Abstract
Natural populations of diploid Arabidopsis lyrata exhibit the sporophytic type of self-incompatibility system characteristic of Brassicaceae, in which complicated dominance interactions among alleles in the diploid parent determine self-recognition phenotypes of both pollen and stigma. The purpose of this study was to investigate how polyploidy affects this already complex system. One tetraploid population (Arabidopsis lyrata ssp kawasakiana from Japan) showed complete self-compatibility and produced viable selfed progeny for at least three generations subsequent to field collection. In contrast, individuals from a second tetraploid population (A. lyrata ssp petraea from Austria) were strongly self-incompatible (SI). Segregation of SI genotypes in this population followed Mendelian patterns based on a tetrasomic model of inheritance, with two to four alleles per individual, independent segregation of alleles, and little evidence of dosage effects of alleles found in multiple copies. Similar to results from diploids, anomalous compatibility patterns involving particular combinations of individuals occurred at a low frequency in the tetraploids, suggesting altered dominance in certain genetic backgrounds that could be due to the influence of a modifier locus. Overall, dominance relationships among S-alleles in self-incompatible tetraploid families were remarkably similar to those in related diploids, suggesting that this very important and complicated locus has not undergone extensive modification subsequent to polyploidization.
Similar content being viewed by others
Introduction
Genetically controlled self-incompatibility (SI) systems in flowering plants promote outcrossing by preventing self-pollination, or cross-pollination among individuals sharing expressed alleles. Such systems have attracted the attention of both molecular biologists interested in the biochemistry of cell–cell recognition signaling pathways and population geneticists interested in the dynamics of multiallelic systems maintained by the forces of balancing selection (eg Charlesworth et al, 2000). These tend to be highly complex systems, with individual male and female recognition components physically linked in multi-gene regions characterized by low recombination, which allows maintenance of the same specificity in male and female genes. These regions are often referred to as S-loci, with variants referred to as S-alleles or S-haplotypes. In one of the best-characterized systems, gametophytic SI (GSI: where the recognition phenotype of the pollen is determined by its own haploid genotype), polyploidization has been reported to disrupt the process, via competition among alleles in diploid pollen (eg, Lewis, 1947; Chawla et al, 1997; Stone, 2002). This is apparently a genotype-dependent effect in at least some species (reviewed in Mable, 2004b), but there are a relatively large number of tetraploid GSI species with increased potential for self-compatibility compared to their diploid relatives (Miller and Venable, 2000). Recent evidence from tetraploid sour cherries suggests that there may be more flexibility in such systems than has previously been assumed, with segregation for self-compatible and self-incompatible individuals in some cultivars resulting from the presence of both functional and nonfunctional S-alleles that may be due to alterations in either male or female genes (Hauck et al, 2002; Yamane et al, 2003).
Sporophytic self-incompatibility (SSI) is even more complex: the self-recognition phenotype of both pollen and stigmas is determined by the diploid genotype of the parent, so that dominance interactions among alleles in both male and female components (the genes SCR and SRK, respectively) determine recognition phenotypes (eg reviewed by Nasrallah, 2002; Hiscock and Tabah, 2003). In diploids, SI alleles show very complex dominance relationships, with nonlinear dominance hierarchies, partial expression of SI, differential dominance in pollen and stigmas, genotype-dependent deviations from expected compatibility patterns, and distinctive classes of alleles showing particular dominance patterns (Bateman, 1952; Thompson and Taylor, 1966, 1971; Sampson, 1967; Stevens and Kay, 1989; Kowyama et al, 1994; Mehlenbacher, 1997; Hatakeyama et al, 1998; Schierup, 1998; Brennan et al, 2002; Mable et al, 2003). It seems unlikely that competition among S-alleles in tetraploids with SSI would result in the breakdown observed in some species with GSI. In fact, a loss of SI is not seen as frequently with SSI (reviewed in Mable, 2004b), with some tetraploids showing no such breakdown (eg Ranunculus repens, Ranunculaceae: Lundqvist, 1998; Rutidosis leptorrhynchoides, Asteraceae: Young et al, 2000) and others showing variation in outcrossing rates by accession or variant (Senecio vulgaris, Asteraceae: Abbott et al, 1998; Brassica napus, Brassicaceae: Brugière et al, 2000). However, the population genetics and molecular control of SSI have recently been characterized for diploids of Arabidopsis lyrata in the family Brassicaceae (Charlesworth et al, 2000, 2003a, 2003b; Kusaba et al, 2001, 2002; Schierup et al, 2001; Nasrallah et al, 2002; Mable et al, 2003). Dominance interactions among S-alleles (i.e. variants in male and female genes) in this species are known to be complex, but have been resolved sufficiently to allow predictions about interactions among alleles falling into different classes (Nasrallah et al, 2000; Kusaba et al, 2001, 2002; Charlesworth et al, 2003a; Mable et al, 2003; Bechsgaard et al, 2004). The species has been promoted as a model because it has been thought to be a diploid and obligately outcrossing relative of the model species Arabidopsis thaliana (eg Nasrallah, 2000; Mitchell-Olds, 2001), but this is an oversimplification. Polyploidy has been described in populations from three of the four geographically recognized subspecies (Al-Shehbaz et al, 1999; Koch et al, 1999): A. lyrata petraea from Europe (Polatschek, 1966), A. lyrata kawasakiana from Asia (Miyashita et al, 1998), and A. lyrata kamchatica from Asia and Western North America (Rollins, 1993). Preliminary data suggest variation in the strength of SI among populations of tetraploids from two different regions (Europe and Japan), but also among populations of diploids from North America (Mable, 2004a). This species group therefore makes an excellent model to study how polyploidy influences the expression of SI, and factors that contribute to a loss of SI in both diploids and tetraploids.
Study of the breakdown of SI can provide valuable insights into the mechanisms controlling functional SI systems, as has been described for self-compatible Brassica napus (Goring et al, 1993; Robert et al, 1994; Yu et al, 1996; Ciu et al, 1999), and may provide clues to how SI evolved in these groups. Study of dominance interactions among alleles in tetraploids with functional SI systems has the added benefit of combining multiple allelic classes in the same individual to test hypotheses about intransitiveness (ie nonlinearity) of allelic interactions. Whereas previous studies have described the inheritance of SI in tetraploids with sporophytic control (Lundqvist, 1998; Young et al, 2000), there has been no detailed study of dominance interactions among alleles in natural populations, which is necessary before the number and frequency of alleles in tetraploid populations can be studied and compared to those in diploid populations.
The purpose of this study was to ask: (1) How does polyploidy affect the strength of SI in tetraploids derived from diploids with sporophytic control of SI? and (2) How does polyploidy affect segregation and dominance interactions among alleles at the locus controlling SI?
Materials and methods
Samples
Data are presented for two populations of tetraploids: (1) a self-compatible population from Japan (A. lyrata ssp kawasakiana: collected from Lake Biwa by N Miyashita) and (2) a self-incompatible population from Austria (A. lyrata ssp petraea: collected from Mödling and Aggsbach by S Ansell). For both populations, multiple open-pollinated fruits were collected from each maternal plant (seven from Japan and seven from Austria); field-collected seed sets thus include a mixture of full-sib and half-sib families. Segregation analyses were performed on progeny from crosses between plants from different maternal seed sets. To test the inheritance of SI in the Austrian population, crosses were performed between plants that were strongly SI and those that showed higher levels of self-compatible. Plants were grown in the greenhouse for a minimum of 12 h (normally 16) with a preferred temperature of 18°C day and 17°C nights, although higher temperatures were reached in summer months. Seeds were germinated without vernalization by sowing directly into a 3:1 mixture of soil:clay (Promix: Turface).
DNA content
Ploidy level of all plants was assessed using flow cytometry, and confirmed by chromosome counts as described in detail in Dart et al (2004). Flow cytometry was performed according to a protocol modified from Bino et al (1992), and Arumuganathan and Earle (1991) on a BD Biosciences FACSCalibur flow cytometer (BD Biosciences, San Jose, CA, USA), using three to five young leaves per plant (approx. 15–20 mm in length, resulting in approximately 70 mg of tissue) stained with propidium iodide. Data were acquired and analyzed using CellQuest and CellQuestPro software (Becton Dickinson and Co., 1996). The lowest peak in each sample was used to estimate DNA content, as these plants have the high degree of endopolyploidy common to other Brassicaceae (Barow and Meister, 2003). To confirm chromosome numbers, staining of both meiotic and mitotic tissue from whole buds (described in Dart et al, 2004) was carried out following a modified protocol by Ross et al (1996), based on a previous protocol by Henderson and Lu (1968).
Pollinations
For both tetraploid populations, progeny from seeds raised from field-collected fruits were selfed by rubbing stigmas with anthers from the same plant (six replicates per plant) to assess strength of SI. Outcross pollinations (ie between individuals from different maternal seed sets) were performed opportunistically as plants flowered to ensure fertility. Cross-pollinations were performed by the following method. As plants began to flower, open flowers were removed, and the plants were placed under bridal veil covers for a minimum of 3 days prior to pollination. Pollinations were performed by rubbing anthers on stigmas of emasculated flowers. Each combination of parents was pollinated reciprocally, using three replicate flowers whenever possible. Compatibility was scored by fruit set 7–10 days after pollination according to the criteria described in Mable et al (2003). Rather than scoring pollinations by fluorescent staining of pollen tube growth, fruit set was monitored in order to produce second-generation progeny to test dominance relationships. Progeny from first-generation crosses were raised to flowering and used for segregation analyses based on a complete diallele pollination design within each family tested. Each individual was also selfed to assess the strength of SI. Progeny were sorted into incompatibility groups based on fruit set as described, in Mable et al (2003) to infer phenotype segregation of alleles at the S-locus.
S-allele genotyping
Genotypes at the SRK locus (referred to as Aly13 in previous papers: Schierup et al, 2001; Charlesworth et al, 2003a, 2003b) were determined according to Mable et al (2003). In brief, two approaches were used to determine SRK genotypes: (1) screening with a set of allele-specific primers previously designed for diploid populations, and (2) cloning and screening PCR products amplified using general primers (situated in the extracellular or S-domain) designed to amplify a large number of SRK alleles in A. lyrata. The full set of primers used for screening are described previously (Schierup et al, 2001; Charlesworth et al, 2003a; Mable et al, 2003); primers that were used to amplify alleles described in this study are listed in Table 1. Clones were screened using the restriction endonucleases AluI and RsaI; at least three clones of each variant detected were sequenced using Big Dye (Applied Biosystems, Foster City, CA, USA) on an ABI 377 automated sequencer. Products from allele-specific primers were sequenced directly from gel bands purified using QiaQuick extraction kits (Qiagen Inc, Mississauga, ON, USA). Consensus sequences from three clones per sequence type and sequences from direct PCR products were aligned to previously characterized alleles (Charlesworth et al, 2003b) manually using Se-Al, version 1.0 (Rambaut, 1996) and MacClade, version 4.0 (Maddison and Maddison, 2000). When sequences were amplified in tetraploids that did not appear in the diploid populations, new allele-specific forward primers were designed (Table 1) and linkage of these putative alleles to the S-phenotype was tested in the families used for segregation analyses.
Segregation analyses
For diploid populations, segregation analyses of S-phenotypes (from pollination studies) and S-genotypes were initially performed separately to look for congruence without bias (Schierup et al, 2001; Mable et al, 2003). In tetraploids, however, due to the more complex nature of segregation possible with four gene copies per individual, segregation of S-genotypes could be performed independently, but grouping of phenotypes could not be completely assessed without considering genotype information. Multiple complex models of allelic segregation in tetraploids are possible (eg Jackson and Hauber, 1982; Wu et al, 2001), but for simplicity only the two most extreme models were considered here: (1) tetrasomic inheritance, which results when there is completely independent segregation of each allele (expected to be more common in autotetraploids); and (2) disomic inheritance, which results when parental alleles tend to segregate together so that inheritance is effectively diploid (expected to be more common in allotetraploids). Fit of these models to the expected genotype and phenotype classes was evaluated using χ2 with degrees of freedom equal to n−1, where n is the number of expected genotype classes. Copy number of each allele in parents was inferred based on segregation of alleles in offspring.
Phylogenetic analysis
The combined alignment of new tetraploid alleles with previously published diploid alleles (Charlesworth et al, 2003a; Popset alignment accession: gi:27545451) was used to estimate a hypothesis of phylogenetic relationships, using PAUP* 4.0b10. (Swofford, 2000). Previous data have shown that relationships among alleles at this locus are not well resolved because of their high divergence from one another, but that alleles fall into two broad groups (Charlesworth et al, 2003a). A maximum likelihood tree was constructed using an HKY85 model of evolution (ie estimating transition/transversion ratio and base frequencies: Hasegawa et al, 1985) to determine which of these groups the new tetraploid alleles fell into. A neighbor-joining (Saitou and Nei, 1987) bootstrap (1000 replicates) was performed to determine relative support for these relationships. Three paralogous genes from A. lyrata (Aly 10.1, Aly 10.2, and Aly 8) and their respective orthologues in A. thaliana (Ark 1, Ark 2, Ark 3) were used to root the tree (see Popset Alignment gi:27545451).
Results
Japanese tetraploids
Strength of SI
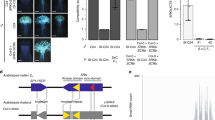
Polyploidy was first suspected in the Japanese population, due to the very large plant size compared to diploid populations. In addition, all progeny raised from field-collected seed sets from seven maternal plants produced autogamous fruits in the same greenhouse where self-incompatible diploids showed little or no fruit development without hand pollinations. Complete self-compatibility was maintained through two selfed generations (Figure 1) and chromosome counts and flow cytometry on these later generations confirmed that the plants were tetraploid (4x=32: Dart et al, 2004).
Phenogram showing relationships among alleles at the Aly 13 (SRK) locus of A. lyrata. The tree is rooted with paralogous genes from A. lyrata (Aly 10.1, Aly 10.2, and Aly 8) and their respective orthologues in A. thaliana (Ark 1, Ark 2, Ark 3). The tree was reconstructed using maximum likelihood under an HKY85 model of evolution and relative bootstrap support was estimated using neighbor-joining. The extensive divergence and possible gene conversion among alleles at this locus do not allow definitive estimation of relationships among individual alleles, but they can be broadly separated into two groups, one of which includes unlinked alleles 13-2 and 13-7 in a strongly supported clade (group B). Alleles found in tetraploid populations are underlined. The Austrian tetraploid alleles 13-28 and 13-29 fell into group B and were recessive to the alleles 13-30 and 13-33, which fell into group A. The Japanese tetraploids had the group A allele 13-22. The 13-1 allele was found in both populations.
Segregation of S-alleles
Only three SRK alleles were found in four first-generation individuals screened from Japan, with one of the alleles (Aly 13-1) found in a single individual and the other two (Aly 13-22 and 13-7) found in all individuals screened (Figure 1; Table 1). The first two of these alleles have been demonstrated to be linked to the S-locus in diploids (Aly 13-1 and 13-22: Schierup et al, 2001), while the third (13-7) is thought to be a paralogous gene that is not linked to the S-locus (Charlesworth et al, 2003a). Both the 13-1 and 13-22 sequences were identical to those found in diploid populations, but the unlinked allele had a reading-frame-altering single-base pair deletion in all individuals examined. Segregation analyses of alleles in the S2 generation indicated that at least one other linked allele must be present at the SRK locus because only 12/27 individuals (44%) showed positive amplification of the 13-22 allele found in the parent plant.
Austrian tetraploids
Strength of SI
All progeny raised from field-collected seeds from the seven maternal Austrian plants (first generation: J2000 seed set; Table 2) were confirmed to be tetraploid (4x=32: Dart et al, 2004), as originally inferred from allozyme data (S Ansell, personal communication). All individuals tested in subsequent generations (Table 3) were also tetraploid. Although most first-generation individuals were highly self-incompatible, progeny from two maternal plants (J2000/1 and J2000/7 families) showed evidence of partial self-compatibility (Table 2). This was not, however, found in the next generation. Crosses and selfings among plants within families between an apparently self-compatible plant (J2000 1/8, which produced 4/4 fruits on selfing) and a strongly self-incompatible plant (J2000 4/2, which produced 0/6 fruits on selfing) indicated that while plants within this second generation (C7 family; Table 4) were capable of setting seed when crossed to individuals with different SRK alleles, they were strongly self-incompatible. Similar results were found in the third generation, with all individuals tested showing strong SI (E5 family: Table 5).
Segregation and dominance of S-alleles
Screening for S-alleles at the SRK locus (Aly13) for first-generation plants (J2000) revealed four new alleles in the Austrian tetraploids (13-28, 13-29, 13-30, 13-33) that had not been found in previous surveys of diploid populations and one allele, 13-1, that has been found in all populations tested so far (Mable et al, 2003). Two of these new alleles (13-28 and 13-29) were originally found by sequencing PCR products amplified using an allele-specific primer for an allele found in a diploid population from Iceland (Aly 13-18). The alleles 13-30 and 13-33 were found by cloning and screening with the general primers 13F1 and 13FBM. The parents used to initiate subsequent generations (J2000 1/8 and 4/2) had the genotypes 1/28/33 and 1/29/30, respectively, at the Aly 13 (SRK) locus (Table 3). Assuming that each individual carries four alleles per locus, it was inferred that the 13-1 allele was present in two copies in both plants, since putative homozygotes for 13-1 appeared in the progeny; assignment of progeny genotypes was based on this assumption. It is possible that an unidentified recessive allele was present in the J2000 4/2 parent, but this does not substantially affect interpretation of results (it would have to be an allele that is recessive to all others). The unlinked allele, 13-7, was also present in some individuals but did not have the 1-bp deletion found in the Japanese population.
Diallele crosses for two third-generation families were made among progeny from seeds raised from crosses within the C7 seed family (E3 and E5 families; Table 3), but only the results for E5 are presented in detail here (Table 6). Concordance was high between the six SI phenotypes and 11 SI genotypes for the E5 family, allowing prediction of the following dominance relationships among parental alleles:
-
1)
Alleles 13-30 and 13-33 are codominant in both pollen and stigma (ie both are expressed when found together), but are dominant to 13-28, 13-29, and 13-1, as indicated by reciprocal incompatibility of SI phenotype groups I and V (which share allele 13-30), and groups I and IV (which share allele 13-33) compared with compatibility of groups sharing the other alleles.
-
2)
Alleles 13-28 and 13-29 are coexpressed in both pollen and stigma when they occur together, as indicated by reciprocal incompatibility of groups II and III (which share allele 13-29) and groups III and VI (which share allele 13-28). However, relative expression may differ in pollen and stigma in some individuals (see footnotes in Table 5 and details below).
-
3)
Allele 13-1 is recessive to 13-30 and 13-33 and probably recessive to 13-28 and 13-29 (even when present in three copies), based on reciprocal crosses among groups II, III, and VI.
Close examination of compatibility results (Table 6) indicated that some individuals appeared to show deviations from the above scheme in combination with certain other genotypes. For example, when used as pollen donors, two individuals with the genotype 1/1/28/29 showed different compatibilities when crossed to plants with the genotypes: (1) 1/1/1/29 (A4 produced 13/13 fruits; B6 produced 0/9 fruits), (2) 1/28/30/33 (A4 produced 2/3 fruits; B6 produced 0/3 fruits), and (3) 1/1/28/33 (A4 produced 6/12 fruits; B6 produced 2/12 fruits). The corresponding reciprocal crosses did not show such discrepancies. These results can be explained as follows: (1) 13-29 is coexpressed with 13-28 in both pollen and stigmas as predicted by the dominance scheme above in individual B6, whereas 13-28 is dominant to 13-29 in pollen and codominant in stigmas in individual A4; (2) 13-28 (and/or 13-1) is expressed with the dominant alleles 13-30 and 13-33 in stigmas (but not in pollen) of the individual that B6 was crossed to as pollen donor (A4 and B6 were crossed to two different individuals because flowering stopped before the full diallele could be completed); and (3) 13-28 (and/or 13-1) is at least partially expressed in stigmas (but not in pollen) with 13-33 for all four individuals with the genotype 1/1/28/33. It cannot be determined from these results whether it is 13-28 or 13-1 (or both) that has altered expression but the anomaly described in footnote b (Table 5) suggests that 13-1 is expressed with 13-33 in the pollen of individual A12 and with 13-30 in stigmas of individual A11, but that 13-1 is not expressed in both in the reciprocal combination. Footnote c suggests that 13-1 is also expressed in the pollen of individual A11, but that only one out of three individuals with the genotype 1/1/1/33 expresses 13-1 in its stigma. In addition, the fact that pollen from A11 is incompatible with stigmas of individuals with the genotype 1/1/1/28 but compatible with those with the genotype 1/1/1/29 suggests that 13-29 is dominant to 13-1 in these individuals and/or that 13-1 and 13-28 are coexpressed in pollen. Unfortunately, crossing information with individuals with the genotype 1/1/1/28 was incomplete because one individual showed reduced fertility and the rest did not flower long enough to be crossed to all other individuals. Similar individual variation in allele expression was seen in family E3, which segregated for alleles 13-1, 28, 30, and 33. In this family, a cross between a homozygous individual with genotype 1/1/1/1 and an individual with genotype 1/1/1/33 confirmed that 13-1 is expressed with 13-33 in stigmas, but not in pollen. However, when the cross was repeated with another 1/1/1/33 individual, the cross was fully compatible, suggesting that 13-1 was not expressed in stigmas in this plant.
The proposed dominance model is consistent with the observed segregation of phenotypes and genotypes in family E5 (Table 6), based on a tetrasomic model of inheritance (P=0.29 and 0.12, respectively). A disomic model of inheritance would predict a maximum of only four genotypes, but 16 were seen here. The genotypes 28/29/30/33, 1/28/29/30, 1/1/1/30, and 1/1/1/1 were missing, but this is not unexpected given the large number of expected classes. To increase the sample size and evaluate the possibility that some genotypes might be at a disadvantage, a larger family (D1 family n=41; Table 3) sharing the same five alleles was studied. All expected genotypes were found; the only genotype that was consistently under-represented in both families was 1/1/1/30 and segregation was consistent with a tetrasomic model of inheritance (P=0.10). Additional families screened for segregation of SRK alleles but not used in pollinations also supported a tetrasomic model of inheritance at this locus (Table 3). Deviation from expected patterns in family E1 appeared to be due to excess of genotypes 1/1/1/29, 1/1/1/33, and 1/1/30/33 and deficiency of 1/1/1/30, 1/29/30/33, and 1/1/29/33, but this was not seen in the reciprocal cross (family E2) involving the same parents.
Phylogenetic analysis
The new tetraploid alleles from the Austrian population fell into the same distinctive classes of alleles previously found for diploid populations: group B alleles form a strongly supported (based on bootstrap analysis) cluster that are closely related to a group of unlinked alleles (including the 13-7 allele discussed above), whereas group A alleles do not form a tight cluster but are distinguishable from group B sequences based on DNA sequences and conserved protein motifs (Charlesworth et al, 2003a). The tetraploid alleles in group B (13-28 and 13-29) appeared to be most closely related to diploid alleles 13-3 (found in North American populations) and 13-18 (found in Icelandic populations), respectively. Relationships among group A sequences were not resolved by phylogenetic analyses because of their extensive divergence from one another, but 13-33 appeared to be most closely related to diploid alleles 13-4 and 13-10 (found in Scottish populations). The Japanese tetraploid allele 13-22 was identical in sequence to an allele (found in Icelandic diploids) previously demonstrated to be dominant to the group B allele 13-3, and to the group A alleles 13-1 and 13-13 (Schierup et al, 2001).
Discussion
Polyploidy and SI
Our results suggest that, as for other species with sporophytic control of SI (eg Lundqvist, 1998; Young et al, 2000), polyploidy per se probably does not cause breakdown of SI in A. lyrata. Although tetraploids from Japan were completely self-compatible, those from a population in Austria showed strong SI, with dominance of SI alleles apparently operating similar to that in related diploid populations. Our results also suggest that, contrary to recent findings of dramatic changes in gene expression and rearrangements in polyploids (reviewed by Osborn et al, 2003), inheritance and dominance of a very complicated genetic system appears to function remarkably similarly to that in related diploid populations. In contrast to a disomic inheritance pattern of S-alleles (but without considering dominance relationships) suggested in tetraploid Raphanus repens (Lundqvist, 1998), our results suggest independent segregation of all four alleles at the S-locus in self-incompatible tetraploids (ie tetrasomic inheritance). These data provide the potential for a more detailed study of the population genetics of genes under balancing selection in tetraploids. The fate of duplicated genes in polyploids has been investigated for the most widely studied vertebrate cell–cell recognition system (the major histocompatibility locus or MHC: Sammut et al, 2002), but similar studies have not been conducted previously on a plant system.
In the diploid selfing species A. thaliana, a functional SI response is produced when SRK/SCR genes are introduced from self-incompatible A. lyrata (Nasrallah et al, 2002), suggesting that loss of SI is not due to loss of the downstream machinery but is associated with particular haplotypes at the S-locus. Complete genotypes were not obtained for the self-compatible Japanese tetraploids (segregation analyses indicated that one additional SRK allele is present in the families examined). However, all individuals screened had a 13-22 allele that was identical in sequence to that from diploid populations from Iceland, but also shared a 13-7 allele with a 1-bp deletion that would change the reading frame and presumably make the protein nonfunctional. In diploids, this latter allele is not linked to the SI recognition phenotype (Charlesworth et al, 2003a), and it is tempting to speculate that it could represent a locus involved in complete expression of SI or one which interacts with functional alleles through gene conversion, such as occurs among genes in the MHC complex of vertebrates (Charlesworth et al, 2003a). It is not known where this gene resides in relation to the S-locus. For RNase-based gametophytic SI systems, modifier loci have been suggested to influence the expression of SI (Lewis, 1943) and result in weakened SI (pseudocompatibility: Levin, 1993), such as seen in some crosses involving the Austrian population (Table 6). Similar genotype-dependent variation in SSI phenotypic responses in diploid Senecio squalidus have been explained on the basis of influences from a cryptic gametophytic system (Brennan et al, 2002), as suggested by Lewis (1994) for Brassica. Insufficient data exist to determine whether the locus represented by allele 13-7 or some other unidentified locus influences the expression of dominance, but the pattern is intriguing.
The apparent partial self-compatibility of some of the individuals raised from field-collected seeds from Austria may be attributable to altered interactions among alleles in some genetic contexts, as described for their progeny (and grand-progeny). Individual J2000 1/8 produced 4/4 full-sized fruits on selfing (15% of these seeds were viable) and was used as a parent for the C7 family (Tables 2 and 3). The genotype of this individual was 1/1/28/33, which was suggested as one of the genotypes that showed different dominance of alleles 13-28 and/or 13-1 in pollen and stigma relative to allele 13-33 (and 13-30) in some individuals in the E5 family. Two out of four individuals with this genotype in family E3 also produced full-sized fruits on selfing and several crosses between individuals with this genotype were unexpectedly compatible. Several individuals with the genotype 1/1/28/30 also showed partial self-compatibility in this family. One explanation is that dominance relationships among alleles differ in pollen and stigma, with 13-33 (or 13-30) expressed in one and 13-28 (or 13-1) in the other. Theoretically, linkage to modifiers has been implicated in the evolution of dominance (Ewens, 1966), suggesting that it is possible that altered dominance in pollen and stigma in some individuals could be due to the influence of such a locus. The possible involvement of the locus including 13-7 in such expression changes is currently being investigated in these families (two slightly different sequences are segregating) and in individuals from diploid populations of A. lyrata that show similar genotype-dependent changes in the expression of SI. Crosses between the Japanese and Austrian tetraploids have produced only inviable seeds and triploid individuals resulting from crosses between the Japanese tetraploids and SI diploids are sterile, so it has not been possible to examine inheritance of the loss of SI between these populations, but it may be possible to conduct more extensive comparisons within the Austrian populations.
Polyploidy and dominance of SI alleles
In diploid Brassica, there are two recognizably distinct dominance classes of S-alleles, with class I alleles dominant to class II alleles in pollen but codominant in stigma. However, nonlinear dominance hierarchies, partial expression of SI, and different dominance in pollen and stigma have also been described (Thompson and Taylor, 1966, 1971; Hatakeyama et al, 1998; Shiba et al, 2002; Kakizaki et al, 2003). In diploid A. lyrata, there may be at least four dominance classes, but relationships within and among these classes are still under study (N Prigoda and BK Mable, unpublished; Bechsgaard et al, 2004). One class appears to be dominant to all others (including allele 13-22 found in the Japanese population), but there appear to be three recessive classes, the most recessive being represented only by allele 13-1. Geneological relationships among diploid alleles indicate that one well-supported group of alleles that includes the 13-7 locus (Group B: Figure 2, Charlesworth et al, 2003a) are recessive in both pollen and stigma (N Prigoda and BK Mable, unpublished) to alleles in the second major group, which includes a putative orthologous pseudogene from A. thaliana (Group A: Figure 2, Charlesworth et al, 2003a). Dominance relationships among the group A alleles are more complex, with some alleles showing intermediate patterns of dominance and evidence of partial SI (Mable et al, 2003), as has been described for Brassica Type I alleles (eg Ripley and Beversdorf, 2003). Given these potential complexities, it is rather remarkable that tetraploid alleles fall so cleanly into the same classes predicted for diploids (Tables 3 and 6). The tetraploid group B alleles (13-28 and 13-29) are recessive (in both pollen and stigma) to those in group A (13-30 and 13-33), and alleles generally appear to be codominant within classes (Figure 1).
The apparent presence of 13-1 in multiple copies in individuals from field-collected seeds supports evidence from diploids that the recessive 13-1 allele is present at much higher frequency than other alleles in the population (Mable et al, 2003), and the results presented here suggest that dosage effects do not alter its dominance relationships in most crosses. More detailed crossing experiments are required to assess whether the anomalies described for some individuals involve relative expression of 13-1 or 13-28 in some contexts, and to investigate dosage effects in detail. However, overall, 13-1 appears to remain recessive in combination with all other alleles. In the E5 family, it is theoretically possible that a fourth, unidentified allele was present in one of the parents (the other parent had to have two copies of 13-1 to explain segregation results in the progeny), but this would have to be recessive to all other alleles, including 13-1.
In both diploid A. lyrata and diploid Brassica, dominance has been found to depend on the relative expression of the male (SCR) rather than the female (SRK) component of SI (Hatakeyama et al, 1998; Kusaba et al, 2002; Kakizaki et al, 2003), but only S-alleles within group A were included in these studies. It would be interesting to see if all four alleles for both male and female genes are expressed in tetraploids, and to determine if this relates to observed dominance relationships in all four dominance classes.
An added benefit of examining segregation in tetraploids is that it may be possible to look at the intransitiveness of dominance by combining four different alleles within individuals and examining the effects of dosage of particular allelic combinations. The progeny from the crosses described here could be used for this purpose, as they should include individuals with multiple copies of alleles other than 13-1. The genotype-dependent anomalies in dominance relationships noted here in particular deserve more detailed study.
Conclusions
It does not appear that polyploidy necessarily disrupts the functioning of SI alleles in the sporophytic system described here. Since nothing is known about the history of these tetraploids (ie whether they are autopolyploids or allopolyploids, the age of the polyploidization event, or parental origins), it is unclear whether the current mating systems remained the same with polyploidization or whether these systems have evolved subsequent to the genome duplication event. The best test of this would be to examine the strength of SI and dominance relationships among alleles in neopolyploids generated from diploids with alleles with known dominance relationships. Colchicine itself has been found to affect the strength of SI (Pandey, 1968), so it would be necessary to induce polyploids through natural means for this purpose. It is also not known whether independent segregation (ie tetrasomic inheritance) is found across the genome or whether it is only found at some loci, including the S-locus. Segregation analyses of microsatellite loci are currently underway to assess this. Comparing all possible combinations of compatibility types (SC and SI) and ploidy levels (2n and 4n) in one species would provide the opportunity to better our understanding of the evolution and genetic implications of sporophytic SI. With populations such as these, one could examine in detail the factors that control cell–cell recognition in this species and what factors might lead to a breakdown in the process. Finally, understanding dominance interactions in tetraploids will help to elucidate how interactions among duplicated gene copies may influence traits in new and old tetraploids.
References
Abbott RJ, Bretagnolle FC, Thebaud C (1998). Evolution of a polymorphism for outcrossing rate in Senecio vulgaris: influence of germination behavior. Evolution 52: 1593–1601.
Al-Shehbaz IA, O’Kane S, Price RA (1999). Generic placement of species excluded from Arabidopsis (Brassicaceae). Novon 9: 297–307.
Arumuganathan K, Earle ED (1991). Nuclear DNA content of some important plant species. Plant Mol Biol Reporter 9: 208–218.
Barow M, Meister A (2003). Endopolyploidy in seed plants is differently correlated to systematics, organ, life strategy and genome size. Plant Cell Environ 26: 571–584.
Bateman AJ (1952). Self-incompatibility systems in angiosperms. I. Theory. Heredity 6: 285–310.
Bechsgaard J, Bataillon T, Schierup MH (2004). Uneven segregation of sporophytic self-incompatibility alleles in Arabidopsis lyrata. J Evol Biol 17: 554–561.
Bino RJ, De Vries JN, Kraak HL, Van Pijlen JG (1992). Flow cytometric determination of nuclear replication stages in tomato seeds during priming and germination. Ann Bot 69: 231–236.
Brennan A, Harris S, Tabah D, Hiscock S (2002). The population genetics of sporophytic self-incompatibility in Senecio squalidus L. (Asteraceae) I: S allele diversity in a natural population. Heredity 89: 430–438.
Brugière N, Cui Y, Bi Y-M, Arnoldo M, Jackman L, Rothstein SJ (2000). Molecular genetics of self-incompatibility in Brassica napus. Ann Bot 85: 133–140.
Charlesworth D, Awadalla P, Mable BK, Schierup MH (2000). Population-level studies of multiallelic self-incompatibility loci, with particular reference to Brassicaceae. Ann Bot 85 (Suppl A): 227–239.
Charlesworth D, Bartolomê-Husson C, Schierup MH, Mable BK (2003a). Haplotype structure of the stigmatic self-incompatibility gene in natural populations of Arabidopsis lyrata. Mol Biol Evol 20: 1741–1753.
Charlesworth D, Mable BK, Schierup MH, Bartolomê-Husson C, Awadalla P (2003b). Diversity and linkage of genes in the self-incompatibility gene family in Arabidopsis lyrata. Genetics 164: 1519–1535.
Chawla B, Bernatzky R, Liang W, Marcotrigiano M (1997). Breakdown of self-incompatibility in tetraploid Lycopersicum peruvianum: inheritance and expression of S-related proteins. Theor Appl Genet 95: 992–996.
Ciu Y, Brugière N, Jackman L, Bi Y-M, Rothstein SJ (1999). Structural and transcriptional comparative analyses of the S locus regions in two self-incompatible Brassica napus lines. Plant Cell 11: 2217–2231.
Dart S, Kron P, Mable BK (2004). Characterizing polyploidy in Arabidopsis lyrata using chromosome counts and flow cytometry. Can J Bot 82: 185–197.
Ewens WJ (1966). Linkage and the evolution of dominance. Heredity 21: 363–370.
Goring DR, Gavin TL, Schafer U, Rothstein SJ (1993). An S receptor kinase gene in self-compatible Brassica napus has a 1-bp deletion. Plant Cell 5: 531–539.
Hasegawa M, Kishino K, Yano T (1985). Dating the human–ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22: 160–174.
Hatakeyama K, Watanabe M, Takasaki T, Ojima K, Hinata K (1998). Dominance relationships between S-alleles in self-incompatible Brassica campestris L. Heredity 80: 241–247.
Hauck NR, Yamane H, Tao R, Iezzoni AF (2002). Self-compatibility and incompatibility in sour cherry (Prunus cerasus L.). Sex Plant Reprod 15: 39–46.
Henderson SA, Lu BC (1968). The use of haematoxylin for squash preparations of chromosomes. Stain Technol 43: 233–236.
Hiscock S, Tabah D (2003). The different mechanisms of sporophytic self-incompatibility. Philos Trans R Soc Lond B 358: 1037–1045.
Jackson RC, Hauber DP (1982). Autotriploid and autotetraploid cytogenetic analyses: correction coefficients for proposed binomial models. Am J Bot 69: 644–646.
Kakizaki T, Takada Y, Ito A, Suzuki G, Shiba H, Takayama S et al (2003). Linear dominance relationship among four class-II S haplotypes in pollen is determined by the expression of SP11 in Brassica self-incompatibility. Plant Cell Physiol 44: 70–75.
Koch MA, Bishop JG, Mitchell-Olds T (1999). Molecular systematics and evolution of Arabidopsis and Arabis. Plant Biol 1: 529–537.
Kowyama Y, Takahasi H, Muraoka K, Tani T, Hara K, Shiotani L (1994). Number, frequency and dominance relationships of S-alleles in diploid Ipomoea trifida. Heredity 73: 275–283.
Kusaba M, Dwyer KG, Hendershot J, Vrebalov J, Nasrallah JB, Nasrallah ME (2001). Self-incompatibility in the genus Arabidopsis: characterization of the S locus in the outcrossing A. lyrata and its autogamous relative A. thaliana. Plant Cell 13: 627–643.
Kusaba M, Tung C-W, Nasrallah ME, Nasrallah JB (2002). Monoallelic expression and dominance interactions in anthers of self-incompatible Arabidopsis lyrata. Plant Phys 128: 17–20.
Levin DA (1993). S-gene polymorphism in Phlox drummondii. Heredity 71: 193–198.
Lewis D (1943). Physiology of incompatibility in plants. III. Autopolyploids. J Genet 45: 171–185.
Lewis D (1947). Competition and dominance of incompatibility alleles in diploid pollen. Heredity 1: 85–108.
Lewis D (1994). Gametophytic-sporophytic incompatibility. In: Williams EG, Clarke AE, Knox RB (eds) Genetic Control of Self-incompatibility and Reproductive Development in Flowering Plants, Kluwer Press: Dordrecht. pp 88–101.
Lundqvist A (1998). Disomic control of self-incompatibility in the tetraploid Ranunculus repens (Ranunculaceae). Hereditas 128: 181–183.
Mable BK (2004a). Evolutionary genetics of self-incompatibility in a new ‘model’ plant: Arabidopsis lyrata. In: Cronk QC, Whitton J, Taylor IEP (eds) Plant Adaptation: Molecular Biology and Ecology, NRC Research Press: Ottawa, (in press).
Mable BK (2004b). Polyploidy and self-compatibility: Is there an association? New Phyt 162: 803–811.
Mable BK, Schierup MH, Charlesworth D (2003). Estimating the number, frequency, and dominance of S-alleles in a natural population of Arabidopsis lyrata (Brassicaceae) with sporophytic control of self-incompatibility. Heredity 90: 422–431.
Maddison DR, Maddison WP (2000). MacClade 4: Analysis of phylogeny and character evolution. Version 4.0, Sinauer Associates: Sunderland, MA.
Mehlenbacher S (1997). Revised dominance hierarchy for S-alleles in Corylus avellana. Theor Appl Genet 94: 360–364.
Miller JS, Venable DL (2000). Polyploidy and the evolution of gender dimorphism in plants. Science 289: 2335–2338.
Mitchell-Olds T (2001). Arabidopsis thaliana and its wild relatives: a model system fo ecology and evolution. Trends Ecol Evol 16: 693–699.
Miyashita NT, Kawabe A, Innan H, Terauchi R (1998). Intra- and interspecific DNA variation and codon bias of the alcohol dehydrogenase (Adh) locus in Arabis and Arabidopsis species. Mol Biol Evol 15: 1420–1429.
Nasrallah JB (2002). Recognition and rejection of self in plant reproduction. Science 296: 305–308.
Nasrallah ME (2000). Arabidopsis species hybrids – emerging model systems for the analysis of species differences. J Plant Growth Regul 19: 326–333.
Nasrallah ME, Kandasamy M, Chang M-C, Stadler Z, Lim S, Nasrallah JB (2000). Identifying genes for pollen-stigma recognition in crucifers. Ann Bot 85 (Suppl A): 125–132.
Nasrallah ME, Liu P, Nasrallah JB (2002). Generation of self-incompatibile Arabidopsis thaliana by transfer of two S locus genes from A. lyrata. Science 297: 247–249.
Osborn TC, Pires JC, Birchler JA, Auger DL, Chen ZJ, Lee H-S et al (2003). Understanding mechanisms of novel gene expression in polyploids. Trends Genet 19: 141–147.
Pandey KK (1968). Colchicine-induced changes in the self-incompatibility behaviour of Nicotiana. Genetica 39: 257–271.
Polatschek A (1966). Cytotaxonomische Beitraege zur Flora der Ostalpenlaender. Oesterr Botan ZeitschriftOesterr Botan Zeitschrift Bd: 113.
Rambaut A (1996) Se-Al: Sequence alignment editor, version 1.0 alpha1. Distributed over the world wide web (http://evolve.zps.ox.ac.uk ).
Ripley VL, Beversdorf WD (2003). Development of self-incompatible Brassica napus: (II) Brassica oleracea S-allele expression in B. napus. Plant Breed 122: 6–11.
Robert LS, Allard S, Franklin TM, Trick M (1994). Sequence and expression of endogenous S-locus glycoprotein genes in self-compatible Brassica napus. Mol Gen Genet 242: 209–216.
Rollins RC (1993). The Cruciferae of Continental North America, Stanford University Press: Stanford, CA.
Ross KJ, Fransz P, Jones GH (1996). A light microscope atlas of meiosis in Arabidopsis thaliana. Chromosom Res 4: 507–516.
Saitou N, Nei M (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4: 406–425.
Sammut B, Marcuz A, Pasquier L (2002). The fate of duplicated major histocompatibility complex class Ia genes in a dodecaploid amphibian, Xenopus ruwenzoriensis. Eur J Immunol 32: 2698–2709.
Sampson DR (1967). Frequency and distribution of self-incompatibility alleles in Raphanus raphanistrum. Genetics 56: 151–241.
Schierup MH (1998). The number of self-incompatibility alleles in a finite, subdivided population. Genetics 149: 1153–1162.
Schierup MH, Mikkelsen AM, Hein J (2001). Recombination, balancing selection and phylogenies in MHC and self-incompatibility genes. Genetics 159: 1833–1844.
Shiba H, Iwano M, Entani T, Ishimoto K, Shimosato H, Che F-S et al (2002). The dominance of alleles controlling self-incompatibility in Brassica pollen is regulated at the RNA level. Plant Cell 14: 491–504.
Stevens JP, Kay QON (1989). The number, dominance relationships and frequencies of self-incompatibility alleles in a natural population of Sinapis arvensis L. in South Wales. Heredity 62: 199–205.
Stone JL (2002). Molecular mechanisms underlying the breakdown of gametophytic self-incompatibility. Quart Rev Biol 77: 17–32.
Swofford DL (2000). PAUP* 4.0, Sinauer Associates: Sunderland, MA.
Thompson KF, Taylor JP (1966). Non-linear dominance relationships between S alleles. Heredity 21: 345–362.
Thompson KF, Taylor JP (1971). Self-compatibility in kale. Heredity 27.
Wu SS, Wu R, Ma CX, Zeng ZB, Yang MCK, Casella G (2001). A multivalent pairing model of linkage analysis in autotetraploids. Genetics 159: 1339–1350.
Yamane H, Ikeda K, Hauck NR, Iezzoni AF, Tao R (2003). Self-incompatibility (S) locus region of the mutated S6-haplotype of sour cherry (Prunus cerasus) contains a functional pollen S allele and a non-functional pistil S allele. J Exp Bot 54: 2337–2431.
Young AG, Millar C, Gregory E, Langston A (2000). Sporophytic self-incompatibility in diploid and tetraploid races of Rutidosis leptorrhynchoides (Asteraceae). Austr J Bot 48: 667–672.
Yu K, Schafer U, Glavin TL, Goring TR, Rothstein SJ (1996). Molecular characterization of the S locus in two self-incompatible Brassica napus lines. Plant Cell 8: 2369–2380.
Acknowledgements
We thank N Prigoda and L Mercer for help with pollinations, S Dart and M Lesosky for help with plant maintenance, P Kron for collecting the flow cytometry data, D Tabah for interesting discussions about explanations for anomalous crossing results and especially, D Charlesworth for helpful comments on the manuscript, assistance with pollinations, and continued support. Two anonymous reviewers provided helpful comments on the manuscript. This research was supported by the Canadian Foundation for Innovation, the Ontario Innovation Trust, and the Natural Sciences and Engineering Council of Canada.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mable, B., Beland, J. & Di Berardo, C. Inheritance and dominance of self-incompatibility alleles in polyploid Arabidopsis lyrata. Heredity 93, 476–486 (2004). https://doi.org/10.1038/sj.hdy.6800526
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800526
Keywords
This article is cited by
-
Ancestral self-compatibility facilitates the establishment of allopolyploids in Brassicaceae
Plant Reproduction (2023)
-
Analysis of genetic variation using ISSR and the development of SCAR marker in synthetic autotetraploids of Vigna mungo (L.) Hepper
Vegetos (2019)
-
What causes mating system shifts in plants? Arabidopsis lyrata as a case study
Heredity (2017)
-
Taming the wild: resolving the gene pools of non-model Arabidopsislineages
BMC Evolutionary Biology (2014)
-
Population fragmentation causes randomly fixed genotypes in populations of Arabidopsis kamchatica in the Japanese Archipelago
Journal of Plant Research (2012)