer Society, the Netherlands Organization for Scientific Research, the FA Research Fund, Inc., Eugene, Oregon, the European FA patient support organizations, and the Medical Research Council. We apologize to colleagues whose work could not be cited because of space limitations.
Author information
Authors and Affiliations
Supplementary information
Related links
Related links
DATABASE LINKS
hereditary non-polyposis colorectal cancer
NADPH cytochrome P450 reductase
FURTHER INFORMATION
Glossary
- APLASTIC ANAEMIA
-
Bone marrow failure. This results in a peripheral deficiency of all bone-marrow-derived haematopoietic lineages, such as red blood cells, platelets and leukocytes. There are many causes of this potentially fatal clinical syndrome.
- ACUTE MYELOID LEUKAEMIA
-
(AML) Also known as acute non-lymphocytic leukaemia (ANLL), this is a heterogeneous group of leukaemias in which the lymphoblast cells that give rise to granulocytes do not mature and become too numerous. These immature blast cells are then found in the blood and the bone marrow.
- SQUAMOUS CELL CARCINOMAS
-
Cancers that arise from squamous epithelial cells or 'keratinocytes', which cover the skin and oral cavity.
- CROSSLINKING AGENTS
-
These agents form a specific type of lesion in double-stranded DNA — chemical crosslinks between the two DNA strands (interstrand crosslinks) and between adjacent bases on the same strand (intrastrand crosslinks). These modifications obstruct DNA replication and, because they can modify both strands, specific repair pathways are required to maintain genomic integrity in cells exposed to them.
- CARETAKER GENES
-
Genes that encode the proteins that function to protect the genome against alterations.
- NUCLEOTIDE EXCISION REPAIR
-
A DNA-repair pathway that removes DNA damage (such as ultraviolet-light-induced thymidine dimers) and bulky DNA adducts by excising the region of DNA that contains the damaged base(s). The single-stranded gap is filled in by using the intact strand as a template.
- MICROCELL
-
An experimentally produced cell-like structure, which contains a micronucleus in which one or a few chromosomes are present. They can be used as a vector to transfer genetic material to a normal cell by cell fusion.
- SERIAL TRANSPLANTATION
-
When a donor tissue is used for another transplant after engraftment in a recipient.
- MITOTIC RECOMBINATION
-
A crossover between two homologous double-stranded DNA molecules that leads to a physical exchange of DNA and genetic information. This recombination occurs frequently during meiosis, but is relatively rare during mitosis. As a consequence of mitotic recombination, cells can undergo a 'loss of heterozygosity' or gene conversion.
- GENE CONVERSION
-
A specific type of recombination, which results in non-reciprocal genetic exchange, in which the sequence of one DNA strand is used to alter the sequence of the other.
- RAD51
-
The human homologue of bacterial RecA. RAD51 is required for homologous recombination, during which it promotes strand invasion. RAD51 forms nucleoprotein filaments around single-stranded DNA.
- RAD50
-
RAD50 forms a complex with MRE11 and NBS1 and is essential for double-stranded DNA-break repair by homologous recombination and non-homologous end-joining.
- RING-FINGER MOTIF
-
A structure seen in many DNA-binding proteins; it is composed of a polypeptide loop held in a hairpin bend bound to a zinc atom.
- UBIQUITIN LIGASE
-
An enzyme that couples the small (∼7 kDa) protein ubiquitin to lysine residues on a target protein, thus targeting it for degradation by the proteasome. It is less clear how monoubiquitylation targets proteins to cellular compartments and modifies their activity.
- V(D)J RECOMBINATION
-
A specialized form of recombination that assembles the genes that encode lymphocyte antigen receptors from variable (V), diversity (D) and joining (J) gene segments. DNA double-stranded breaks are introduced between the V, D and J segments and DNA repair/recombination proteins then join the segments together.
- ERROR-PRONE REPAIR PATHWAYS
-
DNA lesions can be repaired by many competing pathways. Some of these pathways repair damaged DNA with reduced fidelity, giving rise to new mutations.
- TRANSCRIPTION-COUPLED REPAIR
-
A specialized pathway that facilitates the repair of damaged DNA during transcription, in which the transcribed strand of active genes is preferentially repaired. This process is defective in the genetic disease, Cockayne syndrome.
Rights and permissions
About this article
Cite this article
Joenje, H., Patel, K. The emerging genetic and molecular basis of Fanconi anaemia. Nat Rev Genet 2, 446–458 (2001). https://doi.org/10.1038/35076590
Issue Date:
DOI: https://doi.org/10.1038/35076590
This article is cited by
-
A novel frame-shift deletion in FANCF gene causing autosomal recessive Fanconi anemia: a case report
BMC Medical Genetics (2019)
-
Differentiation of MISSLA and Fanconi anaemia by computer-aided image analysis and presentation of two novel MISSLA siblings
European Journal of Human Genetics (2019)
-
Map of synthetic rescue interactions for the Fanconi anemia DNA repair pathway identifies USP48
Nature Communications (2018)