Abstract
Chinese indigenous sheep can be classified into three types based on tail morphology: fat-tailed, fat-rumped, and thin-tailed sheep, of which the typical breeds are large-tailed Han sheep, Altay sheep, and Tibetan sheep, respectively. To unravel the genetic mechanisms underlying the phenotypic differences among Chinese indigenous sheep with tails of three different types, we used ovine high-density 600K SNP arrays to detect genome-wide copy number variation (CNV). In large-tailed Han sheep, Altay sheep, and Tibetan sheep, 371, 301, and 66 CNV regions (CNVRs) with lengths of 71.35 Mb, 51.65 Mb, and 10.56 Mb, respectively, were identified on autosomal chromosomes. Ten CNVRs were randomly chosen for confirmation, of which eight were successfully validated. The detected CNVRs harboured 3130 genes, including genes associated with fat deposition, such as PPARA, RXRA, KLF11, ADD1, FASN, PPP1CA, PDGFA, and PEX6. Moreover, multilevel bioinformatics analyses of the detected candidate genes were significantly enriched for involvement in fat deposition, GTPase regulator, and peptide receptor activities. This is the first high-resolution sheep CNV map for Chinese indigenous sheep breeds with three types of tails. Our results provide valuable information that will support investigations of genomic structural variation underlying traits of interest in sheep.
Similar content being viewed by others
Introduction
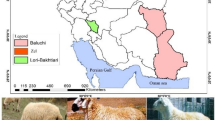
Chinese indigenous sheep breeds can be classified into fat-tailed, fat-rumped, and thin-tailed sheep breeds. Worldwide, more than 25% of sheep breeds are fat-tailed or fat-rumped and store large amounts of fat in the tail1. It is thought that fat-tailed sheep, which appeared during the process of domestication, are the result of breeding by humans and natural selection2. Fat-tailed sheep were first documented approximately 5000 years ago2. The tails of fat-tailed and fat-rumped sheep are considered to be adaptations to hostile environments and act as energy reserves to support migration and survival during cold winters3. In addition, the fat in the tails of fat-tailed and fat-rumped sheep can be used by humans as a high-energy food during periods of drought and famine2,4. Therefore, natural and artificial selection has led to an increased prevalence of fat in sheep tails across generations3,4.
Copy number variation (CNV), a type of genomic structural variation, refers to DNA segments with sizes ranging from 1 kilobase (Kb) to several megabases (Mb) in which duplication or deletion events have occurred5. CNVs are ubiquitously distributed in the human genome and can influence phenotypic variation via changes in gene expression and gene dosage6,7,8. CNV is a major source of genetic variation and phenotypic diversity8,9. CNV in the ASIP gene determines the white and grey coat phenotypes in goats10. In addition, CNV in the KIT gene leads to a white coat colour in pigs11, whereas the phenotype of the pea comb in chickens is influenced by CNV in intron 1 of SOX512. Chinese indigenous sheep breeds show tremendous phenotypic differences, especially in the tail. Large-tailed Han sheep have the largest and fattiest tail of any Chinese sheep breed, characterized by being long, broad, and fat tail with a thin, twisted end turned upwards between two lobes and a broad base. Altay sheep have fat deposits in the buttocks that look like a tail; thus, Altay sheep are known as fat-rumped sheep, while Tibetan sheep have a thin tail the shape of a flattened cone13. To understand the genetic basis of these three types of tails, we aimed to identify the potential contribution of CNV to phenotypic variation.
Therefore, the primary aim of this study is to detect CNV in Chinese indigenous sheep breeds with different types of tails. Genome-wide CNV detection was carried out by high-density 600K SNP genotype arrays. The PennCNV program was used to analyse the data and validate the CNVRs by qPCR. To the best of our knowledge, this is the first sheep CNVR map constructed based on high-density 600K SNP genotype data. We also compared our results with those from previous studies of sheep CNV. This study provides a comprehensive map of CNV in the ovine genome that will guide studies aimed at elucidating the causes of complex traits.
Results
SNP genotyping
A total of 120 sheep were genotyped using the Illumina Ovine SNP 600K BeadChip. Samples with low-quality signal intensity were excluded based on the CNV quality control filter criteria. A total of 110 individuals from three sheep breeds (36 large-tailed Han sheep, 37 Altay sheep, and 37 Tibetan sheep) were analysed.
Genome-wide detection of CNVs in sheep
Table 1 summarizes the initial numbers of CNVs and CNVRs as identified by PennCNV software in the three selected breeds. After merging overlapping CNVs, a total of 371, 301, and 66 CNVRs with lengths of 71.35 Mb, 51.65 Mb, and 10.56 Mb were detected on 26 pairs of autosomal chromosomes in large-tailed Han sheep, Altay sheep, and Tibetan sheep, respectively (Supplementary File 1: Tables S1–S3). A number of differences were found in the numbers of CNVRs among the three breeds. A total of 216 CNVRs were shared by large-tailed Han sheep and Altay sheep, 56 CNVRs shared by large-tailed Han sheep and Tibetan sheep, 55 CNVRs shared by Altay sheep and Tibetan sheep, and 55 CNVRs shared by all three breeds, as shown in Fig. 1. A CNV map of sheep with different types of tails was constructed by removing repetitive CNV regions, as shown in Fig. 2. CNVRs were not uniformly distributed across chromosomes. Among the 490 CNVRs, 93 were gains, 390 were losses, and 7 were losses and gains within the same region. The length of the CNVRs ranged from 100.11 to 804.18 Kb, with an average length of 165.39 Kb and a median length of 133.17 Kb (Supplementary File 1: Table S4). The ratio of CNVRs on 26 pairs of autosomal chromosomes ranged from 1.16% to 82.50%. Chromosome 3 had the most CNVRs (42), as shown in Table 2. No CNVRs were identified on chromosome 25 or 26 in Tibetan sheep (Supplementary File 2: Fig. S1).
CNV Validation by qPCR
To confirm the identified CNVRs, we randomly chose 10 CNVRs for validation by qPCR. The 10 selected CNVRs represented different types of CNV (gain, loss, and gain/loss) and were chosen from all three of the tested sheep breeds. Eight (80%) of the randomly selected CNVRs were confirmed in agreement with PennCNV (Fig. 3).
The y-axis shows the NR values obtained by QPCR, and the x-axis shows the sample names in the different CNV regions. Samples with NRs of approximately 2 denote normal individuals (two copy), samples with NRs of approximately 1 denote one-copy-loss individuals (one copy), and samples with NRs of approximately 3 or more denote copy-number-gain individuals.
Gene content of sheep CNVRs
In large-tailed Han sheep, a total of 2881 annotated genes within 329 CNVRs (88.67%) were identified from the Ensembl Genes 64 Database using the BioMart data management system, while the remaining 42 CNVRs (11.32%) lacked annotated genes. Within the 301 CNVRs that were detected in Altay sheep, 244 CNVRs (81.06%) harboured 1795 annotated genes, whereas the other 57 CNVRs (18.94%) did not contain annotated genes. In Tibetan sheep, 293 annotated genes were contained within 60 CNVRs (90.90%), whereas the remaining 6 CNVRs (9.09%) lacked annotated genes. A total of 1839 common genes were shared among large-tailed Han sheep, Altay sheep, and Tibetan sheep. Therefore, a total of 3130 genes resided in the detected CNVRs.
To provide insight into the functions of the genes in the identified CNVRs, GO and KEGG pathway analyses were performed using the DAVID tool. After the conversion of Ovis aries Ensembl gene IDs to human orthologue Ensembl gene IDs using BioMart, 3016 corresponding human gene IDs remained in DAVID. ‘Human’ was selected as the background for subsequent analysis. After Bonferroni correction, 50 GO terms were identified as significantly enriched (Supplementary File 2: Table S5). The significantly enriched GO terms were involved in thyroid hormone receptor activator activity (GO: 0010861) and coactivator activity associated with energy expenditure (GO: 0030375)14. In addition, some significantly enriched GO terms were involved in GTPase regulator activity (GO: 0030695), GTPase activator activity (GO: 00005096), regulation of GTPase activity (GO: 0043087), regulation of Ras protein signal transduction (GO: 0046578), skeletal system development (GO: 0001501), embryonic morphogenesis (GO: 0048706), embryonic limb morphogenesis (GO: 0048598), transcription regulator activity (GO: 0030528), and other biological processes. KEGG pathway analysis revealed that the genes in the identified CNVRs were enriched in 2 significant pathways: ribosome and notch signalling (Supplementary File 2: Table S6).
Difference between fat-tailed and thin-tailed sheep breeds
Among the breeds that were selected for this study, large-tailed Han sheep and Altay sheep are fat-tailed and fat-rumped sheep, respectively, while Tibetan sheep are thin-tailed sheep. There were many differences in the CNV of fat-tailed and thin-tailed sheep. Fat-tailed sheep harboured 211 CNVRs covering 56 Mb of the sheep genome sequence and 123 genes. After GO enrichment analysis, some genes associated with fat were identified (Table 3). Seven genes associated with fat were identified in the CNVRs of large-tailed Han sheep, including peroxisome proliferator-activated receptor-α (PPARA), retinoic X receptor A (RXRA), Kruppel-like factor 11 (KLF11), adipocyte determination and differentiation factor 1 (ADD1), fatty acid synthase (FASN), phosphoprotein phosphatase 1 catalytic subunit A (PPP1CA), and platelet-derived growth factor alpha (PDGFA). In Altay sheep, 5 genes in CNVRs were associated with fat deposition, including peroxin 6 (PEX6), RXRA, FASN, PPP1CA, and PDGFA. Tibetan sheep are thin-tailed sheep native to a high-altitude plateau. Sixty-six CNVRs were identified in Tibetan sheep; however, no CNVRs were detected on chromosome 25 or 26. In Tibetan sheep, only 1 gene related to fat deposition was found within the identified CNVRs: RXRA. Two genes related to adaptation to high altitude were found within the identified CNVRs in Tibetan sheep: α-ketoglutarate-dependent dioxygenase alkB homologue 5 (ALKBH5) and nuclear prelamin A recognition factor-like (NARFL).
Comparison with other studies on CNV in sheep
Our results were compared to those of previous reports on sheep genomic CNV. The first study on sheep genome CNV was conducted by Fontanesi et al.15 using a tiling oligonucleotide array with approximately 385,000 probes that was used to detect 135 CNVRs from 11 ewes belonging to 6 different Italian dairy or dual-purpose sheep breeds. Eleven of the 135 CNVRs identified were also identified in our results (Supplementary File 1: Table S7, Table 4). Liu16 identified 238 CNVRs by assessing 329 individuals from 3 breeds using the Ovine SNP 50 BeadChip, but only 12 of these CNVRs on 10 chromosomes were also identified in our results (Supplementary File 1: Table S8). The most recent study on CNV in sheep was carried out by Ma17, who identified 111 CNVRs by subjecting 160 individuals from 8 breeds to analysis using the Ovine SNP 50 BeadChip; only 8 of these CNVRs on 4 chromosomes were also identified in our results (Supplementary File 1: Table S9).
Discussion
In recent years, with the development of high-throughput genotyping technology, CNV detection using SNP chip data has been conducted in humans and animals18,19,20. Several algorithms for inferring CNV based on SNP chip data have been developed, including PennCNV, cnvPartition, and QuantiSNP. No single approach can capture all CNV; thus, one may be complementary to another. However, most studies on CNV using SNP arrays in animals and humans have used only PennCNV software21,22,23, especially for high-density SNP data22,23,24,25,26,27,28. PennCNV detects CNV more reliably than do some other algorithms29 because PennCNV can incorporate multiple sources of information, including total signal intensity and allelic frequency SNPs30,31. When multiple algorithms are used to detect CNV, determining the appropriate number of CNVs may be difficult. If we accept only CNVs that are commonly detected by all algorithms, many CNVs are missed, but many false-positive CNVs are identified if we accept all CNVs detected by different algorithms. Finally, high-density SNP chips and stricter filtering criteria could ensure a high positive validation rate. Although we used only PennCNV to detect CNVs, two strict criteria were adopted to reduce the risk of false-positive results: SD of LRR < 0.30 and BAF = 0.01. These strict criteria and high-density SNP array data likely resulted in the high qPCR confirmation percentage of 80% in the present study. In our study, most of the predicted positive samples identified by the PennCNV program agreed well with the qPCR experiments and showed a high level of copy number concordance between them. The discrepancy between PennCNV and qPCR findings may indicate uncertain boundaries of CNV based on the SNP array, as well as the potential impacts of SNPs and small indels.
It is notable that few of the CNVRs that were identified in this study were identified in other sheep CNV studies, similar to the results of other CNV studies in humans and mammals26,36. The inconsistency between results from different studies might be due to the following reasons: (1) different genetic backgrounds and sample sizes; (2) different chip platforms (Illumina SNP chips or CGH arrays); and (3) SNP chips with different densities and CNV-calling algorithms.
In this study, we selected three sheep breeds distributed in different parts of China with obviously different types of tails: Altay sheep, large-tailed Han sheep and Tibetan sheep. Altay sheep live in the Gobi Desert, where the annual average temperature and extreme minimum temperature are 4.0 °C and −42.7 °C, respectively, and the ground is snow-covered for 200–250 days of the year. In such an environment, fat tends to be deposited on the rump of the animal, forming a substantial buttock that is an adaptation to the harsh environment of the Gobi Desert. Our results indicate that the process of adaptation to extreme climates in Altay sheep is principally mediated through complex, integrated metabolic responses, as has been confirmed in rodents33. Climate has a significant impact on animal fitness and physiology, especially in ruminants34. Climate factors, including solar radiation, temperature, UV radiation, humidity, and precipitation, have direct impacts on sheep; some of these climate factors, such as temperature and precipitation, also directly affect the quality, quantity, and digestibility of food, thus impacting sheep indirectly35. Therefore, long-term thermal stress can lead to energy metabolic adaptation, as well as cold and heat tolerance, in particular breeds36. At the same time, sheep with different types of tails (fat-tailed and thin-tailed) also follow basic thermoregulatory principles to conserve or dissipate body energy during different seasons. The ability to deposit fat in the tail is beneficial in extremely harsh environments and seasons. Large-tailed Han sheep are primarily produced in the hinterland of the northern Chinese plain, which has clear seasonal variation, with cold, dry winters and hot, rainy summers. We found some candidate genes involved in endocrine regulation, consistent with the impact of day length and seasonal timing on sheep physiology and evolutionary fitness. The tails of large-tailed Han sheep show hypertrophy and have a fan-like shape, sagging below the hock and dragging on the ground in some individuals. During periods of famine, the tails of fat-tailed sheep can be used as food.
Thin-tailed Tibetan sheep live in the mountainous region of the Tibetan plateau at an altitude of 3000–5000 metres. Tibetan sheep are relatively strong and tall, with a thin tail in the shape of a flat cone. We found that fat-tailed and fat-rumped sheep had more CNVRs (371 and 301, respectively) than did thin-tailed sheep (66). In the CNVRs of fat-tailed sheep, we detected some genes associated with fat. In the CNVRs of thin-tailed sheep, we found two genes (ALKBH5 and NARFL) associated with altitude adaptation. It is generally believed that the first wild sheep were thin-tailed sheep, from which fat-tailed sheep were produced as the result of selection by humans and by nature37. These results indicate that natural and artificial selection for fat deposition in sheep tails may lead to CNV alterations.
In this study, we identified genes in CNVRs associated with fat synthesis and metabolism in fat-tailed (7 genes), fat-rumped (5 genes), and thin-tailed (1 gene) sheep. RXRA was identified in CNVRs in all three types of sheep, indicating that it may play an important role in the maintenance of basic fat metabolism. PPARA, RXRA, KLF11, ADD1, FASN, PPP1CA, PDGFA, and PEX6 were only identified in fat-tailed and fat-rumped sheep, indicating that these genes may cause tail fat deposition. PPARA is a member of the PPAR family. Peroxisomes play an important role in fatty acid metabolism. Unsaturated fatty acids bind to PPARA, which is highly expressed in the liver, skeletal muscle, heart, and kidney38, where it activates genes that are involved in fatty acid metabolism. RXRA is a member of the nuclear hormone superfamily and forms heterodimers with a number of nuclear receptors, including the thyroid hormone receptor, vitamin D receptor, GR, and PPAR39. RXRA can mediate heterodimerization with retinoic acid receptors (RARs) and is essential for the functional activation of RARs by their ligands40. RXRA plays a role in lipid homeostasis41,42 and foetal development39. KLF11 (Kruppel-like factor 11) is a member of the Kruppel-like factor family and is expressed in many human tissues43. Loft44 reported that KLF11 is a browning transcription factor in human adipocytes. PPAR directly induces KLF11 and, together with KLF11, activates and maintains the brite-selective gene programme. ADD1 (adipocyte determination- and differentiation-dependent factor 1), a member of the basic helix-loop-helix (bHLH) family of transcription factors, is associated with adipocyte differentiation and cholesterol homeostasis45. ADD1 plays an important role in the process of adipocyte differentiation46, as well as in the activation of PPAR and C/EBPs. FASN (fatty acid synthase) plays a key role in the de novo synthesis of fatty acids in mammals. FASN catalyses reaction steps in the conversion of acetyl-CoA, malonyl-CoA, and NADPH into saturated fatty acids. FASN is expressed in the mammalian liver and adipose tissue, where it regulates body fat deposition and fatty acid anabolism47. PPP1CA (phosphoprotein phosphatase 1 catalytic subunit alpha isozyme) is an isoform of PPP1C48, which was first identified as the enzyme that converts phosphorylase A into phosphorylase B49. A number of roles have been attributed to PPP1C in the regulation of important cellular events. PDGFA (platelet-derived growth factor A) is a PDGF subunit50. PDGF has been directly implicated in developmental and physiological processes51. PEX6 (peroxin 6) is a member of the PEX family. Peroxidase is involved in fatty acid metabolism, glycometabolism, toxic material degradation, and oxygen concentration regulation. The biosynthesis and proliferation of the peroxisome are regulated by PPARs and PEX. Because super-long-chain fatty acids cannot enter mitochondria52, they must be oxidized into short fatty acids by peroxidase. Some of the differences may be due to selection because of different environments. We also identified Rab family protein TBC1D13, a GTPase activator that plays an important role in increasing GTPase activity53. GTP hydrolysis is catalysed by GTPases, after which GDP serves as an energy source in metabolic reactions within the cell.
To better understand the molecular function of these candidate genes, we examined their GO classification. Many of the genes that were detected in our study were consistent with prior expectations because they are involved in adipose tissue energy balance, skeletal system development, and metabolic activities. Two intriguing candidate GO terms were thyroid hormone receptor activator activity (GO: 0010861) and coactivator activity (GO: 0030375) related to energy expenditure. These observations should be explored and verified in future studies.
Conclusions
We performed genome-wide CNV detection based on ovine high-density genotyping data from 120 sheep with three types of tails and provide the highest resolution CNV map of the sheep genome produced to date. Among the 490 CNVRs, 93 were gains, 390 were losses, and 7 were losses and gains within the same region. The length of the CNVRs ranged from 100.11 to 804.18 Kb, with an average length of 165.39 Kb and a median length of 133.17 Kb. A total of 371, 301, and 66 CNVRs were identified in large-tailed Han sheep, Altay sheep, and Tibetan sheep, respectively. The CNVR regions harboured genes associated with fat deposition and fat synthesis in fat-tailed sheep and fat-rumped sheep. The CNVR regions harboured genes associated with adaptation to the low oxygen environment of the Tibetan plateau in Tibetan sheep. These candidate genes included PPARA, RXRA, KLF11, ADD1, FASN, PPP1CA, PDGFA, and PEX6. Moreover, multilevel bioinformatics analyses of the detected candidate genes showed that they were likely involved in fat deposition, GTPase regulation, and peptide receptor activities. This study is an important step towards the generation of a CNV map of the ovine genome and provides an important resource for studies of ovine genomic variation. In addition, our findings provide valuable information that will facilitate studies investigating genomic structural variation underlying traits of interest in sheep, including types of sheep tails, fat deposition in sheep tails, and environmental adaptability.
Materials
Ethics Statement
All of the animal procedures were performed in strict accordance with the guidelines proposed by the China Council on Animal Care and the Ministry of Agriculture of the People’s Republic of China. All of the animal experiments were approved by the Chinese Academy of Agricultural Sciences (CAAS) (Beijing, China).
DNA sample collection
For this study, 120 individuals from 3 breeds, including 40 large-tailed Han sheep (20 rams and 20 ewes), 40 Altay sheep (20 rams and 20 ewes), and 40 Tibetan sheep (20 rams and 20 ewes), were collected from Liaocheng in Shandong Province, Altay in Xinjiang Province, and Tianzhu in Gansu Province, respectively. All of the specimens were randomly selected.
Genomic DNA samples were extracted from blood using the TIANamp Blood DNA Kit (Tiangen Biotech Co. Ltd., Beijing, China). The purity and concentration of genomic DNA were measured using a NanoVue Spectrophotometer.
Genotyping and quality control
The genomic DNA of each specimen was genotyped with the Illumina Ovine SNP 600 BeadChip (Illumina Inc., CA, USA), which contains 604,715 SNPs spanning the ovine genome with an average distance of 4.28 Kb.
To increase the accuracy of CNV inference, stringent quality control criteria were applied: (1) individual call rate >95% and call frequency >90%; (2) LRR standard deviation <0.30; (3) BAF drift <0.01; and (4) default waviness factor.
Detection of CNV
Similar to previous studies27, chromosomes X and Y were excluded from the analysis. PennCNV software was only used to detect CNV on autosomes and was downloaded from http://penncnv.openbioinformatics.org/en/latest/user-guide/download/. The PennCNV algorithm incorporates multiple information sources, including the total signal intensity of the log R ratio (LRR), B allele frequency (BAF), and population frequency of the B allele (PFB)54. The LRR and BAF of each SNP for every sample were exported from Illumina GenomeStudio software. The PFB was generated based on the BAF of each SNP marker. Genomic waves were adjusted with the sheep GC model file, which was generated by calculating the GC content of 1-Mb genomic regions surrounding each marker (500 Kb on each side), after the program argument ‘gcmodel’ was used to adjust the results. According to previous research using high-density SNP chips to detect CNV in humans55,56, the CNV filter was based on three criteria: (1) the CNV must contain ten or more consecutive SNPs; (2) the length of the CNV must be at least 100 K; and (3) the CNV must be present in at least one animal.
Finally, following the method reported by Redon et al., CNVRs were obtained by merging CNVs with overlapping regions that were identified across all samples7.
Experimental validation of CNVRs by quantitative PCR
Quantitative real-time PCR (qPCR) was used to validate the CNVRs that were detected in this study. First, qPCR was performed to validate 10 randomly selected CNVRs that were identified by PennCNV using genomic DNA of the same sheep as analysed by the Illumina chip. The GAPDH gene was chosen as a reference gene for all of the qPCR experiments15. The primers were designed using the Primer 3 webtool (http://frodo.wi.mit.edu/primer3/). All of the PCR primers were designed based on NCBI reference sequences (Supplementary File 1: Table S10). PCR reactions were prepared using the Power SYBR Green PCR reagent kit (Takara Bio). PCR was carried out in a total volume of 20 μL containing 1 μL of DNA (approximately 60 ng), forward primer and reverse primers (2 μL total), 10 μL of Master Mix (2×), and 15 μL of water. PCR was performed with the following cycling conditions: 5 min at 95 °C, followed by 40 cycles at 95 °C for 10 sec and 60 °C for 10 sec. All of the reactions were run in triplicate and included blank controls. The 2−ΔΔCt method was used to calculate the copy number57,58,59, where Ct is the threshold cycle. The copy number of the target gene in the test sample was calculated as 2 × 2−ΔΔCt, where ΔΔCt was defined as in a previous report60. The corresponding equation was

where Ct is the threshold cycle, sample A is the tested individual, and sample B is the control individual.
Gene detection and functional analysis
Genes harboured in the inferred CNVRs were obtained from the Ensembl Genes 64 Database using BioMart software based on the Ovis aries (Oar_v3.1) gene sequence assembly. The Database for Annotation, Visualization and Integrated Discovery (DAVID) (http://david.abcc.ncifcrf.gov/) was used to perform the gene ontology (GO)61 enrichment analysis and Kyoto Encyclopedia of Gene and Genome (KEGG)62 pathway analysis. To better understand the functions of the genes within the detected CNVRs, the Ovis aries Ensembl gene IDs were converted into human orthologue Ensembl gene IDs using BioMart because the annotation of the sheep genome is limited.
Additional Information
How to cite this article: Zhu, C. et al. Genome-wide detection of CNVs in Chinese indigenous sheep with different types of tails using ovine high-density 600K SNP arrays. Sci. Rep. 6, 27822; doi: 10.1038/srep27822 (2016).
References
Safdarian, M., Zamiri, M. J., Hashemi, M. & Noorolahi, H. Relationships of fat-tail dimensions with fat-tail weight and carcass characteristics at different slaughter weights of Torki-Ghashghaii sheep. Meat Sci. 80, 686–689 (2008).
Moradi, M. H., Nejati-Javaremi, A., Moradi-Shahrbabak, M., Dodds, K. G. & McEwan, J. C. Genomic scan of selective sweeps in thin and fat tail sheep breeds for identifying of candidate regions associated with fat deposition. BMC Genet. 13, 10 (2012).
Ermias, E., Yami, A. & Rege, J. Fat deposition in tropical sheep as adaptive attribute to periodic feed fluctuation. J. Anim. Breed. Genet. 119, 235–246 (2002).
Kashan, N. E. J., Azar, G. H. M., Afzalzadeh, A. & Salehi, A. Growth performance and carcass quality of fattening lambs from fat-tailed and tailed sheep breeds. Small Ruminant Res. 60, 267–271 (2005).
Mills, R. E. et al. Mapping copy number variation by population-scale genome sequencing. Nature 470, 59–65 (2011).
Conrad, D. F. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010).
Redon, R. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006).
Zhang, F., Gu, W., Hurles, M. E. & Lupski, J. R. Copy number variation in human health, disease, and evolution. Annu. Rev. Genomics Hum. Genet. 10, 451–481 (2009).
Stankiewicz, P. Structural variation in the human genome and its role in disease. Annu. Rev. Med. 61, 437–455 (2010).
Fontanesi, L. Copy number variation and missense mutations of the Agouti signaling protein (ASIP) gene in goat breeds with different coat colors. Cytogenet. Genome Res. 126, 333–347 (2009).
Giuffra, E. et al. A large duplication associated with dominant white color in pigs originated by homologous recombination between LINE elements flanking KIT. Mamm. Genome 13, 569–577 (2002).
Wright, D. et al. Copy number variation in intron 1 of SOX5 causes the Pea-comb phenotype in chickens. PLoS Genet. 5, e1000512 (2009).
China National Commission of Animal Genetic Resources. Animal genetic resource in China-Sheep and Goats. Vol. 1, (eds Du, L. et al.) 24–121 (Chinese Agricultural Press, 2011).
Mullur, R., Liu, Y.-Y. & Brent, G. A. Thyroid hormone regulation of metabolism. Physiol. Rev. 94, 355–382 (2014).
Fontanesi, L. et al. A first comparative map of copy number variations in the sheep genome. Genomics 97, 158–165 (2011).
Liu, J. et al. Analysis of copy number variations in the sheep genome using 50K SNP BeadChip array. BMC Genomics 14, 433–440 (2013).
Ma, Y., Zhang, Q., Lu, Z., Zhao, X. & Zhang, Y. Analysis of copy number variations by SNP50 BeadChip array in Chinese sheep. Genomics 106, 295–300 (2015).
Diskin, S. J. et al. Copy number variation at 1q21.1 associated with neuroblastoma. Nature 459, 987–991 (2009).
Kirov, G. The role of copy number variation in schizophrenia. Expert. Rev. Neurother. 10, 25–32 (2010).
Fontanesi, L. et al. Copy Number Variation and Missense Mutations of the Agouti Signaling Protein (ASIP) Gene in Goat Breeds with Different Coat Colors. Cytogenet. Genome Res. 126, 333–347 (2009).
Wang, M. D., Dzama, K., Hefer, C. A. & Muchadeyi, F. C. Genomic population structure and prevalence of copy number variations in South African Nguni cattle. BMC Genomics 16, 1 (2015).
Wang, Y. et al. Analysis of genome-wide copy number variations in Chinese indigenous and western pig breeds by 60 K SNP genotyping arrays. PLoS One 9, e106780 (2014).
Bae, J. S. et al. Identification of copy number variations and common deletion polymorphisms in cattle. BMC Genomics 11, 232 (2010).
Zhang, Q., Ma, Y., Wang, X., Zhang, Y. & Zhao, X. Identification of copy number variations in Qinchuan cattle using BovineHD Genotyping Beadchip array. Mol. Genet. Genomics 290, 319–327 (2014).
Yi, G., Qu, L., Chen, S., Xu, G. & Yang, N. Genome‐wide copy number profiling using high‐density SNP array in chickens. Anim. Genet. 46, 148–157 (2015).
Jiang, L. et al. Genome-wide detection of copy number variations using high-density SNP genotyping platforms in Holsteins. BMC Genomics 14, 131 (2013).
Hou, Y. et al. Genomic characteristics of cattle copy number variations. BMC Genomics 12, 127 (2011).
Schiavo, G. et al. Copy number variants in Italian Large White pigs detected using high-density single nucleotide polymorphisms and their association with back fat thickness. Anim. Genet. 45, 745–749 (2014).
Marenne, G. et al. Assessment of copy number variation using the Illumina Infinium 1M SNP-array: a comparison of methodological approaches in the Spanish Bladder Cancer/EPICURO study. Hum. Mutat. 32, 240–248 (2011).
Winchester, L., Yau, C. & Ragoussis, J. Comparing CNV detection methods for SNP arrays. Brief. Funct. Genomics Proteomics 8, 353–366 (2009).
Pinto, D. et al. Comprehensive assessment of array-based platforms and calling algorithms for detection of copy number variants. Nat. Biotechnol. 29, 512–520 (2011).
Wu, Y. et al. A genome-wide scan for copy number variations using high-density single nucleotide polymorphism array in Simmental cattle. Anim. Genet. 46, 289–298 (2015).
Rezende, E. & Bozinovic, F. T. Climatic adaptation and the evolution of basal and maximum rates of metabolism in rodents. Evolution 58, 1361–1374 (2004).
Hofmann, R. R. Evolutionary steps of ecophysiological adaptation and diversification of ruminants: a comparative view of their digestive system. Oecologia 78, 443–457 (1989).
Mcmanus, C. et al. Skin and coat traits in sheep in Brazil and their relation with heat tolerance. Trop. Anim. Health Pro. 43, 121–126 (2011).
Lv, F. H. et al. Adaptations to Climate-Mediated Selective Pressures in Sheep. Mol. Biol. Evol. 31, 3324–3343 (2014).
Moradi, M. H., Nejati-Javaremi, A., Moradi-Shahrbabak, M., Dodds, K. G. & McEwan, J. C. Genomic scan of selective sweeps in thin and fat tail sheep breeds for identifying of candidate regions associated with fat deposition. BMC Genetics 13, 10 (2012).
Braissant, O., Foufelle, F., Scotto, C., Dauça, M. & Wahli, W. Differential expression of peroxisome proliferator-activated receptors (PPARs): tissue distribution of PPAR-alpha,-beta, and-gamma in the adult rat. Endocrinology 137, 354–366 (1996).
Harvey, N. C. et al. Childhood bone mineral content is associated with methylation status of the RXRA promoter at birth. J. Bone Miner. Res. 29, 600–607 (2014).
Zhang, J., Ma, X., Wang, H., Ma, D. & Huang, G. Elevated methylation of the RXRA promoter region may be responsible for its downregulated expression in the myocardium of patients with TOF. Pediatr. Res. 75, 588–594 (2014).
Moreau, A., Vilarem, M. J., Maurel, P. & Pascussi, J. M. Xenoreceptors CAR and PXR activation and consequences on lipid metabolism, glucose homeostasis, and inflammatory response. Mol. Pharm. 5, 35–41 (2008).
Wada, T. PXR and CAR in energy metabolism. Trends Endocrin. Met. 20, 273–279 (2009).
Yin, K. J. et al. KLF11 mediates PPAR cerebrovascular protection in ischaemic stroke. Brain 136, 1274–1287 (2013).
Loft, A. et al. Browning of human adipocytes requires KLF11 and reprogramming of PPARγ superenhancers. Gene Dev. 29, 7–22 (2015).
Kim, J. B. & Dual D. N. A. binding specificity of ADD1/SREBP1 controlled by a single amino acid in the basic helix-loop-helix domain. Mol. Cell. Biol. 15, 2582–2588 (1995).
Spiegelman, B. M. & Flier, J. S. Adipogenesis and obesity: rounding out the big picture. Cell 87, 377–389 (1996).
Jun, L. et al. Structural and Functional Analysis of Fatty Acid Synthase Gene Promoter of Goat. Chin. J. Anim. Vet. Sci. 43, 204–210 (2012).
Silva, J. V., Freitas, M. J. & Fardilha, M. Phosphoprotein phosphatase 1 complexes in spermatogenesis. Curr. Mol. Pharmacol. 7, 136–146 (2014).
Krebs, E. G. & Fischer, E. H. Phosphorylase activity of skeletal muscle extracts. J. Biol. Chem. 216, 113–120 (1955).
Bhandari, B., Wenzel, U. O., Marra, F. & Abboud, H. E. A nuclear protein in mesangial cells that binds to the promoter region of the platelet-derived growth factor-A chain gene. Induction by phorbol ester. J. Biol. Chem. 270, 5541–5548 (1995).
O’Neill, G. T. & Akhurst, R. J. Growth Factor Disturbance in Handbook of Experimental Pharmacology, Vol. 124 (eds Kavlock, R. J. et al.) Ch. 11, 301–324 (Springer, 1997).
Titorenko, V. I. & Rachubinski, R. A. The life cycle of the peroxisome. Nat. Rev. Mol. Cell Biol. 2, 357–368 (2001).
Ishibashi, K., Kanno, E., Itoh, T. & Fukuda, M. Identification and characterization of a novel Tre-2/Bub2/Cdc16 (TBC) protein that possesses Rab3A-GAP activity. Genes Cells 14, 41–52 (2009).
Wang, K. et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 17, 1665–1674 (2007).
Jakobsson, M. et al. Genotype, haplotype and copy-number variation in worldwide human populations. Nature 451, 998–1003 (2008).
Glessner, J. T. et al. Strong synaptic transmission impact by copy number variations in schizophrenia. Proc. Natl. Acad. Sci. USA 107, 10584–10589 (2010).
Bodin, L., Beaune, P. H. & Loriot, M. A. Determination of Cytochrome P450 2D6 (CYP2D6) Gene Copy Number by Real-Time Quantitative PCR. J. Biomed. Biotechnol. 2005, 248–253 (2005).
D’Haene, B., Vandesompele, J. & Hellemans, J. Accurate and objective copy number profiling using real-time quantitative PCR. Methods 50, 262–270 (2010).
Livak, K. J. & Schmittgen, T. D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 261ΔΔCT Method. Methods 25, 402–408 (2001).
Fadista, J. o., Nygaard, M., Holm, L. E., Thomsen, B. & Bendixen, C. A Snapshot of CNVs in the Pig Genome. Plos One 3, e3916 (2008).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Kanehisa, M., Goto, S., Furumichi, M., Tanabe, M. & Hirakawa, M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 38, D355–360 (2010).
Acknowledgements
We would like to thank the two anonymous reviewers for their constructive comments and suggestions, which greatly improved our manuscript. This work was supported by the Agricultural Science and Technology Innovation Program (No. ASTIP-IAS-TS-6), the National Natural Science Foundation of China (No. 31572357), and the National Modern Agricultural Industry Technology Fund for Scientists in Sheep Industry System (No. CARS-39-04B).
Author information
Authors and Affiliations
Contributions
F.Z. and L.D. conceived and designed the experiments. C.Z. performed the experiments. C.Z., H.F. and Z.Y. analysed the data. C.Z. and H.F. carried out experimental validation. S.H., X.M., J.X., H.W., L.Z., C.W. and Q.Z. contributed reagents/materials/analysis tools. C.Z. and F.Z. wrote the paper. All of the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhu, C., Fan, H., Yuan, Z. et al. Genome-wide detection of CNVs in Chinese indigenous sheep with different types of tails using ovine high-density 600K SNP arrays. Sci Rep 6, 27822 (2016). https://doi.org/10.1038/srep27822
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep27822
This article is cited by
-
Whole-body adipose tissue multi-omic analyses in sheep reveal molecular mechanisms underlying local adaptation to extreme environments
Communications Biology (2023)
-
Genetics of the phenotypic evolution in sheep: a molecular look at diversity-driving genes
Genetics Selection Evolution (2022)
-
Genome-wide identification of copy number variation and association with fat deposition in thin and fat-tailed sheep breeds
Scientific Reports (2022)
-
A global analysis of CNVs in Chinese indigenous fine-wool sheep populations using whole-genome resequencing
BMC Genomics (2021)
-
Genome-wide characterization of copy number variations in the host genome in genetic resistance to Marek’s disease using next generation sequencing
BMC Genetics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.