Abstract
The enzyme L-asparaginase (L-ASNase) is used in the treatment of Acute Lymphoblastic Leukemia. The preparations of this enzyme for clinical use are derived from bacterial sources and its use is associated with serious adverse reactions. In this context, it is important to find new sources of L-ASNase. In this work, the Placket-Burman Experimental Design (PBD) was used to determine the influence of the variables on the L-ASNase production then it was followed by a 28–4 Factorial Fractional Design (FFD). The results obtained from PBD have shown a range of L-ASNase activity, from 0.47 to 1.77 U/gcell and the results obtained from FFD have showed a range of L-ASNase activity, from 1.10 to 2.36 U/gcell. L-proline and ammonium sulfate were identified as of significant positive variables on this production enzyme by Penicillium cerradense sp. nov. The precise identification of this new species was confirmed by morphological characteristics and sequence comparisons of the nuclear 18S-5.8S-28S partial nrDNA including the ITS1 and ITS2 regions, RNA polymerase II, β-tubulin and calmodulin genomic regions. The genetic sequence coding for the L-ASNase was obtained after carrying out a full genome sequencing. The L-ASNase expressed by P. cerradense sp. nov may have promising antineoplastic properties.
Similar content being viewed by others
Introduction
L-Asparaginase (L-ASNase) is an enzyme used for the treatment of acute lymphoblastic leukemia (ALL). In the human body this enzyme (L-asparagine amidohydrolase, EC 3.5.1.1) is responsible for the selective catalysis of the asparagine hydrolysis reaction in aspartic acid and ammonia1. Neoplasic cells cannot synthesize L-asparagine unlike normal cells due to the low expression or absence of the L-asparagine synthetase gene, therefore they obtain the required asparagine from circulating pools2. Acute lymphoblastic leukemia (ALL) is the most common cancer in childhood, with a prevalence up to 25% of cancers in children who are under the age of 15 years3.
Asparaginase is listed in the 21st WHO List of Essential Medicines as a cytotoxic and adjuvant medicine for acute lymphoblastic leukemia4.
However, important adverse reactions and toxicity associated with the use of an enzyme from prokaryote are observed5, including anaphylaxis, coagulation disorders, neurological crises, hyperglycemia, hepatotoxicity, leukopenia, pancreatitis and thrombosis6,7. Another factor that contributes to asparaginase-associated toxic side effects is its glutaminase activity8. Among the reported asparaginase formulations, there are asparaginases with undetected glutaminase activity, others with low to moderate activity, and some others with augmented glutaminase activities. Of the three asparaginases licensed by the US Food and Drug Administration, all of which are fermentation products, E. coli asparaginases have relatively low glutaminase activity, while Erwinia asparaginase has a higher glutaminase moiety, approximately tenfold higher than that of E. coli and, therefore, KM and VMAX more favorable for deamination of glutamine6.
Considering the importance of clinical application and the severe side effects presented as a result of the treatment, the search for alternative sources of L-asparaginase is of relevant interest for the development of new biopharmaceuticals. Microorganism are a better source than animals or plants, considering their ability to grow easily on rather simple and inexpensive substrates9. Nowadays, new L-asparaginase have been identified in eukaryotic sources. Among filamentous fungi that produce this enzyme, species of the genus Aspergillus, Penicillium, Fusarium and Cladosporium have been frequently reported in the literature10,11,12. El-Hadi et al.13 studied the effect of nutritional and environmental factors using the PB design and it was concluded that K2HPO4, sucrose, and time of fermentation were the most important factors that influence L-asparaginase activity within their tested limits. Niharika and Supriya14, found that the presence of sucrose as a carbon source in the fermentation medium was an effective inducer for L-asparaginase production using F. oxysporum. Souza, et al.15 described that fungi L-asparaginase production is extremely influenced by the composition of the fermentation medium, especially carbon and nitrogen sources, and physical factors such as temperature, pH, agitation, inoculum concentration and fermentation time.
Several researchers have reported on the optimization of growth conditions, such as carbon and nitrogen sources, pH or temperature for L-asparaginase production, however no specified medium has been established for optimum enzyme production, once each organism has its own particular conditions for maximum enzymatic production15 and this is important to development an economically viable process.
Some statistical experimental design methods have been employed in bioprocess optimization of asparaginase production from fungi. Making it possible to study the influence of individual factors, and the interaction between them, which can be enable the optimization of the process.
In this work, the morpho-molecular characterization of a new species of Penicillium isolated from the soil of the Cerrado (Brazilian Savannah) was performed and the production of L-asparaginase enzyme by this fungus was evaluated using submerse fermentation. The gene encoding the L-asparaginase of this species was also characterized.
Results
Two isolates of Penicillium were obtained from the soil of the Cerrado for morphological and molecular characterization. The amplification and sequencing of the partial rDNA (including the ITS), RPB2, β-tubulin and calmodulin regions revealed sequences of ca. 1.200, 800, 720, and 570 bp, respectively. The ITS and RPB2 sequences were used in the multigenic species identification. The β-tubulin and calmodulin gene sequences were not included in the phylogenetic analysis because these regions were unavailable for most of the previously described Penicillium species. The new sequences were deposited in GenBank under accession numbers MT006126, MT006127, MT416532 to MT416537. No topological conflicts were found among the phylogenetic trees based on each of the two partial genomic regions, and therefore, the data sets were concatenated (single gene trees are available in TreeBASE). For the multilocus analysis, 86 taxa were used (Table Supplementary S1), with alignments of RPB2 and ITS having 915 and 586 bp in length, respectively. The concatenate alignment (1501 bp) showed 891 conserved characters, 589 variable, and 513 phylogenetically informative sites. The GTR + I + G model was selected for RPB2 and ITS.
Based on morphological and molecular comparisons, a new species of Penicillium belonging to the section Citrina is proposed in this work (Fig. 1).
Bayesian phylogenetic tree based on concatenate sequences (ITS and RPB2) of Penicillium species section Citrina. Bayesian posterior probabilities values are indicated at the nodes and thick lines indicate posterior probability greater than or equal to 0.99. The isolates in this study are highlighted in bold. The tree was rooted with Coccidioides immitis CBS 14656. The specimens in this study are highlighted in bold.
Taxonomy
Penicillium cerradense sp. nov. Cruvinel, Magalhães, P. O., Pinho. (Fig. 2).
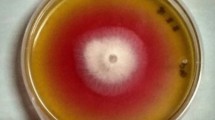
Penicillium cerradense sp. nov. (A) Colony appearance (surface and reverse) after 7 days of growth on malt extract agar at 25 ± 2 °C. (B–D) Conidiophores. (E, F) Conidiogenic apparatus with ampulliform phialides. (G) Conidia. (H) Conidia germinating after 48 h. (I) Sclerotia. Scale Bar: B‒H = 10 μm, I = 50 μm.
MycoBank: MB 835,241.
GenBank: ITS = MT006126, RPB2 = MT416532, TUB = MT416533, CAL = MT416534, L-ASNase = MT742156.
Systematic position: Ascomycota, Pezizomycotina, Eurotiomycetes, Eurotiomycetidae, Eurotiales, Aspergillaceae.
Type: —BRAZIL, Goiás, Água Fria de Goiás. 14°58′10.21′′S, 48°1′19.43″W, on soil of Cerrado, 30 January 2009, coll. F. G. Siqueira (holotype in dried culture UB23977, ex-type DCFS6a).
Etymology: —This species refers to Cerrado.
Colonies of this fungus have grown slowly in MEA culture medium (15 mm), PDA (30 mm), and SDA (37 mm) in 7 days, without aerial and superficial mycelium formation. In MEA the colonies were light green or greyish-green, velvety, white or absent colony edges and light-brown in reverse (Fig. 2A). Conidiophores were formed abundantly, solitary, erect, hyaline, emerging from hypha, consisting of a stipe followed by a penicillate conidiogenic apparatus; hyaline and smooth stipe with 25.0‒240.0 × 2.0‒3.0 μm (average 85.0 × 2.5 μm; Fig. 2B‒D). The penicillated conidiogenic apparatus measured 7.0‒16.0 μm in length and 4.0‒14.0 μm in width. Conidiophore were predominantly monoverticillate, when biverticillate presented 2 branches with 10.0‒20.0 × 2.0‒3.0 μm (average 13 × 2.5 μm), each branch finished in the production of 2‒4 (− 8) phialides. Ampulliform phialides was predominantly in four, hyaline, aseptate, 5.0‒8.0 × 2.0‒3.0 μm (average 7.0 × 2.5 μm; Fig. 2E‒F). Conidia catenulate, subglobose or strongly ellipsoidal, hyaline, smooth with 1.5‒3.0 × 2.0‒3.0 μm (average 2.5 × 2.5 μm; Fig. 2G‒H). Globose sclerotia, solitary to abundant, superficial or immersed in the culture medium, pigmented, pale brownish to brownish with 150.0‒340.0 × 130.0‒320.0 μm (average 250 × 200 μm; Fig. 2I). Sexual form not observed.
In PDA, the colony was greyish-green with a well-defined white border and light-brown roughness in reverse. Finally, in SDA the colony was greyish-green, light gray or brown coloration with white border and light-brown with intense roughness in reverse (Fig. 3).
Additional specimen examined: BRAZIL, Goiás, Água Fria de Goiás. 14°58′10.21″S, 48°1′19.43″W, on soil of Cerrado, 30 January 2009, coll. F. G. Siqueira (culture DCFS6b).
Notes: —The new species groups in a distinct clade with P. sumatrense. Penicillium cerradense sp. nov. is phylogenetically close but clearly distinct of the P. sumatrense. The new species has conidiophores predominantly monoverticillate or biverticillate with two branches, abundant sclerotia, smaller stipe (85 μm) and phialides (5.0‒8.0 × 2.0‒3.0 μm), while P. sumatrense has conidiophores with 3‒6 branches, absent sclerotia, larger stipe (up to 200 μm) and phialides (8.0‒10 × 2.0‒3.5 μm)16.
L-asparaginase gene
The L-asparaginase gene was obtained from the whole genome sequencing of Penicillium cerradense isolate DCFS6a. The L-asparaginase gene was 1251 bp in length, and its nucleotides sequence was analyzed using the Clustal Omega and NCBI's BLAST programs. The nucleotides and amino acids sequences of Penicillium cerradense showed homology to asparaginase genes derived from Penicillium sizovae, Aspergillus niger and Aspergillus ibericus (Fig. Supplementary S1), among other species. Based on amino acid sequences alignment, it was markedly different from other previously reported Penicillium spp. derived-L-asparaginases. It is noteworthy that in the analysis using the Clustal Omega and NCBI's BLAST programs, homology to L-asparaginase genes was found only in this unique sequence presented in this work, thus an enzymatic activity reported in this manuscript is encoded by the only gene identified for L-asparaginase in the P. cerradense sp. nov.
Identifying significant variables affecting L-asparaginase production by
Penicillium cerradense sp. nov. using statistical design
The PBD provides initial indications of how each variable tends to influence the L-asparaginase production17 and it is convenient especially when facing large number of factors that can potentially influence optimal or near optimum responses18. This design is recommended when more than eight factors are under investigation18. This model describes interaction among factors, and it is used to screen and evaluate the important factors that influence asparaginase production. The experiment was conducted to study the effect of each selected variable on the production of L-asparaginase. The design matrix selected for the screening of significant variables for L-asparaginase production and the corresponding response (Y) under culture medium conditions are shown in Table 1. The results obtained from PBD have shown a wide range of L-asparaginase activity, from 0.47 to 1.77 U/gcell. The maximum L-asparaginase activity was achieved on run number 9 with culture medium containing L-asparagine 3.0%, L-proline 3.0%, urea 0.1%, sodium nitrate 2.5%, yeast extract 0.1%, ammonium sulfate 1.5%, peptone 2.0%, glucose 0.2%, sucrose 0.2%, malt extract 0.5% and potassium chloride 0.01%.
The relationship between a set of independent variables and the response (Y) is determined by a mathematical model called the multiple regression model. The determination of the main effects was performed, and the results are presented in Table 2. Eight out of the eleven variables tested (L-asparagine, L-proline, urea, yeast extract, ammonium sulfate, peptone, glucose and sucrose), showed positive effect and improved L-asparaginase production, whereas variables such as sodium nitrate and potassium chloride decreased L-asparaginase activity. The significant variables (p < 0.1) for L-asparaginase activity were identified as L-proline (p = 0.0395) and potassium chloride (p = 0.0768).
The experiment was conducted for the eight variables which presented positive effect. The results obtained from FFD showed a wide range of L-asparaginase activity, from 1.10 to 2.36 U/gcell. The maximum L-asparaginase activity was achieved in run number 10 with culture medium containing L-proline 5.0%, peptone 2.0%, sucrose 0.2%, urea 2.0%, ammonium sulfate 2.0%, yeast extract 1.5%, glucose 1.0%, L-asparagine 3.0% (Table 3). Statistical analysis of the response was performed, and the results are presented in Table 4. The variables with positive effect such as L-proline, ammonium sulfate and L-asparagine induced the highest level of L-asparaginase production, whereas the variables with negative effect (peptone, sucrose, urea, yeast extract and glucose) decreased the activity. The significant variable (p < 0.05) identified was ammonium sulfate (p = 0.02). After 20 runs, L-asparaginase activity was calculated using the proposed model equation: Prediction L-asparaginase activity = 1.92 + 0.00375X1 − 0.045X2 − 0.31X3 − 0.042X4 + 0.29X5 − 0.15X6 − 0.17X7 + 0.04X8. Level of significance of 95%. R2 = 0.5894 and predict R2 = − 0.31. A negative predict R2 implies that the overall mean is a better predictor of your response than the current model. The "Lack of Fit t-value" of 2.58 implies the Lack of Fit is significant. After the analyses of the ANOVA results, the next steps of the study were carried out using one variable at a time.
L-proline and ammonium sulfate were identified as of significant positive variables on the production of L-asparaginase by Penicillium cerradense. The effect of L-proline in different concentrations is represented in Fig. 4. The L-proline substrate concentration of 9% differs significantly from 3 and 7%. The enzymatic activity at 9% corresponds to approximately 1.6 times greater than the one found in lower concentration. In the medium with 7% concentration the morphology of the grown biomass is different from the others at other concentrations. The fungus grew slowly and the final biomass showed a lower yield. The observed decrease in enzyme activity from 5 to 7% of substrate concentration could be related to this morphological difference in the growth of P. cerradense sp. nov. El-Enshasy et al.19 identified the correlation of protein production capacity with morphological differences in the growth of Aspergillus niger in submerged culture medium. In this study, while evaluating the effect of L-proline in concentrations of 15% and 20%, a reduction in L-asparaginase activity was observed.
Effect of L-proline on L-asparaginase activity by Penicillium cerradense cultivated for 4 days at 30 °C. The results are presented as mean of enzyme activities with standard deviation for each run (n = 9). *9% ≠ 3% and 7% of the proline concentration (p < 0.05). 7% vs 9% (p = 0.0004); 3% vs 9% (p < 0.0001).
The same strategy used to verify the effect of L-proline substrate on the L-asparaginase activity was used to identify the effect of substrate ammonium sulfate in enzyme production. The effect of ammonium sulfate in different concentrations is represented in Fig. 5. The analyses were performed in triplicate and the results are presented as mean of enzyme activities with standard deviation for each run. The higher enzymatic activity was observed at 7% ammonium sulfate concentration. The effect of ammonium sulfate in concentrations of 10% and 15% suggested stabilization or decrease in L-asparaginase activity. However, as per as statistical analysis, no significant difference in L-asparaginase activities was identified for the different ammonium sulfate concentrations used in this study.
Discussion
Filamentous fungi are producers of a range of primary and secondary metabolites; therefore, they are commercially exploited as cell sources for the production of a wide variety of enzymes. These high productivity characteristics of filamentous fungi are related to their abilities to grow at high rates and to high biomass densities supported by low-cost substrates in simple fermentation process. Several filamentous fungi have been described as producers of L-asparaginase. Among species that produce this enzyme, the genus Aspergillus, Penicillium, Fusarium and Cladosporium have been frequently reported in the literature7,10,11. Some species of the genus Penicillium have been reported as producing L-asparaginase: Penicillium aculeatum, P. brevicompactum, P. chrysogenum, P. citrinum, P. claviforme, P. cyclopium, P. digitatum, P. expansum, P. granulatum, P. nelicum, P. nigricans, P. olsonii, P. simplicissimum, P. urticae10,11,20,21,22,23. Souza et al.15 reported that only two species have been deposited in a culture collection center, A. terreus MTCC 1782 strain and R. miehei CAU432. The molecular identification of a fungus was mentioned in one study, which species was identified based on 18S rRNA sequence analysis.
The production of L-asparaginase by the fungus Penicillium cerradense, which was identified in this study as a new fungus species belonging to the genus Penicillium, represents an unprecedented work.
Besides that, the analysis of the results obtained by PBD and FFD made it possible to identify L-proline and ammonium sulfate substrates as significant independent variables with positive effect on L-asparaginase production by Penicillium cerradense. In similar studies investigating the culture medium components with filamentous fungi A. terreus, the variables L-proline and ammonium sulfate also had positive effects on L-asparaginase enzyme production after optimization of culture media24. Dias and Sato25, working with another species of the genus Aspergillus built a Plackett–Burman design for selection of significant variables and obtained a result where temperature, inoculum concentration, and pH of the culture medium presented a significant and positive effect for the proposed model.
In a strategy for optimizing L-asparaginase production by P. cyclopium applying experimental design, during the initial design phase it was identified that ammonium sulfate had a significant positive effect, in addition to the variables MgSO4.7H2O and KCl26. In a study with A. terreus27, in the analysis of the effects of nitrogen sources on the production of L-asparaginase, higher enzymatic activities were identified in the presence of ammonium sulfate substrate in solid culture medium, and the largest yield was obtained by supplementing sucrose (1%), ammonium sulfate (1%), NaCl (1%) and L-asparagine (1%). It was observed that L-proline and ammonium sulfate positively affect L-asparaginase production, however enzyme activity is inhibited at high concentrations of these substrates. In an optimization study of L-asparaginase production by Saccharomyces cerevisiae, Lang et al. (1997) verified the amount of nitrogen source in the medium and discussed the effect of undesirable increase of ammonium ions in cellular metabolism. The excess of ammonia molecules can lead to catabolic nitrogen repression in cells28.
Among the statistical designs found in the literature, L-asparagine and L-proline were the amino acids most used as nitrogen source as well as substrate inducer for asparaginase production, these results could be founded in the studies12,25,26,29,30,31,32,33.
Phylogenetic analyses combined with morphological comparisons revealed a new species of Penicillium, P. cerradense sp. nov., isolated from the soil of the Cerrado and with potential for L-asparaginase production. The L-asparaginase gene was sequenced and this identification in eukaryotic sources is important in an effort to find new biopharmaceuticals with fewer side effects for leukemia treatment. The substrates L-proline and ammonium sulfate were found to have positive effects on the production of L-asparaginase by the fungus.
Methods
Fungal strain
Based on work previously developed by the research group at the Laboratory of Quality Control and Natural Products—Faculty of Health Sciences, University of Brasília, Brasília, Brazil—the production of L-Asparaginase by filamentous fungi obtained from soil and plant species of the Cerrado biome was addressed. A former screening of fungi revealed that the isolates addressed in this study have shown potential to produce L-asparaginase (personal communication).
The two isolates were obtained from soil samples of the Cerrado (14°58′10.21″S, 48°1′19.43″W) collected in Água Fria de Goiás city, Goiás state, Brazil. A hyphal-tip culture of each isolate was obtained on Potato Dextrose Agar (PDA) and were deposited within the scope of the SisBiota Brasil (National System of Research in Biodiversity—CNPq) of filamentous fungi with authorization to access or send a sample of the genetic heritage component number 010770/2013-5 and access authorization by the National Genetic Heritage Management System and The Traditional Knowledge Associated Genetic Heritage Management Council in compliance with the provisions of Law No. 13,123/2015 and its regulations (Registration number: AEFBB51 Pérola de Oliveira Magalhães Dias Batista). The strains are maintained at Laboratory of Enzymology of the Institute of Biological Sciences at University of Brasília, Brazil, and were donated by Professor Edivaldo Ximenes Ferreira Filho. The isolates were stored at − 80 °C for later use.
DNA extraction, PCR amplification and sequencing
Genomic DNA was extracted from a pure culture originally grown on PDA at 25 ± 1ºC for 7 days. Fungal mycelium was scraped from colony margins and DNA was extracted using the Wizard Genomic DNA Purification Kit (Promega, Madison, WI, USA). Polymerase Chain Reaction (PCR) was performed with partial sequences of the nuclear 18S-5.8S-28S partial nrDNA, including the ITS1 and ITS2 regions (ITS), with the primers V9G34 and LR535; RNA polymerase II subunit 2 (RPB2) with the primers 5F236 and 7Cr37; β-tubulin (TUB) with the primers T138 and Bt2b39, and calmodulin (CAL) using primers Cal228F40 and Cal2Rd41. Amplification was performed with an initial denaturation of 95ºC for 1 min and 30 s, followed by 35 cycles of 95ºC for 20 s, annealing at 53ºC (for ITS), 54ºC (RPB2), 55ºC (β-tubulin) or 59ºC (calmodulin) for 45 s, initial extension at 72ºC for 45 s and at 72ºC for 5 min on final extension. PCR products were analyzed in 1% agarose electrophoresis gels stained with GelRed (Biotium Inc., Hayward, CA, USA) in a TAE 1X buffer and visualized under UV light to check for amplification size and purity. The PCR products were purified and sequenced by Macrogen (South Korea). For whole genome sequencing, total genomic DNA was extracted from mycelia grown under submerged fermentation for 3 days at 30 °C. DNA concentration and quality were determined by Nanodrop ND-1000 spectrometer and agarose electrophoresis gel. The genome of isolate DCFS6a was sequenced with the Illumina HiSeq 2500 platform (StabVida, Lisbon, Portugal).
Phylogenetic analysis
The new sequences were assembled and manually edited using Geneious v.8.1.9 (https://www.geneious.com/). To determine the Penicillium section which they shared the highest nucleotide identity with, the partial nucleotide sequences and the BLASTn algorithm were used to search the NCBI-GenBank non-redundant nucleotide database. For species identification, a concatenate tree was reconstructed using the ITS and RPB2 sequences from deposited in the GenBank16,42,43 and from the other two isolates obtained in this study. Additionally, Coccidioides immitis CBS 14656 was used as outgroup. To test possible topological incongruences, phylogenetic trees were individually obtained from each genomic region. Multiple alignments were obtained with MAFFT v744. Finally, phylogenetic trees were reconstructed, for the concatenate data (ITS and RPB2), using Bayesian Inference (BI). The best substitution models for each partition were determined with MrModeltest45. The CIPRES web portal46 was used to run MrBayes v3.2.147. The Markov Chain Monte Carlo (MCMC) analysis was run with a total of 10 million generations, sampling every 1,000 generations. The convergence of the log likelihoods was confirmed using TRACER v1.7.143. The first 25% of the sampled trees were discarded as burn-in, with the posterior probability (PP) values calculated with the remaining trees48. The phylogenetic tree was edited in FigTree v1.447,48 (http://tree.bio.ed.ac.uk/software/figtree/) and Inkscape (www.inkscape.org).
Morphological analysis
Macroscopical characters were observed from hyphal tip cultures grown on PDA, 2% Malt Extract Agar (MEA) and Sabouraud Dextrose Agar (SDA) during seven days at 25 ± 1 °C, while microscopical characteristics were studied with the fungus grown in MEA. Microscopical characteristics were analyzed by mounting reproductive structures in clear lactoglycerol, and 30 measurements for each morphological parameter were carried out at a magnification of × 1,000 using a Leica DM2500 light microscope equipped with a Leica DFC 490 digital camera, coupled to a computer containing the Leica Qwin-Plus software.
L-asparaginase activity assay
In most of the microorganisms, L-asparaginase accumulates as an intracellular (periplasmic, cytoplasmic and membrane bound) product49. The L-asparaginase was determined by the formation of L-aspartic acid β-hydroxamate (AHA) from asparagine and hydroxylamine50 with modifications. According to this method, a periplasmic activity of L-asparaginase can be quantified directly in the whole cell without previous extraction51. For analysis, the culture media were filtrated on Whattman #2 filter paper and the cells were washed twice with buffer Tris–HCl buffer (50 mM) pH 8.6 and used as samples. The reaction mixture was: 1.5 mL Tris–HCl buffer (50 mM) pH 8.6, 0.2 mL of L-asparagine solution (100 mM), 0.2 mL hydroxylamine solution (1 M), and 0.1 g biomass. After incubation at 37 °C for 30 min in a temperature-controlled bath, the reaction was ended by the addition of 0.5 mL of ferric chloride reagent 5.0% (w/w) FeCl3, 2.5% (w/v) TCA, and 0.33 M HCl. The reaction between the AHA and FeCl3 led to a brown coloration that could be quantified by absorbance (500 nm). Sample blank was performed without the addition of L-asparagine and hydroxylamine hydrochloride solutions and all analyses were performed in triplicates. The results are presented as distribution of enzyme activities. The calibration curve was performed through multiple dilutions of a 5 mM AHA stock solution and the addition of appropriate amounts of FeCl3/TCA/HCl solution, ranging from 0.01 to 3.0 µmol of ferric AHA mL−1. A unit of L-asparaginase activity (U/gcell) corresponds to 1 µmol of AHA produced per minute per gram of sample cells.
Production culture for screening of variables
Submerse cultivation was performed in 250 mL Erlenmeyer flasks containing 50 mL of culture medium based on the design matrix together with K2HPO4 0.152%, MgSO4.7H2O 0.052%, ZnSO4.7H2O 0.001%, FeSO4.7H2O 0.001% and CuSO4.5H2O 0.052%. A 5 mm diameter disk of the mycelium of the fungus was deposited in the autoclaved media and kept in a temperature controlled orbital shaker at 30 °C and 120 rpm for 4 days. The culture media were filtrated on Whattman #2 filter paper and the cells were washed twice with buffer Tris–HCl buffer (50 mM) pH 8.6 and used as samples.
Evaluation of variables effect with statistical designs
Plackett–Burman experimental design (PBD)
The independent variables such as L-asparagine, L-proline, urea, sodium nitrate, yeast extract, ammonium sulfate, peptone, glucose, sucrose, malt extract and potassium chloride were considered to evaluate their effect on L-asparaginase production by Penicillium cerradense isolate DCFS6a. The culture media were prepared based on PBD52 given in Table 5. The different factors were prepared in two levels: (− 1) for low and (+ 1) for high level. Eleven independent variables were screened in 16 combinations plus a triplicate of the central point, totaling 19 runs organized from the type matrix 2K, K factors at 2 levels, considering the number of runs with n = 4t, where t is an integer. The matrix was built considering the minimum number of 4 trials more than the number of variables under study, allowing a degree of freedom to calculate the standard error18. To determine the significant effects, following the screening objective of the PBD, the fixed significance level was 10% (p < 0.1), minimizing the risk of excluding some important factor for the next step of the process12. However, the variables with positive effect in L-asparaginase production were fixed at a high level and those variables that had a negative effect were excluded to follow with FFD. This model (PBD) could provide us with initial indications of how each variable tends to influence L-asparaginase production, regardless the significancy of the variable18. The experimental design matrix and determination of the main effects were established by Protimiza Experimental Design software (https://experimental-design.protimiza.com.br/).
Fractional factorial design (FFD)
From the results of PBD, the independent variables with positive effect in L-asparaginase production (L-proline, peptone, sucrose, urea, ammonium sulfate, yeast extract, glucose and L-asparagine) were selected, regardless the significancy of the variable18, and the culture media were prepared to the FFD (Table 6). The different factors were prepared in two levels: (− 1) for low and (+ 1) for high level. The FFD matrix was determined in a 28–4 factorial design, generating 16 combinations of the 8 variables and 4 replicates of the central point, totaling 20 runs. To determine the significant effects, following the objective of the FFD, the fixed significance level was 5% (p < 0.05). The experimental design matrix and analysis of variance were established by Protimiza Experimental Design software (https://experimental-design.protimiza.com.br/).
Effect of the significant variables
The experiment was designed to evaluate the influence of L-proline and ammonium sulfate on the production of L-asparaginase by Penicillium cerradense isolate DCFS6a. These nutrients were identified as significant variables according to PBD and FFD, respectively. The culture medium selected for such analysis was based on the highest value of enzymatic activity obtained in PBD (L-asparagine 3.0%, urea 0.1%, sodium nitrate 2.5%, yeast extract 0.1%, peptone 1.5%, glucose 0.2%, sucrose 0.2%, malt extract 0.5% and potassium chloride 0.01%, K2HPO4 0.152%, MgSO4.7H2O 0.052%, ZnSO4.7H2O 0.001%, FeSO4.7H2O 0.001% and CuSO4.5H2O 0.052%) supplemented with different concentrations of L-proline (3%, 5%, 7%, 9%, 15% and 20%), and ammonium sulfate (1.5%, 3%, 5%, 7%, 10% and 15%). As this is an independent determination, the substrate that was not under evaluation had been kept at its lowest concentration. The effects of substrates were evaluated with 3 independent experiments with n = 3, under the same culture conditions.
The statistical analysis was performed using GraphPad Prism Version 6.01 software (GraphPad Software, La Jolla California USA, www.graphpad.com). For comparison by analysis of variance between the samples, after observing the data distribution, parametric test ANOVA followed by Tukey's multiple-comparison posttest was applied. The data were represented by mean and standard deviation (M ± SD). The significant difference was considered for the values of p < 0.05.
Data availability
Data supporting the findings of this manuscript are available from the corresponding author upon reasonable request.
References
Lima, G. M. et al. Glycosylation of L-asparaginase from E. coli through yeast expression and site-directed mutagenesis. Biochem. Eng. J. 156, 107516 (2020).
Costa, I. M. et al. Recombinant L-asparaginase 1 from Saccharomyces cerevisiae: an allosteric enzyme with antineoplastic activity. Sci. Rep. 6, 36239 (2016).
Hunger, S. P. et al. Improved survival for children and adolescents with acute lymphoblastic leukemia between 1990 and 2005: a report from the children’s oncology group. J. Clin. Oncol. 30(14), 1663–1669 (2012).
WHO. World Health Organization Model List of Essential Medicines, 21st List. World Health Organization. (2019). http://www.who.int/medicines/publications/essentialmedicines/. Accessed 27 July 2021.
Shrivastava, A., Khan, A. A., Shrivastav, A., Jain, S. K. & Singhal, P. K. Kinetic studies of L-asparaginase from Penicillium digitatum. Prep. Biochem. Biotechnol. 42, 574–581 (2012).
Avramis, V. I. Asparaginases: biochemical pharmacology and modes of drug resistance. Anticancer Res. 32, 2423–2437 (2012).
Saeed, H. et al. Molecular cloning, structural modeling and production of recombinant Aspergillus terreus asparaginase in Escherichia coli. Int. J. Biol. Macromol. 106, 1041–1051 (2018).
Warrell, R. P. Jr. et al. Phase I evaluation of succinylated Acinetobacter glutaminase-asparaginase in adults. Cancer Res. 40, 4546–4551 (1980).
Lopes, A. M. et al. Therapeutic L-asparaginase: upstream, downstream and beyond. Crit. Rev. Biotechnol. 37, 82–99 (2017).
Imada, A., Igarasi, S., Nakahama, K. & Isono, M. Asparaginase and glutaminase activities of micro-organisms. J. Gen. Microbiol. 76(1), 85–89 (1973).
Gulati, R., Saxena, R. K. & Gupta, R. A rapid plate assay for screnning L-Asparaginase producing micro-organisms. Lett. Appl. Microbiol. 24(1), 23–26 (1997).
Mohan Kumar, N. S. & Manonmani, H. K. Purification, characterization and kinetic properties of extracellular L-asparaginase produced by Cladosporium sp. World J. Microbiol. Biotechnol. 29, 577–587 (2013).
El-Hadi, A., El-Refai, H., Shafei, M., Zaki, R. & Mostafa, H. Statistical optimization of L-asparaginase production by using Fusarium solani. Egypt. Pharm. J. 6, 16–23 (2017).
Niharika, Y. C. & Supriya, S. Production of L-asparaginase by Fusarium oxysporum using submerged fermentation. Int. J. Pharm. Sci. Invent. 3, 32–40 (2014).
Souza, P. M., de Freitas, M. M., Cardoso, S. L., Pessoa, A. & Guerra, E. N. S. Magalhães PO Optimization and purification of L-asparaginase from fungi: a systematic review. Crit. Rev. Oncol. Hematol. 120, 194–202 (2017).
Houbraken, J., Frisvad, J. C. & Samson, R. A. Taxonomy of Penicillium section Citrina. Stud. Mycol. 70, 53–138 (2011).
Yu, X., Hallett, S. G., Sheppard, J. & Watson, A. K. Application of the Plackett-Burman experimental design to evaluate nutritional requirements for the production of Colletotrichum coccodes spores. Appl. Microbiol. Biotechnol. 47, 301–305 (1997).
Rodrigues, M., and Iemma, A. Planejamento de experimentos e otimização de processos, 3rh edn. (ed. Casa do Espírito Amigo Fraternidade Fé e Amor) (São Paulo, 2014).
El-Enshasy, H., Kleine, J. & Rinas, U. Agitation effects on morphology and protein productive fractions of filamentous and pelleted growth forms of recombinant Aspergillus niger. Process Biochem. 41(10), 2103–2112 (2006).
Elshafei, A. M., Hassan, M. M., Abouzeid, M. A., Mahmoud, D. A. & Elghonemy, D. H. Purification, characterization and antitumor activity of L-asparaginase from Penicillium brevicompactum NRC 829. Br. Microbiol. Res. J. 2(3), 158–174 (2012).
Alhussaini, M. S. Mycobiota of wheat flour and detection of a-amylase and L-asparaginase enzymes. J. Life Sci. 10, 1112–1122 (2013).
Yadav, N. C. & Sarkar, S. Production of L-Asparaginase by Fusarium oxysporum using submerged fermentation. Int. J. Pharm. Sci. Invent. 3(6), 32–40 (2014).
Chow, Y. Y. & Ting, A. S. Y. Endophytic L-asparaginase-producing fungi from plant associated with anticancer properties. J. Adv. Res. 6(6), 869–876 (2015).
Sarquis, M. I., Oliveira, E. M., Santos, A. S. & Costa, G. L. Production of L-asparaginase by filamentous fungi. Mem. Inst. Oswaldo Cruz 99, 489–492 (2004).
Dias, F. F. Sato HH Sequential optimization strategy for maximum l-asparaginase production from Aspergillus oryzae CCT 3940. Biocatal. Agric. Biotechnol. 6, 33–39 (2016).
El-Refai, H. A. et al. Statistical optimization of anti-leukemic enzyme L-asparaginase production by Penicillium cyclopium. Curr. Trends Biotechnol. Pharm. 8(2), 130–142 (2014).
Aparna, C. & Raju, K. J. Optimization of process parameters for l-asparaginase production by Aspergillus terreus MTCC 1782 under solid state fermentation using mixed substrate. Int. J. Eng. Technol. 4, 354–360 (2015).
Lang, C., Göllnitz, C., Popovic, M. & Stahl, U. Optimization of fungal polygalacturonase synthesis by Saccharomyces cerevisiae in fed-batch culture. Chem. Eng. J. 65(3), 219–226 (1997).
Baskar, G. & Renganathan, S. Application of Latin square design for the evaluation and screening of supplementary nitrogen source for L-asparaginase production by Aspergillus terreus MTCC 1782. Indian J. Sci. Technol. 2(12), 50–54 (2009).
Baskar, G. & Renganathan, S. Statistical screening of process variables for the production of L-asparaginase from cornflour by Aspergillus terreus MTCC 1782 in submerged fermentation. Indian J. Sci. Technol. 2(5), 45–48 (2009).
Baskar, G. & Renganathan, S. Statistical and evolutionary optimization of operating conditions for enhanced production of fungal L-asparaginase. Chem. Pap. 65(6), 798–804 (2011).
Baskar, G. & Renganathan, S. Optimization of L -asparaginase production by Aspergillus terreus MTCC 1782 using response surface methodology and artificial neural network-linked genetic algorithm. Asia Pac. J. Chem. Eng. 7(2), 212–220 (2012).
Baskar, G., Sriharini, C., Sripriya, R. & Renganathan, S. Statistical screening of supplementary nitrogen source for enhanced production of L-Asparaginase by Aspergillus Terreus 1782. Chem. Biochem. Eng. Q. 24(4), 467–472 (2010).
Hoog, G. D. & Ende, A. H. G. Molecular diagnostics of clinical strains of filamentous Basidiomycetes. Mycoses 41(5–6), 183–189 (1998).
Vilgalys, R. & Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 172, 4239–4246 (1990).
Sung, H., Sung, J., Hywel-Jones, N. L. & Spatafora, J. W. A multi-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): Identification of localized incongruence using a combinational bootstrap approach. Mol. Phylogenet. Evol. 44(3), 1204–1223 (2007).
Liu, Y. J., Whelen, S. & Hall, B. D. Phylogenetic relationships among ascomycetes: evidence from an RNA polymerase II subunit. Mol. Biol. Evol. 16(12), 1799–1808 (1999).
O’donnell, K., Cigelnik, E. & Nirenberg, H. I. Molecular systematics and phylogeography of the Gibberella fujikuroi species complex. Mycologia 28, 465–493 (1997).
Glass, N. L. & Donaldson, G. C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 61(4), 1323–1330 (1995).
Carbone, I. & Kohn, L. M. A. Method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 91(3), 553–556 (1999).
Quaedvlieg, W. et al. Zymoseptoria gen. nov.: a new genus to accommodate Septoria-like species occurring on graminicolous hosts. Persoonia 26, 57–69 (2011).
Houbraken, J. et al. Classification of Aspergillus, Penicillium, Talaromyces and related genera (Eurotiales): An overview of families, genera, subgenera, sections, series and species. Stud. Mycol. 95, 5–169 (2020).
Katoh, K. & Standley, D. M. MAFFT Multiple sequence alignment software version 7: Improvements in performace and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Posada, D. & Buckley, T. R. Model selection and model averaging in phylogenetics: advantages of Akaike information criterion and Bayesian approaches over likelihood ratio tests. Syst. Biol. 53(5), 793–808 (2004).
Miller, M.A., Pfeiffer, W. and Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the Gateway Computing Environments Workshop (GCE). New Orleans, LA, pp. 1–8 (2010).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3.2.5: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2014).
Rambaut and Drummond. Tracer MCMC trace analysis tool. http://tree.bio.ed.ac.uk/software/tracer/ (2018).
Rannala, B. & Yang, Z. Probability distribution of molecular evolutionary trees: A new method of phylogenetic inference. J. Mol. Evol. 43(3), 304–311 (1996).
Kumar, S., Dasu, V. V. & Pakshirajan, K. Localization and production of novel L-asparaginase from Pectobacterium carotovorum MTCC. Process Biochem. 45(2), 223–229 (2010).
Drainas, C., Kinghorn, J. R. & Pateman, J. A. Aspartic hydroxamate resistance and Asparaginase regulation in the fungus Aspergillus nidulans. J. Gen. Microbiol. 98(2), 493–501 (1977).
Frohwein, Y. Z., Friedman, M., Reizer, J. & Grossowicz, N. Sensitive and Rapid Assay for L-Asparaginase. Nat. New Biol. 230(13), 158–159 (1971).
Plackett, R. L. & Burman, J. P. The design of optimum multifactorial experiments. Biometrika 33(4), 305–325 (1946).
Acknowledgements
This work was supported by the Federal District Foundation (FAPDF)—process 193-0000919/2020-07 (Chamada Conjunta FAPDF E FAPESP No 01/2019), The São Paulo Research Foundation (FAPESP)—process number 2019/23620-0 and Higher Education Personnel Improvement Coordination (CAPES). The authors sincerely thank University of Brighton (UoB) and University of Brasilia (UnB) for providing funds to perform the present research.
Author information
Authors and Affiliations
Contributions
K.C.R.A. planned and performed all experiments. E.X.F.F. donated soil isolates. M.M.F and R.A.F. performed experiments. D.B.P. and R.A.F. assisted in data evaluation. K.C.R.A., P.OM. and D.B.P. wrote the manuscript. AP revised the statistics results and wrote part of the document. J.I.S., D.B.P. and P.O.M. conceived the research project. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Andrade, K.C.R., Fernandes, R.A., Pinho, D.B. et al. Sequencing and characterization of an L-asparaginase gene from a new species of Penicillium section Citrina isolated from Cerrado. Sci Rep 11, 17861 (2021). https://doi.org/10.1038/s41598-021-97316-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-97316-1
This article is cited by
-
Optimizing recombinant production of L-asparaginase 1 from Saccharomyces cerevisiae using response surface methodology
Folia Microbiologica (2024)
-
Cacti as low-cost substrates to produce L-asparaginase by endophytic fungi
World Journal of Microbiology and Biotechnology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.