Abstract
Clonal hematopoiesis (CH) increases the risk of atherosclerotic cardiovascular disease possibly due to increased plaque inflammation. Human studies suggest that limitation of interleukin-6 (IL-6) signaling could be beneficial in people with large CH clones, particularly in TET2 CH. Here we show that IL-6 receptor antibody treatment reverses the atherosclerosis promoted by Tet2 CH, with reduction of monocytosis, lesional macrophage burden and macrophage colony-stimulating factor 1 receptor (CSF1R) expression. IL-6 induces expression of Csf1r in Tet2-deficient macrophages through enhanced STAT3 binding to its promoter. In mouse and human Tet2-deficient macrophages, IL-6 increases CSF1R expression and enhances macrophage survival. Treatment with the CSF1R inhibitor PLX3397 reversed accelerated atherosclerosis in Tet2 CH mice. Our study demonstrates the causality of IL-6 signaling in Tet2 CH accelerated atherosclerosis, identifies IL-6-induced CSF1R expression as a critical mechanism and supports blockade of IL-6 signaling as a potential therapy for CH-driven cardiovascular disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and its supplementary information. Data regarding the single cell RNA sequence of CSF1R expression can be accessed at https://zenodo.org/record/7876218#.ZEvhOHbMIVA. Source data are provided with this paper.
References
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Busque, L. et al. Recurrent somatic TET2 mutations in normal elderly individuals with clonal hematopoiesis. Nat. Genet. 44, 1179–1181 (2012).
Steensma, D. P. et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 126, 9–16 (2015).
Jaiswal, S. et al. Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med. 377, 111–121 (2017).
Tall, A. R. & Fuster, J. J. Clonal hematopoiesis in cardiovascular disease and therapeutic implications. Nat. Cardiovasc. Res. 1, 116–124 (2022).
Bhattacharya, R. et al. Clonal hematopoiesis is associated with higher risk of stroke. Stroke 53, 788–797 (2022).
Dorsheimer, L. et al. Association of mutations contributing to clonal hematopoiesis with prognosis in chronic ischemic heart failure. JAMA Cardiol. 4, 25–33 (2019).
Pascual-Figal, D. A. et al. Clonal hematopoiesis and risk of progression of heart failure with reduced left ventricular ejection fraction. J. Am. Coll. Cardiol. 77, 1747–1759 (2021).
Yu, B. et al. Supplemental association of clonal hematopoiesis with incident heart failure. J. Am. Coll. Cardiol. 78, 42–52 (2021).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Ridker, P. M. et al. Antiinflammatory therapy with canakinumab for atherosclerotic disease. N. Engl. J. Med. 377, 1119–1131 (2017).
Tardif, J.-C. et al. Efficacy and safety of low-dose colchicine after myocardial infarction. N. Engl. J. Med. 381, 2497–2505 (2019).
Nidorf, S. M. et al. Colchicine in patients with chronic coronary disease. N. Engl. J. Med. 383, 1838–1847 (2020).
Fuster, J. J. et al. Clonal hematopoiesis associated with TET2 deficiency accelerates atherosclerosis development in mice. Science 355, 842–847 (2017).
Fidler, T. P. et al. The AIM2 inflammasome exacerbates atherosclerosis in clonal haematopoiesis. Nature 592, 296–301 (2021).
Ridker, P. M. & Rane, M. Interleukin-6 signaling and anti-interleukin-6 therapeutics in cardiovascular disease. Circ. Res. 128, 1728–1746 (2021).
Heink, S. et al. Trans-presentation of IL-6 by dendritic cells is required for the priming of pathogenic TH17 cells. Nat. Immunol. 18, 74–85 (2017).
Tipping, C. C-reactive protein concentration and risk of coronary heart disease, stroke, and mortality: an individual participant meta-analysis. Lancet 375, 132–140 (2010).
Ridker, P. M., Hennekens, C. H., Buring, J. E. & Rifai, N. C-reactive protein and other markers of inflammation in the prediction of cardiovascular disease in women. N. Engl. J. Med. 342, 836–843 (2000).
Ridker, P. M., Rifai, N., Stampfer, M. J. & Hennekens, C. H. Plasma concentration of interleukin-6 and the risk of future myocardial infarction among apparently healthy men. Circulation 101, 1767–1772 (2000).
Volpato, S. et al. Cardiovascular disease, interleukin-6, and risk of mortality in older women: the women’s health and aging study. Circulation 103, 947–953 (2001).
Bick, A. G. et al. Genetic interleukin 6 signaling deficiency attenuates cardiovascular risk in clonal hematopoiesis. Circulation 141, 124–131 (2020).
Bick, A. G. et al. Inherited causes of clonal haematopoiesis in 97,691 whole genomes. Nature 586, 763–768 (2020).
Tosato, G. & Jones, K. D. Interleukin-1 induces interleukin-6 production in peripheral blood monocytes. Blood 75, 1305–1310 (1990).
Ridker, P. M. et al. Modulation of the interleukin-6 signalling pathway and incidence rates of atherosclerotic events and all-cause mortality: analyses from the Canakinumab Anti-Inflammatory Thrombosis Outcomes Study (CANTOS). Eur. Heart J. 39, 3499–3507 (2018).
Swerdlow, D. I. et al. The interleukin-6 receptor as a target for prevention of coronary heart disease: a Mendelian randomisation analysis. Lancet 379, 1214–1224 (2012).
Scheller, J. & Rose-John, S. The interleukin 6 pathway and atherosclerosis. Lancet 380, 338 (2012).
Nachun, D. et al. Clonal hematopoiesis associated with epigenetic aging and clinical outcomes. Aging Cell 20, e13366 (2021).
Kessler, M. D. et al. Common and rare variant associations with clonal haematopoiesis phenotypes. Nature 612, 301–309 (2022).
Vlasschaert, C., Heimlich, J. B., Rauh, M. J., Natarajan, P. & Bick, A. G. Interleukin-6 receptor polymorphism attenuates clonal hematopoiesis-mediated coronary artery disease risk among 451 180 individuals in the UK Biobank. Circulation 147, 358–360 (2023).
Huber, S. A., Sakkinen, P., Conze, D., Hardin, N. & Tracy, R. Interleukin-6 exacerbates early atherosclerosis in mice. Arterioscler. Thromb. Vasc. Biol. 19, 2364–2367 (1999).
Schieffer, B. et al. Impact of interleukin-6 on plaque development and morphology in experimental atherosclerosis. Circulation 110, 3493–3500 (2004).
Song, L. & Schindler, C. IL-6 and the acute phase response in murine atherosclerosis. Atherosclerosis 177, 43–51 (2004).
Zhang, Q. et al. Tet2 is required to resolve inflammation by recruiting Hdac2 to specifically repress IL-6. Nature 525, 389–393 (2015).
Piconese, S. et al. Mast cells counteract regulatory T-cell suppression through interleukin-6 and OX40/OX40L axis toward Th17-cell differentiation. Blood 114, 2639–2648 (2009).
Tsukamoto, H., Senju, S., Matsumura, K., Swain, S. L. & Nishimura, Y. IL-6-mediated environmental conditioning of defective Th1 differentiation dampens antitumour immune responses in old age. Nat. Commun. 6, 6702 (2015).
Busque, L. et al. High-sensitivity C-reactive protein is associated with clonal hematopoiesis of indeterminate potential. Blood Adv. 4, 2430–2438 (2020).
Welsh, C. et al. Association of total and differential leukocyte counts with cardiovascular disease and mortality in the UK Biobank. Arterioscler. Thromb. Vasc. Biol. 38, 1415–1423 (2018).
Murphy, A. J. et al. ApoE regulates hematopoietic stem cell proliferation, monocytosis, and monocyte accumulation in atherosclerotic lesions in mice. J. Clin. Invest. 121, 4138–4149 (2011).
Randolph, G. J. The fate of monocytes in atherosclerosis. J. Thromb. Haemost. 7, 28–30 (2009).
Swirski, F. K. et al. Ly-6Chi monocytes dominate hypercholesterolemia-associated monocytosis and give rise to macrophages in atheromata. J. Clin. Invest. 117, 195–205 (2007).
Libby, P. Collagenases and cracks in the plaque. J. Clin. Invest. 123, 3201–3203 (2013).
Cochain, C. et al. Single-cell RNA-seq reveals the transcriptional landscape and heterogeneity of aortic macrophages in murine atherosclerosis. Circ. Res. 122, 1661–1674 (2018).
Clinton, S. K. et al. Macrophage colony-stimulating factor gene expression in vascular cells and in experimental and human atherosclerosis. Am. J. Pathol. 140, 301–316 (1992).
Rosenfeld, M. E. et al. Macrophage colony-stimulating factor mRNA and protein in atherosclerotic lesions of rabbits and humans. Am. J. Pathol. 140, 291–300 (1992).
Sinha, S. K. et al. Local M-CSF (macrophage colony-stimulating factor) expression regulates macrophage proliferation and apoptosis in atherosclerosis. Arterioscler. Thromb. Vasc. Biol. 41, 220–233 (2021).
Rajavashisth, T. et al. Heterozygous osteopetrotic (op) mutation reduces atherosclerosis in LDL receptor- deficient mice. J. Clin. Invest. 101, 2702–2710 (1998).
Dutta, P. et al. Macrophages retain hematopoietic stem cells in the spleen via VCAM-1. J. Exp. Med. 212, 497–512 (2015).
Qiao, J. H. et al. Role of macrophage colony-stimulating factor in atherosclerosis: studies of osteopetrotic mice. Am. J. Pathol. 150, 1687–1699 (1997).
Robbins, C. S. et al. Local proliferation dominates lesional macrophage accumulation in atherosclerosis. Nat. Med. 19, 1166–1172 (2013).
Hume, D. A. & MacDonald, K. P. A. Therapeutic applications of macrophage colony-stimulating factor-1 (CSF-1) and antagonists of CSF-1 receptor (CSF-1R) signaling. Blood 119, 1810–1820 (2012).
Cai, Z. et al. Inhibition of inflammatory signaling in Tet2 mutant preleukemic cells mitigates stress-induced abnormalities and clonal hematopoiesis. Cell Stem Cell 23, 833–849 (2018).
Cai, Z. et al. Hyperglycemia cooperates with Tet2 heterozygosity to induce leukemia driven by proinflammatory cytokine-induced lncRNA Morrbid. J. Clin. Invest. 131, e140707 (2021).
Zhong, Z., Wen, Z. & Darnell, J. E. Jr. Stat3: a STAT family member activated by tyrosine phosphorylation in response to epidermal growth factor and interleukin-6. Science 264, 95–98 (1994).
Zhuang, S. Regulation of STAT signaling by acetylation. Cell. Signal. 25, 1924–1931 (2013).
Yuan, Z.-L., Guan, Y.-J., Chatterjee, D. & Chin, Y. E. Stat3 dimerization regulated by reversible acetylation of a single lysine residue. Science 307, 269–273 (2005).
Wang, R., Cherukuri, P. & Luo, J. Activation of Stat3 sequence-specific DNA binding and transcription by p300/CREB-binding protein-mediated acetylation. J. Biol. Chem. 280, 11528–11534 (2005).
Ray, S., Lee, C., Hou, T., Boldogh, I. & Brasier, A. R. Requirement of histone deacetylase1 (HDAC1) in signal transducer and activator of transcription 3 (STAT3) nucleocytoplasmic distribution. Nucleic Acids Res. 36, 4510–4520 (2008).
Shi, Y. et al. Epigenetic regulation in cardiovascular disease: mechanisms and advances in clinical trials. Signal Transduct. Target. Ther. 7, 200 (2022).
Hegde, M., Guruprasad, K. P., Ramachandra, L., Satyamoorthy, K. & Joshi, M. B. Interleukin-6-mediated epigenetic control of the VEGFR2 gene induces disorganized angiogenesis in human breast tumors. J. Biol. Chem. 295, 12086–12098 (2020).
Wei, J., Sun, Z., Chen, Q. & Gu, J. Serum deprivation induced apoptosis in macrophage is mediated by autocrine secretion of type I IFNs. Apoptosis 11, 545–554 (2006).
Lee, A. W.-M. & States, D. J. Colony-stimulating factor-1 requires PI3-kinase-mediated metabolism for proliferation and survival in myeloid cells. Cell Death Differ. 13, 1900–1914 (2006).
Linton, M. F., Moslehi, J. J. & Babaev, V. R. Akt signaling in macrophage polarization, survival, and atherosclerosis. Int. J. Mol. Sci. 20, 2703 (2019).
Tap, W. D. et al. Pexidartinib versus placebo for advanced tenosynovial giant cell tumour (ENLIVEN): a randomised phase 3 trial. Lancet 394, 478–487 (2019).
Di Gregoli, K. & Johnson, J. L. Role of colony-stimulating factors in atherosclerosis. Curr. Opin. Lipidol. 23, 412–421 (2012).
Singhal, A. & Subramanian, M. Colony stimulating factors (CSFs): complex roles in atherosclerosis. Cytokine 122, 154190 (2019).
Verma, N. et al. TET proteins safeguard bivalent promoters from de novo methylation in human embryonic stem cells. Nat. Genet. 50, 83–95 (2018).
Kotini, A. G. et al. Functional analysis of a chromosomal deletion associated with myelodysplastic syndromes using isogenic human induced pluripotent stem cells. Nat. Biotechnol. 33, 646–655 (2015).
Ancuta, P. et al. Transcriptional profiling reveals developmental relationship and distinct biological functions of CD16+ and CD16− monocyte subsets. BMC Genomics 10, 403 (2009).
Rohde, D. et al. Bone marrow endothelial dysfunction promotes myeloid cell expansion in cardiovascular disease. Nat. Cardiovasc. Res. 1, 28–44 (2022).
Randolph, G. J. Mechanisms that regulate macrophage burden in atherosclerosis. Circ. Res. 114, 1757–1771 (2014).
Li, Z.-Y., Howarth, S. P. S., Tang, T. & Gillard, J. H. How critical is fibrous cap thickness to carotid plaque stability? A flow–plaque interaction model. Stroke 37, 1195–1199 (2006).
Schuett, H. et al. Transsignaling of interleukin-6 crucially contributes to atherosclerosis in mice. Arterioscler. Thromb. Vasc. Biol. 32, 281–290 (2012).
Akita, K. et al. An interleukin-6 receptor antibody suppresses atherosclerosis in atherogenic mice. Front. Cardiovasc. Med. 4, 84 (2017).
Kim, P. G. et al. Dnmt3a-mutated clonal hematopoiesis promotes osteoporosis. J. Exp. Med. 218, e20211872 (2021).
Agrawal, M. et al. TET2-mutant clonal hematopoiesis and risk of gout. Blood 140, 1094–1103 (2022).
Giles, J. T. et al. Cardiovascular safety of tocilizumab versus etanercept in rheumatoid arthritis: a randomized controlled trial. Arthritis Rheumatol. 72, 31–40 (2020).
Gierens, H. et al. Interleukin-6 stimulates LDL receptor gene expression via activation of sterol-responsive and Sp1 binding elements. Arterioscler. Thromb. Vasc. Biol. 20, 1777–1783 (2000).
Dawoud, A. A. Z., Gilbert, R. D., Tapper, W. J. & Cross, N. C. P. Clonal myelopoiesis promotes adverse outcomes in chronic kidney disease. Leukemia 36, 507–515 (2022).
Kotini, A. G. et al. Stage-specific human induced pluripotent stem cells map the progression of myeloid transformation to transplantable leukemia. Cell Stem Cell 20, 315–328 (2017).
Wang, W. et al. Macrophage inflammation, erythrophagocytosis, and accelerated atherosclerosis in Jak2V617F mice. Circ. Res. 123, e35–e47 (2018).
Heimlich, J. B. et al. Mutated cells mediate distinct inflammatory responses in clonal hematopoiesis. Preprint at bioRxiv https://doi.org/10.1101/2022.12.01.518580 (2022).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Acknowledgements
This work was supported by grants from the National Institutes of Health (nos. HL-118567 and HL-148071 to N.W.; no. HL155431 to A.R.T.) and the Leducq Foundation (no. TNE-18CVD04 to A.R.T.). P.L. receives funding support from the National Heart, Lung, and Blood Institute (nos. 1R01HL134892 and 1R01HL163099-01), the RRM Charitable Fund and the Simard Fund.
Author information
Authors and Affiliations
Contributions
W.L. designed and performed the experiments, analyzed the data and wrote the manuscript. M.Y. performed the experiments and analyzed the data. I.F.M. and P.S.V. provided the mouse bone marrow for the transplantation experiments and wrote the manuscript. M.O. and E.P.P. designed and conducted the experiments related to human ESCs and iPSC-derived macrophages. P.B.A. performed the experiments and analyzed the data. J.B.H. and A.G.B. provided the human TET2 CH RNA-seq data. R.W. and T.X. performed the experiments and analyzed the data. R.L., E.P.P. and I.T. shared the reagents, designed the experiments and provided scientific feedback about the manuscript. N.W. and A.R.T. designed the experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.R.T. is a consultant for Amgen, CSL Behring, AstraZeneca and Foresite Laboratories, and is on the scientific advisory board of Staten Biotechnology, Fortico Biotech and Beren Therapeutics. P.L. is an unpaid consultant to, or involved in clinical trials for Amgen, AstraZeneca, Baim Institute, Beren Therapeutics, Esperion Therapeutics, Genentech, Kancera, Kowa Pharmaceuticals, MedImmune, Merck, Moderna, Novo Nordisk, Novartis, Pfizer and Sanofi Regeneron. P.L. is a member of the scientific advisory board for Amgen, Caristo Diagnostics, Cartesian Therapeutics, CSL Behring, DalCor Pharmaceuticals, Dewpoint Therapeutics, Elucid Bioimaging, Kancera, Kowa Pharmaceuticals, Olatec Therapeutics, MedImmune, Novartis, PlaqueTec, TenSixteen Bio, Soley Thereapeutics and XBiotech. His laboratory has received research funding in the last 2 years from Novartis, Novo Nordisk and Genentech. P.L. is on the board of directors of XBiotech. He has a financial interest in XBiotech, a company developing therapeutic human antibodies, in TenSixteen Bio, a company targeting somatic mosaicism and CHIP to discover and develop new therapeutics to treat age-related diseases, and in Soley Therapeutics, a biotechnology company that is combining artificial intelligence with molecular and cellular response detection to discover and develop new drugs, currently focusing on cancer therapeutics. His interests were reviewed and are managed by Brigham and Women’s Hospital and Mass General Brigham in accordance with their conflict-of-interest policies. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Kathryn Moore, Siddhartha Jaiswal and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
a. Ldlr−/− male mice were transplanted with WT or Tet2KO male mice bone marrow; after 6 weeks reconstitution, WT→Ldlr−/− (n = 9) and Tet2KO→Ldlr−/− (n = 10) mice were fed a western diet for 12 weeks. Bar graph shows total lesion area in the aortic root. Two-tailed unpaired t-test. b-c. WT and Tet2KO macrophage were pretreated with or without IL-1β (100 μg/ml) blocking antibodies for 1 hr, then treated with LPS (20 ng/ml) for 4 hours followed by nigericin (5 μg/ml) for another 30 mins. Secreted IL-1β and IL-6 were determined by ELISA. n = 5 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. d. Bone marrow derived macrophages were pre-incubated with IL-6R blocking antibodies (100 μg/ml) for 1 hour and then challenged with or without IL-6 (25 ng/ml). qPCR was used to measure the mRNA level of Socs3 expression. n = 3 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. e-g. Peripheral blood white blood cells counts, RBC counts and platelet counts after 12 weeks western diet. n = 15 mice per group. One-way ANOVA followed by Tukey’s multiple-comparison test. h-l. Body weight, plasma cholesterol and triglycerides, liver/body weight ratio, spleen/body weight ratio after 12 weeks western diet. m. Immunoblot of isolated splenic monocytes and neutrophil Caspase1, cleaved IL-1β and β-actin after 12 weeks western diet. n = 6 mice per group. One-way ANOVA followed by Tukey’s multiple-comparison test or two-tailed unpaired t-test. n. Immunofluorescence staining of macrophage (anti-Mac2, Green), TREM2 (Red) in aortic roots and quantification of TREM2+Mac2+ cells as the percentage of total Mac2+ cells per lesion area in aortic root cross-sections. Scale Bar, 50μm. n = 15 mice per group. One-way ANOVA followed by Tukey’s multiple-comparison test. o. Immunofluorescence staining of macrophage (anti-Mac2, Green), IL-1β expression (Red) in aortic roots and quantification of IL-1β mean fluorescence intensity per lesion area in aortic root cross-sections. Scale Bar, 50μm. n = 15 mice per group. One-way ANOVA followed by Tukey’s multiple-comparison test. p. Quantification of CSF1R expression in monocytes of Ctrl and Tet2+/− mice. n = 5 mice in ctrl group. n = 6 mice in Tet2 + /- group. Two-tailed unpaired t-test. All the Data are Mean ± SEM.
Extended Data Fig. 2
a. Gating strategy for LSK cells and Hematopoietic progenitor cells (HPC, the first row), LSK (Sca1+c-Kit+) subpopulation (middle row) and HPC (Sca1−c-Kit+) subpopulation (the third row). For the LSK subpopulation, CD150+CD48−, CD150−CD48−, CD150−CD48+ LSK fractions are long term HSC (LT-HSC), short term HSC (ST-HSC) and multipotent progenitors (MPP) respectively. For the HPC subpopulation, the cells are defined as common myeloid progenitors (Lin−Sca1−c-Kit+CD34+CD16/32int; CMP), granulocyte and macrophage progenitors (Lin−Sca1−c-Kit+CD34+CD16/32hi, GMP). b. Hematopoietic stem and progenitor cells in BM. n = 12 mice per group. One-way ANOVA followed by Tukey’s multiple-comparison test. c-h. CSF1R expression on stem and progenitor cells as determined by flow cytometry shown as mean fluorescence intensity and percentage of positive cells. n = 12 mice per group. One-way ANOVA followed by Tukey’s multiple-comparison test. All the Data are Mean ± SEM.
Extended Data Fig. 3
a. Immunoblot and quantification of p-STAT3/STAT3 with or without IL-6 (25 ng/ml). n = 3 independent experiments. Two-tailed unpaired t-test. b. BMDMs were pretreated with trichostatin A (TSA, 100 nM) or DMSO for 1 hour and then incubated with IL-6 (25 ng/ml) for 2 hours. The level of nuclear STAT3 acetylation was determined by western blot. n = 3 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. c. BMDMs were incubated with IL-6 (25 ng/ml) for indicated time. Methylation levels of the STAT3 binding site in the Csf1r promoter region of macrophages were analyzed. n = 3 independent experiments. Two-tailed unpaired t-test. d. Immunoblot and quantification of DNMT1, DNMT3A and DNMT3B expression in BMDMs with or without IL-6 (25 ng/ml) treatment for 4 hours. n = 5 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. e. BMDMs were pretreated with 5-Azacytidine (5-aza, 1 μM) or DMSO for 1 hour and then incubated with or without IL-6 (25 ng/ml) for 8 hours. mRNA level of Csf1r was tested by qPCR. n = 4 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. f. BMDMs were pretreated with 5-aza (1 μM) or DMSO for 1 hour and then incubated with IL-6 (25 ng/ml) for 4 hours. STAT3 binds to the Csf1r promoter area was revealed by ChIP-qPCR assay. Shown is percentage of input. n = 4 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. g. BMDMs were pretreated with TSA (100 nM) or DMSO for 1 hour and then incubated with (25 ng/ml) for 4 hours. Methylation levels of the STAT3 binding site in the Csf1r promoter region of macrophages. n = 5 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. All the Data are Mean ± SEM.
Extended Data Fig. 4
Immunoblot of CSF1R and cleaved Caspase 3 in macrophage with or without IL-6 (25 ng/ml) treatment for 48 hours. Bar graph shows the quantification of immunoblots. n = 5 mice per group. One-way ANOVA followed by Tukey’s multiple-comparison test. All the Data are Mean ± SEM.
Extended Data Fig. 5
Ctrl and Tet2CH mice were fed a western diet for 7 weeks followed by another 5 weeks with or without PLX3397 (200 mg/kg) formulated WD feeding. a. Food intake per day per mouse during 5 weeks PLX3397 WD feeding period. n = 3 independent cages per group. One-way ANOVA followed by Tukey’s multiple-comparison test. b. Plasma cholesterol levels. n = 11 mice in Ctrl+Control group, n = 12 mice in Tet2CH+Control group, n = 13 mice in Ctrl+PLX3397 group, n = 13 mice in Tet2CH + PLX3397 group. One-way ANOVA followed by Tukey’s multiple-comparison test. c-f. Body weight, liver/body weight ratio, spleen/body weight ratio and peripheral blood white blood cells counts, RBC counts and platelet counts after 12 weeks western diet. n = 11 mice in Ctrl+Control group, n = 12 mice in Tet2CH+Control group, n = 13 mice in Ctrl+PLX3397 group, n = 13 mice in Tet2CH + PLX3397 group. One-way ANOVA followed by Tukey’s multiple-comparison test. g. Control and Tet2KO BMDMs were treated with or without oxidized LDL (oxLDL, 50 μg/ml), acetylated LDL (acLDL, 50 μg/ml) or cholesterol crystal (CHO-C,100 μg/ml) overnight and then challenged with LPS (20 ng/ml) for 4 hours with fresh media. Secreted IL-6 level was quantified by ELISA. n = 4 independent experiments. Two-tailed unpaired t-test. All the Data are Mean ± SEM.
Extended Data Fig. 6
a. Flow cytometric analysis of cell surface molecular expression on human embryonic stem cell-derived TET2 deficient macrophages and its isogenic control in day 24. n = 3 independent experiments. Two-tailed unpaired t-test. b. Strategy for gating and confirmation of induced pluripotent stem cell (iPSC) with TET2 haploinsufficiency (TET+/−) and isogenic control for the progenitors (CD34+) and lineage-committed (CD34 + CD45+) cells differentiation on Day1. c. Control and TET2+/− iPSCs were treated with or without hIL-6 (25 ng/ml) for 48 hours. CSF1R expression was analyzed by flow cytometry. n = 3 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. d. Flow cytometric analysis of cell surface molecular expression on iPSC derived TET+/− and isogenic control macrophage after 24 days of differentiation. n = 3 independent experiments. Two-tailed unpaired t-test. e. Control and TET2+/− iPSC-derived macrophages were treated with or without hIL-6 (25 ng/ml) for 48 hours. CSF1R expression was quantified by flow cytometry. n = 3 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. f. Immunoblot and quantification of CSF1R and cleaved Caspase 3 in control and TET2+/− iPSC-derived macrophages with or without hIL-6 (25 ng/ml) treatment for 48 hours. n = 3 independent experiments. One-way ANOVA followed by Tukey’s multiple-comparison test. g. TET2+/− and its isogenc control iPSCs were treated with or without oxidized LDL (oxLDL, 50 μg/ml), acetylated LDL (acLDL, 50 μg/ml) or cholesterol crystal (CHO-C,100 μg/ml) overnight and then challenged with LPS (20 ng/ml) for 4 hours with fresh media. Secreted IL-6 level was quantified by ELISA. n = 4 independent experiments. Two-tailed unpaired t-test. All the Data are Mean ± SEM.
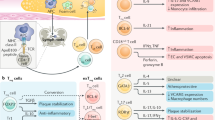
Extended Data Fig. 7 A schematic summary of the main findings.
The schema illustrating proposed mechanisms underlying the benefit of IL-6R or CSF1R inhibition in Tet2 CH-accelerated atherosclerosis.
Supplementary information
Source data
Source Data Figs. 2, 3 and 5 and Extended Data Figs. 1, 3, 4 and 6.
Uncropped immunoblots for Figs. 2, 3 and 5 and Extended Data Figs. 1, 3, 4 and 6.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, W., Yalcinkaya, M., Maestre, I.F. et al. Blockade of IL-6 signaling alleviates atherosclerosis in Tet2-deficient clonal hematopoiesis. Nat Cardiovasc Res 2, 572–586 (2023). https://doi.org/10.1038/s44161-023-00281-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-023-00281-3
This article is cited by
-
Herz und Blut: klonale Hämatopoese
Herz (2024)
-
IL-6R antagonism ameliorates atherosclerosis linked to Tet2-related clonal haematopoiesis
Nature Reviews Cardiology (2023)