Abstract
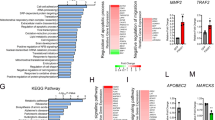
The superfamily of small GTP-binding proteins has expanded dramatically in recent years. The Ras family has long been associated with signaling pathways contributing to normal and aberrant cell growth, while Rho-related protein function is to integrate extracellular signals with specific targets regulating cell morphology, cell aggregation, tissue polarity, cell motility and cytokinesis. Recent findings suggest that certain Rho proteins, including RhoA, Rac1 and Cdc42, can also play a role in signal transduction to the nucleus and cell growth control. However, the nature of the genes regulated by Ras and Rho GTPases, as well as their contribution to their numerous biological effects is still largely unknown. To approach these questions, we investigated the global gene expression pattern induced by activated forms of H-Ras, RhoA, Rac1 and Cdc42 using cDNA microarrays comprising 19 117 unique elements. Using this approach, we identified 1184 genes that were up- or downregulated by at least twofold. Hierarchical cluster analysis revealed the existence of patterns of gene regulation both unique and common to H-Ras V12, RhoA QL, Rac1 QL and Cdc42 QL activation. For example, H-Ras V12 upregulated osteopontin and Akt 1, and H-Ras and RhoA stimulated cyclin G1, cyclin-dependent kinase 8, cyclin A2 and HMGI-C, while Rac1 QL and Cdc42 QL upregulated extracellular matrix and cell adhesion proteins such as alpha-actinin 4, procollagen type I and V and neuropilin. Furthermore, H-Ras V12 downregulated by >eightfold 52 genes compared to only three genes by RhoA QL, Rac1 QL and Cdc42 QL. These results provide key information to begin unraveling the complexity of the molecular mechanisms underlying the transforming potential of Ras and Rho proteins, as well as the numerous morphological and cell cycle effects induced by these small GTPases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bar-Sagi D and Hall A . (2000). Cell, 103, 227–238.
Bos JL . (1989). Cancer Res., 49, 4682–4689.
Brenner KA, Corbett SA and Schwarzbauer JE . (2000). Oncogene, 19, 3156–3163.
Chen H, He Z and Tessier-Lavigne M . (1998). Nat. Neurosci., 1, 436–439.
Coso OA, Chiariello M, Yu JC, Teramoto H, Crespo P, Xu N, Miki T and Gutkind JS . (1995). Cell, 81, 1137–1146.
Fedele M, Berlingieri MT, Scala S, Chiariotti L, Viglietto G, Rippel V, Bullerdiek J, Santoro M and Fusco A . (1998). Oncogene, 17, 413–418.
Fukuhara S, Chikumi H and Gutkind JS . (2001). Oncogene, 20, 1661–1668.
Hall A . (1998). Science, 279, 509–514.
Hill CS, Wynne J and Treisman R . (1995). Cell, 81, 1159–1170.
Liu C, Yao J, Mercola D and Adamson E . (2000). J. Biol. Chem., 275, 20315–20323.
Macara IG, Lounsbury KM, Richards SA, McKiernan C and Bar-Sagi D . (1996). FASEB J., 10, 625–630.
Malek RL, Irby RB, Guo QM, Lee K, Wong S, He M, Tsai J, Frank B, Quackenbush J, Jove R, Yeatman TJ and Lee NH . (2002). Oncogene, 21, 7256–7265.
Michael D and Oren M . (2002). Curr. Opin. Genet. Dev., 12, 53–59.
Miyamoto S, Teramoto H, Coso OA, Gutkind JS, Burbelo PD, Akiyama SK and Yamada KM . (1995). J. Cell Biol., 131, 791–805.
Narumiya S . (1996). J. Biochem. (Tokyo), 120, 215–228.
Nobes CD and Hall A . (1995). Cell, 81, 53–62.
Perona R, Esteve P, Jimenez B, Ballestero RP, Ramon Y, Cajal S and Lacal JC . (1993). Oncogene, 8, 1285–1292.
Porter AC and Vaillancourt RR . (1998). Oncogene, 17, 1343–1352.
Soker S, Takashima S, Miao HQ, Neufeld G and Klagsbrun M . (1998). Cell, 92, 735–745.
Takai Y, Sasaki T and Matozaki T . (2001). Physiol. Rev., 81, 153–208.
Tanaka TS, Jaradat SA, Lim MK, Kargul GJ, Wang X, Grahovac MJ, Pantano S, Sano Y, Piao Y, Nagaraja R, Doi H, Wood III WH, Becker KG and Ko MS . (2000). Proc. Natl. Acad. Sci. USA, 97, 9127–9132.
Teramoto H, Crespo P, Coso OA, Igishi T, Xu N and Gutkind JS . (1996). J. Biol. Chem., 271, 25731–25734.
van Leeuwen FN, van der Kammen RA, Habets GG and Collard JG . (1995). Oncogene, 11, 2215–2221.
Vara Prasad MV and Dhanasekaran N . (1999). Oncogene, 18, 1639–1642
Yang IV, Chen E, Hasseman JP, Liang W, Frank BC, Wang S, Sharov V, Saeed AI, White J, Li J, Lee NH, Yeatman TJ and Quackenbush J . (2002). Genome Biol., 3, research 0062.
Zhang J, Takahashi K, Takahashi F, Shimizu K, Ohshita F, Kameda Y, Maeda K, Nishio K and Fukuchi Y . (2001). Cancer Lett., 171, 215–222.
Zentner MD, Lin HH, Deng HT, Kim KJ, Shih HM and Ann DK . (2001). J. Biol. Chem., 276, 29805–29815.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is a ‘United States Government Work’ paper as defined by the US Copyright Act.
Rights and permissions
About this article
Cite this article
Teramoto, H., Malek, R., Behbahani, B. et al. Identification of H-Ras, RhoA, Rac1 and Cdc42 responsive genes. Oncogene 22, 2689–2697 (2003). https://doi.org/10.1038/sj.onc.1206364
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206364
Keywords
This article is cited by
-
Akt-mediated Ephexin1–Ras interaction promotes oncogenic Ras signaling and colorectal and lung cancer cell proliferation
Cell Death & Disease (2021)
-
TLX1 and NOTCH coregulate transcription in T cell acute lymphoblastic leukemia cells
Molecular Cancer (2010)
-
Transcriptomal profiling of the cellular transformation induced by Rho subfamily GTPases
Oncogene (2007)
-
Hypoxia upregulates osteopontin expression in NIH-3T3 cells via a Ras-activated enhancer
Oncogene (2005)
-
Autocrine activation of an osteopontin-CD44-Rac pathway enhances invasion and transformation by H-RasV12
Oncogene (2005)