Abstract
Converting biomass to biofuels is a key strategy in substituting fossil fuels to mitigate climate change. Conventional strategies to convert lignocellulosic biomass to ethanol address the fermentation of cellulose-derived glucose. Here we used super-resolution fluorescence microscopy to uncover the nanoscale structure of cell walls in the energy crops maize and Miscanthus where the typical polymer cellulose forms an unconventional layered architecture with the atypical (1, 3)-β-glucan polymer callose. This raised the question about an unused potential of (1, 3)-β-glucan in the fermentation of lignocellulosic biomass. Engineering biomass conversion for optimized (1, 3)-β-glucan utilization, we increased the ethanol yield from both energy crops. The generation of transgenic Miscanthus lines with an elevated (1, 3)-β-glucan content further increased ethanol yield providing a new strategy in energy crop breeding. Applying the (1, 3)-β-glucan-optimized conversion method on marine biomass from brown macroalgae with a naturally high (1, 3)-β-glucan content, we not only substantially increased ethanol yield but also demonstrated an effective co-fermentation of plant and marine biomass. This opens new perspectives in combining different kinds of feedstock for sustainable and efficient biofuel production, especially in coastal regions.
Similar content being viewed by others
Introduction
An increasing worldwide demand for energy combined with decreasing fossil energy resources not only fosters climate change1 but also explorations for fossil energy in sensitive ecosystems2. A key strategy to mitigate climate change is to substitute fossil by renewable energy sources. Because liquid fuels play a predominant role in the transportation sector, second generation biofuels from lignocellulosic feedstock reveal a high potential in substituting fossil fuels3. Restrictions in the production of ethanol from biomass mainly derive from the plant cell wall’s recalcitrance, which is primarily determined by cellulose crystallinity but also lignin and hemicellulose content4,5.
To improve second generation ethanol production, we tried to identify and increase the content of cell wall polymers that are easily degradable and contain readily fermentable residues. Hence, polymers that only consist of glucose would represent an optimal substrate for ethanol fermentation with efficient microorganisms like yeast (Saccharomyces cerevisiae). Apart from the major (1, 4)-β-glucan cell wall polymer cellulose6, only two other cell wall polymers consist entirely of glucose: (1, 3)-β-glucan, known as callose in plants7 and (1, 3;1, 4)-β-glucan, a mixed-linkage glucan found in plants of the order Poales, in horsetail (Equisetum spp.) and in bryophytes8. (1, 3)-β-glucan is important to maintain the vascular system, for pollen development and cell plate formation in growing tissue as well as for defense responses7. Mixed-linkage glucan can serve as an energy storage and has a growth-related function in vegetative tissues of grasses9. Because of their biological function, the abundance of these β-glucan polymers has been considered low in lignocellulosic biomass10. Therefore, these polymers have not been targeted to improve saccharification. To test whether an undiscovered potential of β-glucan processing to ethanol would exist, we determine the (1, 3)-β-glucan and (1, 3;1, 4)-β-glucan content in lignocellulosic biomass of crops representing a major source of lignocellulosic feedstock from agriculture in temperate climates: barley (Hordeum vulgare), wheat (Triticum aestivum) and maize (Zea mays); as well as in the model plants Arabidopsis (Arabidopsis thaliana) and Brachypodium (Brachypodium distachyon) and the emerging, perennial grass Miscanthus (Miscanthus x giganteus). This low-input energy crop with high biomass yields in temperate climates has been proposed for sustainable lignocellulosic feedstock production11.
Whereas the (1, 3;1, 4)-β-glucan content was relatively low in all tested plant species (0.2–0.5%), the (1, 3)-β-glucan content was exceptionally high in maize and Miscanthus, reaching 2% and 5% of total dry leaf biomass, respectively (Fig. 1a). The specificity of the florescent dye aniline blue for staining (1, 3)-β-glucan rather than (1, 3;1, 4)-β-glucan allowed its usage in assays for (1, 3)-β-glucan quantification and microscopic localization (Supplementary Fig. 1). Confocal laser scanning microscopy (CLSM) revealed an unexpectedly high (1, 3)-β-content in epidermal leaf cells of maize and Miscanthus (Fig. 1b), which we did not observe in the other examined plant species (Supplementary Fig. 2). Moreover, localization microscopy (LM), which we recently established for super-resolution analysis of β-glucan polymer networks in plant cells12, facilitated three-dimensional rendering of (1, 3)-β-glucan and (1, 4)-β-glucan macrofibrils in cell walls (Fig. 1b). This microscopic technique revealed a parallel orientation of β-glucan polymer layers. The generation of videos based on three-dimensional rendered glucan macrofibrils data specifically addressed the visualization of direct interactions between the two cell wall polymers. Whereas in maize, interaction between (1, 3)-β-glucan and (1, 4)-β-glucan was mainly based on direct macrofibrils attachment (Supplementary Video 1), (1, 3)-β-glucan macrofibrils additionally partially surrounded (1, 4)-β-glucan macrofibrils in Miscanthus (Supplementary Video 2). This suggests the establishment of a tight polymer network that we also identified in epidermal leaf cells of Arabidopsis at sites of attempted pathogen penetration12. Our results from maize and Miscanthus provide first evidence that (1, 3)-β-glucan can be a major polymer of unchallenged secondary cell walls outside the vascular tissue. Interestingly, we also found a relatively high resistance of (1, 3)-β-glucan to chemical degradation by diluted trifluoroacetic and sulfuric acid (Supplementary Fig. 3), which supported the idea of an independent cell wall component rather than being part of the hemicellulose fraction10.
Layered cell wall architecture in maize and Miscanthus.
(a) β-Glucan content in senesced leaf biomass. n.d., not detectable. Values represent the mean of three independent biological experiments. Error bars represent ± SE. (b) Confocal laser-scanning microscopy (CLSM) of aniline blue fluorochrome (ABF)-stained (1, 3)-β-glucan (blue channel) and pontamine fast scarlet 4B (S4B)-stained (1, 4)-β-glucan (red channel) cross sections of leaves from maize (Z. mays) amd Miscanthus (M. x giganteus). White arrows indicate epidermal cells with a high (1, 3)-β-glucan content and sites of localization microscopy (LM) application. 3D, surface rendering of β-glucan networks. Scale bars: CLSM, 50 μm; LM ABF/S4B, 5 μm; LM 3D, 2 μm.
Before engineering an improved utilization of (1, 3)-β-glucan-enriched biomass, we developed an equation to estimate the increase in biomass saccharification after optimized (1, 3)-β-glucan hydrolysis:

where A(Px) describes the relative amount of the glucan polymer Px, f (s) the glucose saccharification factor of the biomass B before optimization and of the glucan polymer Px after optimized hydrolysis and f (s)ithe increase in glucose saccharification after optimization. Based on equation (1), we expected an improved saccharification only if f (s)Px > f (s)B.
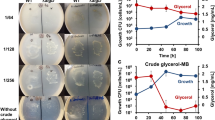
We analyzed the maize leaf biomass broth after dilute-sulfuric acid pretreatment, hydrolysis with the cell wall-degrading enzyme cocktail Accellerase 1500 and subsequent fermentation with a non-adapted, laboratory yeast strain. Here, we detected relative high contents of laminaribiose and –triose (Fig. 2a), which we distinguished from putative glucotrioses deriving from possible (1, 3;1, 4)-β-glucan degradation using a refractive index detector coupled to an HPLC-system (Supplementary Fig. 4). Due to their chemical composition and their relatively small size, we considered these (1, 3)-β-glucan degradation products as a potential, unused glucose source for fermentation. Therefore, we initiated experiments for optimizing (1, 3)-β-glucan hydrolysis and usage of (1, 3)-β-glucan degradation products for ethanol production. In a first step, we changed the yeast strain during fermentation to increase laminaribiose and –triose utilization during fermentation. The application of the yeast strain CEN.PK113-13D (CEN) that has been used to develop strains for optimized biomass fermentation13 significantly improved laminaribiose utilization during fermentation, but did not effected laminaritriose utilization (Fig. 2a). The heterologous expression of the bacterial laminaribiose ABC transporter (LBT) from Clostridium thermocellum14 in the yeast strain CEN (Supplementary Fig. 5a,b) further improved the laminaribiose and laminaritriose utilization (Fig. 2b), resulting in only residual amounts of laminaribiose and –triose in the fermentation broth (Fig. 2a). However, the utilization of laminaritetraose, long-chained (1, 3)-β-glucan, or oligomers deriving from (1, 4)-β-glucan and (1, 3;1, 4)-β-glucan hydrolysis was not facilitated (Supplementary Fig. 5c). Because of the efficient utilization of these two (1, 3)-β-glucan oligomers by CEN+LBT, we considered laminaribiose and –triose as direct contributors to the overall glucose saccharification. Hence, the generation of this yeast strain represented a decisive step in engineering optimized (1, 3)-β-glucan utilization from β-glucan-enriched biomass.
Engineered saccharification and ethanol production of (1, 3)-β-glucan-rich biomass.
(a) Amounts of glucose and the oligosaccharides laminaribiose and –triose remaining in the fermentation broth after 48 h of maize leaf biomass fermentation comparing yeast strains MaV (non-adapted), CEN (adapted) and CEN+LBT (engineered). (b) In vitro growth assays of yeast strains CEN and CEN+LBT on substrates as indicated. (c) Amounts of laminaribiose and –triose (left panel) as well as glucose and the combined amount of these three (1, 3)-β-glucan hydrolysis products (right panel) as indicators for changes in saccharification efficiency of maize and Miscanthus leaf biomass due to additional F. johnsoniae (1, 3)-β-glucanase application (−/+ BGL) after standard biomass pretreatment. (d) Ethanol production after 48 h of fermentation using yeast strains CEN and CEN+LBT and −/+ BGL. Values represent the mean of three independent biological experiments. Letters a, b, c: groups with significant difference, P < 0.05 based on Tukey’s test. Error bars represent ± SE.
To further improve saccharification of (1, 3)-β-glucan-rich biomass, we initially screened 38 (1, 3)-β-glucanases from bacteria, fungi and plants (Supplementary Table 1) that we heterologously expressed and purified from Escherichia coli. Six (1, 3)-β-glucanases showed a higher efficiency in (1, 3)-β-glucan hydrolysis than the Accellerase enzyme cocktail and a commercially available (1, 3)-β-glucanase from Trichoderma reesei, a well-studied and widely used fungus in second generation biofuel production15 (Supplementary Fig. 6a). After confirming expression of these six (1, 3)-β-glucanases in E. coli (Supplementary Fig. 7), we determined their optimal pH and temperature range for enzymatic activity (Supplementary Fig. 8). Under optimal conditions for each enzyme, we identified highest (1, 3)-β-glucanase activity for the enzyme from Flavobacterium johnsoniae (Supplementary Table 1), which was 3.3-times higher than the enzymatic activity of the commercially available (1, 3)-β-glucanase from T. reesei, resulting in an almost 70% hydrolyzing efficiency (Supplementary Fig. 9a). Combining the saccharification efficiency from non-optimized biomass processing (Fig. 2a) and the (1, 3)-β-glucan content of maize and Miscanthus biomass (Fig. 1a), we predicted a 4.0% increase in saccharification efficiency for maize and a 9.3% increase for Miscanthus using equation (1). The experimental data only slightly deviated from our prediction showing a 5.3% increase for maize and a 9.0% increase for Miscanthus (Fig. 2c). The improved second generation ethanol production reflected the efficiency of engineered (1, 3)-β-glucan processing due to i) optimization of its enzymatic hydrolysis and ii) enhanced utilization of its degradation products by the engineered yeast strain CEN+LBT, resulting in an increased ethanol production of 5.7% for maize and 14.4% for Miscanthus (Fig. 2d).
Our results from saccharification and fermentation of maize and Miscanthus biomass revealed a direct correlation between the (1, 3)-β-glucan content of the feedstock and an increased ethanol yield. Hence, we concluded that a further (1, 3)-β-glucan enrichment in biomass would result in increased ethanol yields. To test this hypothesis, we followed two strategies: i) increasing the (1, 3)-β-glucan content in potential feedstock for sustainable biomass production using a biotechnological approach; and ii) identifying new sources of (1, 3)-β-glucan-enriched biomass that could be used in our adapted fermentation process.
To increase the (1, 3)-β-glucan content in feedstock for sustainable biomass production, we overexpressed the GFP-tagged (1, 3)-β-glucan synthase gene PMR4 (POWDERY MILDEW RESISTANT4) from Arabidopsis in Miscanthus (line 35S:PMR4-GFP). PMR4 overexpression in Arabidopsis increased (1, 3)-β-glucan content at infection sites but not in unchallenged tissue16. In contrast, we observed a constitutive increase in (1, 3)-β-glucan content in 35S:PMR4-GFP Miscanthus lines, which was proportional to the relative PMR4 expression level and reached a maximum of 8.5% in leaf tissue (Supplementary Fig. 10a,b). This result suggests different regulatory mechanisms of (1, 3)-β-glucan biosynthesis in Miscanthus and Arabidopsis, which would also explain the strong differences in their overall (1, 3)-β-glucan content (Fig. 1). As expected from Arabidopsis16, PMR4-GFP was localized at the plasma membrane whereas single GFP of a transgenic Miscanthus control line was detectable in cytosolic strands (Supplementary Fig. 10c). We predicted an increase in saccharification efficiency of about 16% in the 35S:PMR4-GFP line with the highest (1, 3)-β-glucan content of 8.5% after optimized hydrolysis, which was relatively close to our experimental results showing a saccharification increase of 14.5%. (Fig. 3b). The improved saccharification of this Miscanthus line resulted in an increase in ethanol production of 20% compared to non-optimized Miscanthus wild-type biomass processing (Fig. 3c). These results revealed a previously undiscovered potential in the layered architecture of maize and Miscanthus leaf cell walls that contain an atypically high content of (1, 3)-β-glucan, which was unleashed by engineering optimized enzymatic hydrolysis and yeast fermentation. Moreover, (1, 3)-β-glucan enrichment represents a new target in breeding energy crops for improved second generation ethanol production.
Enhanced saccharification and ethanol production in (1, 3)-β-glucan-enriched Miscanthus and marine biomass.
(a) (1, 3)-β-glucan content in engineered Miscanthus leaf (35S:PMR4-GFP) and bladderwrack (F. vesiculosus) biomass. (b) Amounts of the oligosaccharides laminaribiose and –triose (left panel) as well as glucose and the combined amount of these three (1, 3)-β-glucan hydrolysis products (right panel) as indicators for changes in saccharification efficiency of Miscanthus wild-type and engineered leaf biomass due to additional F. johnsoniae (1, 3)-β-glucanase application (−/+ BGL) after standard biomass pretreatment. (c) Ethanol production after 48 h of fermentation using yeast strains CEN (adapted) and CEN+LBT (engineered) and −/+ BGL. (d) Thallus morphology of bladderwrack. Photo courtesy of Christian A. Voigt. (e) Micrograph showing cross section of the bladderwrack’s blade after aniline blue fluorochrome-staining of (1, 3)-β-glucan. Micrograph taken by confocal laser-scanning microscopy. Scale bar, 100 μm. (f) Magnification of the blade’s epidermal and cortex cells as indicated in (e). Scale bar = 20 μm. (g) Amounts of laminaribiose, –triose,glucose and the combined amount of these three (1, 3)-β-glucan hydrolysis products as indicators for changes in saccharification efficiency of bladderwrack biomass −/+ BGL. (h) Ethanol production after 48 h of fermentation using the yeast strains CEN and CEN+LBT and −/+ BGL. Values represent the mean of three independent biological experiments. Letters a, b, c: groups with significant difference, P < 0.05 based on Tukey’s test. Error bars represent ± SE.
In our second approach to identify new sources of (1, 3)-β-glucan-enriched biomass, we considered brown macroalgae with a naturally high (1, 3)-β-glucan content as a putative source. To examine the potential of (1, 3)-β-glucan utilization for ethanol production from this marine feedstock, we collected thalli of the brown macroalgae bladderwrack (Fucus vesiculosus) from the German Baltic Sea shore at Eckernförde near Hamburg (Fig. 3d). We determined an astonishing (1, 3)-β-glucan content of 15.3% (Fig. 3a) confirming previous studies of this macroalga17. CLSM revealed (1, 3)-β-glucan deposition in all tissues of the blade (Fig. 3e). However, highest (1, 3)-β-glucan accumulation occurred within epidermal and cortex cells (Fig. 3f). The generation of three-dimensional videos from (1, 3)-β-glucan accumulation within the bladderwrack tissue allowed us to distinguish between different deposition patterns. Whereas elongated cells of the central pith region revealed a scattered (1, 3)-β-glucan deposition pattern (Supplementary Video 3), a relatively compact layer of (1, 3)-β-glucan was intracellular deposited in cortex and especially epidermal cells (Supplementary Video 4). Based on our previous results, we hypothesized that the optimized processing of (1, 3)-β-glucan-enriched bladderwrack biomass would result in increased ethanol production. We first tested the six (1, 3)-β-glucanases with highest activity on brown algae-derived (1, 3)-β-glucan as substrate. Similar to our previous test, the enzyme from F. johnsoniae showed highest activity with a hydrolyzing efficiency of 37.5% (Supplementary Fig. 9b). Using this (1, 3)-β-glucanase in addition to the Accellerase enzyme cocktail for biomass hydrolysis after dilute-sulfuric acid pretreatment, we increased bladderwrack biomass saccharification by 45.6% (Fig. 3g), which was close to the prediction of 46.4% using equation (1). Consequently, the ethanol yield was 50.5% higher in optimized CEN+LBT-driven fermentation compared to non-optimized (1, 3)-β-glucan processing (Fig. 3h).
Similar to Miscanthus, brown macroalgae have been considered for sustainable biomass production18, however, without competing for arable land and food production. Because we demonstrated optimized ethanol production for both, (1, 3)-β-glucan-enriched plant and marine feedstock, we proposed our engineered biomass processing for co-fermentation of Miscanthus and bladderwrack biomass (Fig. 4a). A successful co-fermentation using equal amounts of Miscanthus and bladderwrack biomass proved the applicability of this engineered production approach (Fig. 4b). Regarding saccharification of mixed Miscanthus and bladderwrack biomass, we identified enzymatic hydrolysis as a field of further improvement. Here, the release of laminaribiose from (1, 3)-β-glucan was specifically inhibited during mixed Miscanthus and bladderwrack biomass processing compared to single biomass processing whereas inhibition did not occur for laminaritriose or glucose release in the mixed biomass approach (Fig. 4b).
Co-fermentation of (1, 3)-β-glucan-enriched terrestrial and marine biomass.
(a) Schematic overview of parallel biomass processing. Photos courtesy of Christian A. Voigt. (b) Saccharification efficiency and ethanol production under optimized co-processing of Miscanthus and bladderwrack biomass. Values represent the mean of two independent biological experiments. Letters a, b, c: groups with significant difference, P < 0.05 based on Tukey’s test. Error bars represent ± SE.
Because brown macroalgae do not contain lignin and (1, 3)-β-glucan substantially contributed to improve second generation ethanol production in our study, we considered the produced ethanol as glucanocellulosic.
The effective co-fermentation opens new perspectives for sustainable and efficient ethanol production in bio-refineries, especially in coastal regions that combine the potential of offshore macroalgae aquaculture and proximate energy crop cultivation. An example for a coastal region that would fulfill these prerequisites is Schleswig-Holstein in northern Germany. Existing and planned offshore wind parks in the North and Baltic Sea would facilitate effective macroalgae aquacultures19 and a high potential for Miscanthus cultivation in Schleswig-Holstein was shown in our recent study20. Hence, these coastal regions would represent prototypic sites for future biorefineries for plant and marine biomass co-fermentation, combining short delivery distances of feedstock with a high abundance of renewable electricity for processing, which could help to promote large-scale energy transition projects like the ambitious German Energiewende21.
Methods
Biological material
Brachypodium (Brachypodium distachyon, inbred line Bd 2122), barley (Hordeum vulgare, cultivar Golden Promise23), wheat (Triticum aestivum, cultivar Nandu, Lochow-Petkus, Bergen-Wohlde, Germany), maize (Zea mays, inbred line A18824) and Miscanthus (Miscanthus x giganteus) were cultivated as described in Meineke et al.25. Arabidopsis (Arabidopsis thaliana, wild-type Columbia) was cultivated as described in Stein et al.26. Naturally dried leaf material was harvested manually at its final developmental stage after senescence and 2 additional weeks of drying25. Biomass was additionally dried at 50 °C for 2 days in a drying oven. Washed ashore thalli of the brown macroalga bladderwrack (Fucus vesiculosus) were collected in November from the Baltic Sea shore at Eckernförde (Schleswig-Holstein, Germany, geographical position: 54 °27'57.5″N 9°50'28.0″E) and dried at 50 °C for 3 days. Plant and alga biomass was homogenized with a mill fitted with a 0.5 mm mesh screen prior processing. Material subject to (1, 3)-β-glucan extraction was ground in liquid nitrogen using a mortar and pestle.
(1, 3)-β-Glucan extraction and determination
20 mg of mortared and lyophilized leaf or alga biomass was destained in ethanol (96%) at 50 °C and 600 rpm for 10 min. Subsequent procedures of (1, 3)-β-glucan determination followed the description in Voigt et al.27. Ethanol was removed after centrifugation (2 min, 10,000 g) and the sample was dried using a centrifugal evaporator. After a washing step with H2O, the sample was dried again in a centrifugal evaporator. For (1, 3)-β-glucan extraction, the sample was resuspended in 400 μl of 1 M NaOH and incubated at 80 °C and 600 rpm for 1 h. After centrifugation (10 min, 2000 g), the supernatant was used for the aniline blue fluorescence assay for (1, 3)-β-glucan determination. 5 μl sample were mixed with 45 ml H2O, 5 μl HCl (1 M), 220 μl K2HPO4 (150 mM) and 5 μl aniline blue fluorochrome (ABF, 0.1 mg·ml−1 in H2O, Biosupplies, Australia). Standards ranging from 0 to 20 μg ml−1 were generated from purified (1, 3)-β-glucan from Euglena gracilis (Sigma-Aldrich, Germany) in the same way as described for plant and alga samples. Additional standards were generated accordingly from oat and barley (1, 3;1, 4)-β-glucan deriving from the mixed-linkage beta-glucan kit (Megazyme, Ireland) to verify the specificity of ABF in staining (1, 3)-β-glucan. Fluorescence measurement was performed in 96-well plates with the microplate reader Synergy HT (BioTek, USA; absorbance filter: 380/20 nm, emission filter: 460/40 nm).
(1, 3;1, 4)-β-Glucan determination
Mixed linkage (1, 3;1, 4)-β-glucan in plant biomass was determined according to the manufacturer’s description of the mixed-linkage beta-glucan kit (Megazyme). An additional 5 ml H2O was added to the samples (750 mg milled leaf biomass) in the initial incubation step in a water bath (100°C). Calculation of the (1, 3;1, 4)-β-glucan content followed the instructions of the manufacturer’s manual.
Confocal laser-scanning microscopy
The confocal laser-scanning microscope Zeiss LSM 780 (Carl Zeiss MicroImaging GmbH, Germany) was used for the localization of pontamine fast scarlet 4B (S4B, Sigma-Aldrich)-stained (1, 4)-β-glucan in leaf samples and ABF-stained (1, 3)-β-glucan in leaf and bladderwrack samples. To verify the specificity of ABF in staining (1, 3)-β-glucan, (1, 3)-β-glucan from E. gracilis (Sigma-Aldrich) as well as (1, 3;1, 4)-β-glucan from oat and barley (Megazyme) was suspended in H2O and stained with ABF and used in CLSM. In addition, the green fluorescence protein (GFP)-tagged (1, 3)-β-glucan synthase PMR4 and single GFP in leaves of transformed Miscanthus lines was localized by CLSM. For cross sections of leaves, samples were embedded in 9% Agarose and sections were made with a vibratome (Carl Zeiss MicroImaging). The setup for CLSM analysis of stained cell wall polymers and GFP followed the description in Ellinger et al.16 and Eggert et al.12.
Super-resolution microscopy
A custom modified Nikon stochastic optical reconstruction microscope (N-STORM, Nikon GmbH, Germany) was used to analyze the (1, 3)-β-glucan/(1, 4)-β-glucan polymer network of ABF and S4B stained maize and Miscanthus cross sections. The microscopic setup and image reconstruction was done according to Eggert et al.12.
Treatment and fermentation of plant biomass
For pretreatment, 5 g of milled leaf biomass was mixed with 43 ml sulfuric acid (1.75% (v/v)) and autoclaved for 15 min at 120 °C. Subsequent procedures of enzymatic hydrolysis and fermentation followed the description in Meineke et al.25. To test the impact of the (1, 3)-β-glucanase from F. johnsoniae, 500 ng of the purified enzyme were additionally added to fermentation reactors and incubated for 24 h and 200 rpm at 37 °C. Fermentations were initiated with the inoculation of 2 ml of overnight yeast cultures of the non-adapted, laboratory strain MaV203 (MaV, Life Technologies), CEN, or CEN+LBT (generated in this study). Amounts of glucose, laminaribiose, laminaritriose and ethanol in fermentation supernatants were quantified with a refractive index detector on an ICS-5000 system (Dionex, USA) with a HPX 87H column (Bio-Rad, USA, mobile phase 0.005 M H2SO4, flow rate 0.6 ml·min−1, column temperature: 50 °C, refractive index detector) as described in Meineke et al.25. In addition to laminaribiose and laminaritriose, the trisaccharides glucotriose (I) (β-D-Glc-(1 → 3)-β-D-Glc-(1 → 4)-D-Glc), glucotriose (II) (β-D-Glc-(1 → 4)-β-D-Glc-(1 → 3)-D-Glc) were used as standards (all standard oligosaccharides from Megazyme) to distinguish between possible degradation products from (1, 3)-β-glucan and (1, 3;1, 4)-β-glucan.
Cloning and Miscanthus transformation
We generated two vector constructs for the transformation of Miscanthus: i) overexpression of the callose synthase gene PMR4 from Arabidopsis (At4g03550) fused to GFP and ii) overexpression of single GFP, both under control of the 35S promoter. PMR4-GFP and GFP were amplified from the vector pCAMBIA-35S:PMR4-GFP16 using primers in PCR reactions that provide DNA recombination sequences (attB sites) at their 5′ and 3′ ends (PMR4-5′attB: 5′-GGGGACAAGTTTGTACAAAAAAGCAGGCTATGAGCCTCCGCCACCGC, GFP-5′attB: 5′-GGGGACAAGTTTGTACAAAAAAGCAGGCTTGGAGATC CAAACAATGAGTAAAG, GFP-3′attB: 5′-GGGGACCACTTTGTACAAGAAAGCTGGG TTAAGCTTGAATTCTTATT TGTATA) for utilization with the Gateway cloning technology. PMR4-GFP and GFP were introduced into the plant expression vector pIPKb00428, which provided 35S promoter-driven gene expression. Generated vector constructs containing 35S:PMR4-GFP and 35S:GFP expression cassettes were transformed into Agrobacterium (Agrobacterium tumefaciens, strain GV3101). The generation of transgenic Miscanthus lines followed the principal procedure of Agrobacterium-mediated callus transformation and selection on hygromycin-containing plant cell culture medium. Resistance to hygromycin was provided by the used plant expression vector pIBKb004. A detailed description of the transformation procedure is provided in the Supplementary Information.
Statistical analysis
Descriptive statistics including the mean and the standard error of the mean (SE) along with the Tukey range test for multiple comparison procedures in conjunction with an ANOVA were used to determine significant differences. P < 0.05 was considered significant.
Additional Information
How to cite this article: Falter, C. et al. Glucanocellulosic ethanol: the undiscovered biofuel potential in energy crops and marine biomass. Sci. Rep. 5, 13722; doi: 10.1038/srep13722 (2015).
References
Ash, C. et al. Natural systems in changing climates. Once and future climate change. Introduction. Science 341, 472–473 (2013).
Finer, M., Jenkins, C. N., Pimm, S. L., Keane, B. & Ross, C. Oil and gas projects in the Western Amazon: threats to wilderness, biodiversity and indigenous peoples. Plos One 3, e2932, 10.1371/journal.pone.0002932 (2008).
Sims, R. E., Mabee, W., Saddler, J. N. & Taylor, M. An overview of second generation biofuel technologies. Bioresour. Technol. 101, 1570–1580 (2010).
Yoshida, M. et al. Effects of cellulose crystallinity, hemicellulose and lignin on the enzymatic hydrolysis of Miscanthus sinensis to monosaccharides. Biosci. Biotechnol. Biochem. 72, 805–810 (2008).
Hall, M., Bansal, P., Lee, J. H., Realff, M. J. & Bommarius, A. S. Cellulose crystallinity--a key predictor of the enzymatic hydrolysis rate. FEBS J. 277, 1571–1582 (2010).
Updegraff, D. M. Semimicro determination of cellulose in biological materials. Analyt. Biochem. 32, 420–424 (1969).
Bacic, A., Fincher, G. B. & Stone, B. A. Chemistry, Biochemistry and Biology of (1->3)-β-Glucans and Related Polysaccharides. (Academic Press, 2009).
Burton, R. A. & Fincher, G. B. (1, 3;1, 4)-β-D-glucans in cell walls of the poaceae, lower plants and fungi: a tale of two linkages. Mol. Plant 2, 873–882 (2009).
Buckeridge, M., Rayon, C., Urbanowicz, B., Tine, M. & Carpita, N. Mixed linkage (1→3)(1→4)-β-D-glucans of grasses Cereal Chem. 81, 115–127 (2004).
Scheller, H. V. & Ulvskov, P. Hemicelluloses. Ann. Rev. Plant Biol. 61, 263–289 (2010).
Heaton, E., Dohleman, F. & Long, S. Meeting US biofuel goals with less land: the potential of Miscanthus. Glob. Change Biol. 14, 2000–2014 (2008).
Eggert, D., Naumann, M., Reimer, R. & Voigt, C. A. Nanoscale glucan polymer network causes pathogen resistance. Sci. Rep. 4, 4159, 10.1038/srep04159 (2014).
Wilde, C. et al. Expression of a library of fungal beta-glucosidases in Saccharomyces cerevisiae for the development of a biomass fermenting strain. Appl. Microbiol. Biotechnol. 95, 647–659 (2012).
Nataf, Y. et al. Cellodextrin and laminaribiose ABC transporters in Clostridium thermocellum. J. Bacteriol. 191, 203–209 (2009).
Schuster, A. & Schmoll, M. Biology and biotechnology of Trichoderma. Appl. Microbiol. Biotechnol. 87, 787–799 (2010).
Ellinger, D. et al. Elevated early callose deposition results in complete penetration resistance to powdery mildew in Arabidopsis. Plant Physiol. 161, 1433–1444 (2013).
Mabeau, S. & Kloareg, B. Isolation and analysis of the cell walls of brown algae: Fucus spiralis, F. ceranoides, F. vesiculosus, F. serratus, Bifurcaria bifurcata and Laminaria digitata. J. Exp. Bot. 38, 1573–1580 (1987).
Roesijadi, G., Jones, S. B., Snowden-Swan, L. J. & Zhu, Y. Macroalgae as a Biomass Feedstock: A Preliminary Analysis. Report No. prepared for the U.S. Department of Energy under contract DE-AC05-76RL01830, (Pacific Northwest National Laboratory, 2010).
Buck, B. H. & Buchholz, C. The Offshore-Ring: A new system design for the open ocean aquaculture of macroalgae. J. Appl. Phycol. 16, 355–368 (2004).
Schorling, M., Enders, C. & Voigt, C. A. Assessing the cultivation potential of the energy crop Miscanthus × giganteus for Germany. GCB Bioenergy, 10.1111/gcbb.12170 (2014).
Schiermeier, Q. Renewable power: Germany’s energy gamble. Nature 496, 156–158 (2013).
Vogel, J. P. et al. EST sequencing and phylogenetic analysis of the model grass Brachypodium distachyon. Theoret. Appl. Genet. 113, 186–195 (2006).
Sigburbjorsson, B. & Micke, A. Progress in mutation breeding. In: Induced Mutation in Plants. Proceedings of an International Symposium on Nature, Induction and Utilization of Mutation in Plants, Vienna, 673–697. Interantional Atomic Energy Agency (1969).
Green, C. E. & Philips, R. L. Plant regeneration from tissue cultures of maize. Crop Sci. 15, 417–421 (1975).
Meineke, T., Manisseri, C. & Voigt, C. A. Phylogeny in defining model plants for lignocellulosic ethanol production: A comparative study of Brachypodium distachyon, wheat, maize and Miscanthus x giganteus leaf and stem biomass. Plos One 9, e103580, 10.1371/journal.pone.0103580 (2014).
Stein, M. et al. Arabidopsis PEN3/PDR8, an ATP binding cassette transporter, contributes to nonhost resistance to inappropriate pathogens that enter by direct penetration. Plant Cell 18, 731–746 (2006).
Voigt, C. A., Schäfer, W. & Salomon, S. A comprehensive view on organ-specific callose synthesis in wheat (Triticum aestivum L.): glucan synthase-like gene expression, callose synthase activity, callose quantification and deposition. Plant Physiol Biochem 44, 242–247 (2006).
Himmelbach, A. et al. A set of modular binary vectors for transformation of cereals. Plant Physiol. 145, 1192–1200 (2007).
Acknowledgements
We thank E. Boles from the Goethe University Frankfurt for providing the yeast strain CEN.PK113-13D. This work was supported by a BMBF grant to C.A.V awarded to the Biocenter Klein Flottbek, University of Hamburg (FKZ0315521A).
Author information
Authors and Affiliations
Contributions
C.A.V. conceived the project rationale and wrote the manuscript. C.F. performed pretreatment, hydrolysis and fermentation of plant and marine biomass as well as yeast strain analyses together with C.Z. C.Z. performed glucanase screening and enzyme purification. A.B. generated transgenic Miscanthus lines together with K.W. D.El. performed HPLC analyses. D.Eg. performed super-resolution localization microscopy together with R.R. M.N. performed confocal laser-scanning microscopy together with C.A.V. and C.F.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Falter, C., Zwikowics, C., Eggert, D. et al. Glucanocellulosic ethanol: the undiscovered biofuel potential in energy crops and marine biomass. Sci Rep 5, 13722 (2015). https://doi.org/10.1038/srep13722
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep13722
This article is cited by
-
Conversion of cassava peels into bioethanol using the OSTEP approach
Biomass Conversion and Biorefinery (2022)
-
Production and immobilization of β-glucanase from Aspergillus niger with its applications in bioethanol production and biocontrol of phytopathogenic fungi
Scientific Reports (2021)
-
Dynamics of structural polysaccharides deposition on the plasma-membrane surface of plant protoplasts during cell wall regeneration
Journal of Wood Science (2019)
-
A Chrysoporthe cubensis enzyme cocktail produced from a low-cost carbon source with high biomass hydrolysis efficiency
Scientific Reports (2017)
-
The effect of mechanical pre-processing and different drying methodologies on bioethanol production using the brown macroalga Laminaria digitata (Hudson) JV Lamouroux
Journal of Applied Phycology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.