Abstract
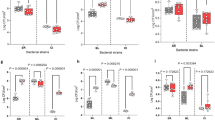
In a pioneering study, Zaura et al. (2009) found that majority of oral microbes fall within the five phyla including, Firmicutes, Proteobacteria, Actinobacteria, Bacteroidetes and Fusobacteria. Subsequent studies further identified a set of microbes that were commonly shared among unrelated individuals (i.e., core). However, these existing studies may have not been designed to investigate the interactions among various core species. Here by harnessing the power of ecological network analysis, we identified some important ecological guilds in the form of network clusters. In particular, we found that the strongest cluster is an alliance between Firmicutes and Bacteroidetes against Actinobacteria (FBA-guild). Within the guild, we further identified two sub-guilds, the Actinobacteria-dominant sub-guild (ASG) and Firmicutes-dominant allied with Bacteroidetes sub-guild (FBSG). Furthermore, we identified so-termed guard nodes in both sub-guilds, and their role may be to inhibit the peer sub-guild given they held competitive interactions only with the outside nodes only but held cooperative interactions only with the internal nodes, which we termed civilian nodes given that they only held cooperative interactions. We postulated that FBA-guild might be to do with protection of oral health against some opportunistic pathogens from Corynebacterium and Actinomyces, the two major genera of Actinobacteria (target of FB alliance).
Similar content being viewed by others
Introduction
The investigation of the human oral microbiome is among the earliest studies of the human microbiome. For example, as early as in the 1990s, scientists and clinicians have already resorted to ecological theories to interpret the etiology of periodontitis1,2. The oral cavity is a complex ecosystem comprised of many habitats, such as tongue, palates, cheeks, teeth and gingival sulcus. These inter-connected intra-oral habitats may have different microbial profiles, and the whole oral microbiome is therefore a meta-microbial community. The oral microbiome plays a critical role in maintaining our oral health, and its dysbiosis can lead to oral diseases such as dental caries, gingivitis and periodontitis3,4,5,6,7,8,9. In addition, recent studies have revealed that oral microbiome may also be associated with many systemic diseases including atherosclerosis10, gastrointestinal cancer11, inflammatory bowel disease12 and diabetes13.
The composition of oral microbiome is influenced by host genetics14, and fluctuates with host health status and lifestyle-related factors15, which could lead to great inter-subject heterogeneity. For example, diet is one of the major factors that disturb the balance of oral microbiota. Adler et al. (2013) showed that, compared with hunter-gatherer diet, carbohydrate-rich farming diet (or modern diet) lead to the modern oral microbiota with low biodiversity and the dominance of cariogenic bacteria16. Moreover, some studies have found that smoking altered the structure of oral microbiome, which increase the risk for periodontitis17,18. Health status and habitats may directly or indirectly influence the factors of host intra-oral environment, such as pH and iron, that has been reported to have significant influences on the oral microbiota19. However, recent studies (e.g., He et al. 2014, Belstrom et al. 2016) have also suggested that the oral microbiome is relatively stable and resistant to species invasions20,21. The maintenance factors of oral microbiota, as Zaura et al. (2014) reviewed, include host-derived and microbe-derived, in which the host immune system plays a key role on the homeostasis of oral microbial community22. In the meantime, oral micro-ecosystem may also help to improve and perfect the host immune system. The invasion-resistance and inter-species interactions as microbe-derived factors are also important to maintain the stability of oral microbiome22.
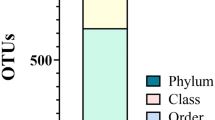
A primary mission of NIH-HMP (human microbiome project) was to answer the question whether there is a core set of species in the human microbiome, and the studies of human oral microbiome were set with a similar goal18,23,24,25,26. In a seminal study on the oral microbiome, Zaura et al. (2009) defined the oral core microbiome as the commonly shared unique sequences (phylotypes) among unrelated individuals24. Through multi-site studies of the healthy oral microbiome, Zaura et al. (2009) detected over 500 species in each individual oral microbial community, and the majority of taxa fall within the five phyla including Firmicutes, Proteobacteria, Actinobacteria, Bacteroidetes and Fusobacteria24. Other studies have also identified the presence of a common core microbiome in the human oral cavity, which is generally defined as the phylotypes or operational taxonomic units (OTUs), in a specific healthy habitat, that are shared among the vast majority of humans24,27,28,29. Nevertheless, the pursuing of core microbiome turned out to be more elusive than initially30,31,32. As Zaura et al. suggested, the studies of oral microbiome should be shifted to functional approaches such as metabolism, and more advanced topics such as the interactions between fungi and bacteria, host environment and microbiota, as well as the inter-species interactions should be paid to more33,34,35.
The objective of this study is to further investigate the inter-species interactions in the core of oral microbiome by detecting and analyzing the important clusters (or ecological guilds) in the oral microbiome network by reanalyzing the datasets originally published by Zaura et al. (2009) for investigating core of the oral microbiome24. An in-depth study of the core oral microbiota should be critical for understanding the structure and stability mechanism of oral microbiome, which can also be significant for investigating the etiology of oral diseases.
Materials and Methods
The oral microbiome datasets
The 16S ribosomal RNA datasets of the human oral microbiome analyzed in this report were first reported in Zaura et al.24. The oral samples were selected from several sites of three healthy male adults, including dental surfaces of upper incisor and upper molar, mucosa of cheek, hard palate and tongue surface, and saliva. A total of 29 oral microbiome samples from three healthy individuals were collected and sequenced with 16s-rRNA amplicon sequencing technology. Each individual were sampled at 9 or 10 oral sites, and on average, 6315 unique sequences were obtained for each sample. There were 818 OTUs (operational taxonomic units) identified at 97% similarity level. Each OTU was labeled after their lowest annotated taxonomic level and a unique number (such as Corynebacterium_767).
Although the number of individuals is relatively small, the sampled sites from each individual (9–10) as well as the reads per sample (6315 on average) are sufficiently large to enable our network analysis for investigating core oral microbiota. In particular, we take advantages of the findings and insights on the oral microbial core obtained from this same datasets by the original scientists24.
The network analysis approach
We adopted standard approach for correlation network analysis to construct and analyze the human oral microbiome network with the 16s-rRNA datasets originally reported by Zaura et al.24,36,37,38,39. To reduce the noise effect of the OTUs with extremely low abundance and potentially spurious OTU reads, we filtered out the OTUs whose total reads in all 29 samples were less than 30, i.e., approximately one read per sample, equivalent to removing the so-called singleton, which is a common practice in ecological analysis. A total of 347 OTUs remain after the filtering operation, and their abundances were utilized to construct the species correlation network, based on Spearman’s rank correlation coefficient (R). The correlation relationships with |R| ≥ 0.6 and p-value ≤ 0.05 (significance level) were set as criteria for selecting network edges (links).
Cytoscape software (Version 2.8.3) was used to visualize the network graphs and MCODE plug-in for Cytoscape for detecting network clusters (modules)36,40,41. MCODE (Molecular Complex Detection) is a graph-theoretic clustering algorithm, which was first introduced by Bader et al. (2003) to identify molecular complexes in large protein interaction networks41. The molecular complexes in a protein network can be considered as the locally dense regions or clusters in a graph. The core algorithm of the MCODE is to detect the clusters of vertex weighted based on the local neighborhood density or cliquishness, which can be measured by the clustering coefficient, Ci,
where ki is the number of neighborhood vertices of vertex i, and n is the number of edges (links) in the neighborhood. There are three main steps in the MCODE algorithms. First, MCODE weights all vertices based on their local neighborhood density, generating the so-termed vertex weighted graph (VWG). Next, the locally highest weighted vertex in the VWG will be set as a seed for a candidate cluster, and the cluster will be isolated by outwardly traversing from the seed to find all the vertices whose weights are within a given threshold. The third step is post-processing to filter vertices in the candidate cluster according to the given parameter sets. The detail interpretation of the algorithm is referred to Bader et al.41.
In addition, iGraph R-package was utilized for computing the network properties42. We also identified the P/N (positive to negative links) ratio in the network, which is a network property proposed by Ma (2017) to measure the balance between cooperative and competitive interactions in the microbiome43.
Results and Discussion
Basic network properties
The oral microbiome network we reconstructed contained 335 nodes (OTUs) and 4335 links (3692 positive links and 643 negative links). Table 1 lists the basic network properties. As shown in Table 1, the ratio of positive to negative correlation relationships is approximately 5.7, which suggests that the oral microbiome network is predominantly cooperative43. Since these network properties do not offer much intuitive insights on the oral microbiome network, we focus on the detection of network clusters, which are equivalent to the guild in ecological community, through which we expect to deepen our understanding on the critical species interactions in the oral microbiome and to further shed light on the structure and functions of core oral microbiota or guilds.
FBA Guild—Firmicutes-Bacteroidetes ally against Actinobacteria
An ecological guild can be defined as a group of species that exploit the same resources or exploit different resources in related manners44. Guild members could be competing for resources and hence hold negative correlation relationships in their abundances in the species correlation network of the oral microbiome. They may also cooperatively exploit other resources and therefore hold positive correlation relationships. Although rigorously defining and identifying microbial guilds can be rather challenging at this stage of human microbiome research mainly because functional studies on the microbiome are still scarce, we believe that the following exploration for bacterial guilds through network cluster detection technique is the best we can perform in order to deepen our understanding on the interspecific interactions and to further shed light on the structure and functions of core oral microbes.
The technique we use for detecting network clusters (modules or ecological guilds) is the MCODE plug-in for Cytoscape40,41. Table 2 lists the 11 clusters we detected with MCODE, including the cluster number, cluster score, number of nodes and number of edges for each cluster. The cluster score is a measure of the cluster density. The higher the cluster score is, and the stronger the corresponding cluster is. The strongest cluster (i.e., No. 1 cluster in Table 2) contains 55 nodes and 648 edges, which is nearly 1/6 of all OTUs in the oral microbiome network.
The strongest cluster primarily consists of the OTUs from three phyla: Actinobacteria, Firmicutes, and Bacteriodetes, and we term the strongest cluster as FBA-cluster with the initials of the three phyla (Fig. 1). As shown in Tables 3, 38.2% (21 out of 55) of the OTUs in the FBA-cluster belong to Actinobacteria, 25.5% (14 out of 55) to Firmicutes, and 16.4% (9 out of 55) to Bacteriodetes. As further illustrated below, FBA cluster or guild is a trio of Firmicutes-Bacteriodete (F-B) ally against Actinobacteria, in which Firmicutes and Bacteriodetes hold positive links and both hold negative links with Actinobacteria in oral microbiome network. In other words, the F-B coalition competes against Actinobacteria, and each of them holds negative relationship with their common ‘enemy’.
The strongest cluster (FBA cluster) in the healthy oral microbiome network. Symbols used: nodes in magenta—the OTUs of Actinobacteria phylum, nodes in yellow—the OTUs of Firmicutes phylum, nodes in cyan— the OTUs of Bacteroidetes phylum, nodes in gray—the OTUs of other phyla; edges in green— positive correlations, edges in red—negative correlations.
A broadly defined ecological guild almost always contains constituent guilds or sub-guilds, and so does our FBA guild that consists of two sub-guilds. Table 3 shows the component taxa of the two sub-guilds (sub-clusters) FBA contains. One is the Actinobacteria-dominant sub-guild (ASG), in which nearly 2/3 (67%) of species belong to Actinobacteria, and no Firmicutes exist and the number (only 2) of species from Bacteroidetes is negligible in the ASG sub-guild. Another is the Firmicutes-dominant sub-guild, in which more than 40% of the species are from the phylum of Firmicutes, and 21% are from Bacteroidetes in this sub-guild. Given the significant presence of Bacteroidetes in the second sub-guild, we term it FBSG (Firmicutes Bacteroidetes sub-guild).
Figure 1 illustrates the topological structure of the two sub-guilds, the left side is the ASG sub-guild and the right side is the FBSG sub-guild. We consider FBSG sub-guild as an ‘alliance’ between Firmicutes and Bacteroidetes against ASG sub-guild. Our justifications include: (i) all interactions between F & B are cooperative, as illustrated in the all positive relationships in the FBSG sub-guild (green edge in the left sub-cluster); (ii) all interactions between the FBSG and ASG sub-guilds are competitive, as illustrated in the all negative relationships between the two sub-clusters (the intermediate red edges); (iii) the numbers of members (network nodes) in both sub-guilds (i.e., 21 A in ASG vs. 14 F + 7B in FBSG) are also on a par with each other.
Figure 2A,B were drawn to facilitate the visualization of the relationships mentioned above by deconstructing graph (Fig. 1) of FBA cluster into two regional blocks. Figure 2A shows the ASG sub-guild, i.e., the left block in Fig. 1. Figure 2B shows the FBSG sub-guild, i.e., the right block in Fig. 1. Figure 3 shows the interactions between both the sub-guilds. While Fig. 2A,B are self-evident, more insights can be revealed by further analyzing the interactions between both the sub-guilds. In remaining part of this section, we focus on further exploring those interactions (in the forms of inter-species and inter-guilds) to complete the objective set for this article as introduced previously.
(A) The ASG sub-guild (Actinobacteria-dominant sub-guild). Symbols used: nodes colored in magenta—the OTUs of Actinobacteria phylum, nodes in cyan—the OTUs of Bacteroidetes phylum, nodes in gray—the OTUs of other phyla; edges in green—positive interactions; no negative links existed here. (B) The FBSG sub-guild (Firmicutes-dominant with Bacteroidetes ally sub-guild). Symbols used: nodes colored in magenta—the OTUs of Actinobacteria phylum, nodes in yellow—the OTUs of Firmicutes phylum, nodes in cyan—the OTUs of Bacteroidetes phylum, nodes in gray—the OTUs of other phyla, edges in green—positive correlations; no negative links existed here.
The negative relationships between the two sub-guilds of the FBA guild. Symbols used: nodes in magenta—the OTUs of Actinobacteria phylum, nodes in yellow—the OTUs of Firmicutes phylum, nodes in cyan—the OTUs of Bacteroidetes phylum, nodes in gray— the OTUs of other phyla; edges in green—positive correlations, edges in red—negative correlations.
FBA Guild—further analysis of the inter-species and inter-guild interactions
In the previous sub-section, we observed the two sub-guilds of the FBA guild, i.e., sub-guild ASG dominated by Actinobacteria, and sub-guild FBSG dominated by Firmicutes and its ally Bacteroidetes. Both sub-guilds compete with each other. The interactions (correlations) within each sub-guild are cooperative (positive), but the interactions between the two sub-guilds are competitive (negative).
To further explore the inter-species and inter-sub-guild interactions, we introduce the concept of ‘guard’ nodes. We define guard nodes of a sub-guild as nodes that have negative relationships with the nodes in another sub-guild, but with full positive relations with nodes within its own sub-guild. That is, guard nodes “guard against” their counterparts in another sub-guild, but are ‘friendly’ to their own sub-guild-members (i.e., their relationships with other sub-guild members are cooperative). Figure 3 shows the interactions between guard nodes from both ASG and FBSG sub-guilds. In Fig. 3, the left column exhibits the 14 guard nodes in the ASG, in which 7 species are from Cornebacterium genus, 2 species from Actinomyces genus, and the remaining 5 guard species from other small phyla but none from Firmicutes or Bacteriodetes. The right column of Fig. 3 displays the 16 guard nodes in the FBSG, in which 7 species are from Firmicutes, 4 from Bacteriodetes, 2 from Actinobacteria, and 3 from other phyla.
The taxonomic information FBA-guild is listed in Table 4, which is tabulated based on Figs. 1 and 3. In Table 4, nodes are classified into two types: one type is the guard node that is always ‘hostile’ (negative interactions) to its counterparts in another sub-guild but always ‘friendly’ to its civilian nodes within the same sub-guild; the other type is, what we called, civilian node who may be ‘friendly’ to any node in the whole FBA guild (or any sub-guild). The distinction between civilian nodes and guard nodes suggests the possible functional differentiations among the nodes in each sub-guild. It should be the differentiation that shape or even determine the interactions between two sub-guilds. Two types of nodes may play rather different roles in the interactions. The role (function) of guard nodes should be to protect their home-sub-guild against invasions from foreign guards, while they should never compete with any nodes in their homeland. Using an analogy, civilian nodes in both sub-guilds, although they have their own citizenships, can ‘friendly’ interact with any nodes regardless of their citizenship. Using another analogy, FBA guild is like a global village, where civilians may friendly trade with each other, but each sub-guild still preserves their military forces (guards) and ‘fight’ each other to keep order. This reminds us that, in the FBA triangle relationship, although FB (Firmicutes & Bacteriodetes) is united against A (Actinobacteria), the competition between FB & A only occurs in military sector and two sides (sub-guilds) cooperate with each other in civilian sectors.
Table 5 further lists all negative interactions (correlations) between both the sub-guilds of the FBA guild. Table 6 further lists the number of positive, negative and total interactions, respectively, between Firmicutes, Bacteriodetes and Actinobacteria. Table 6 also computed the P/N (positive to negative) ratio of links between the three phyla according to Ma (2017) P/N ratio approach43. Table 6 indicates that in the F-B alliance against A, F plays a larger role than B does, given that P/N ratio between F & A is approximately ½ that between B & A and small P/N ratio is resulted from larger number of competitive interactions (the denominator).
FBA Guild—bring back the ‘ugly’ other phyla
In previous sub-sections, we intentionally ignore the “others phyla” that do not belong to Firmicutes, Bacteroidetes and Actinobacteria to simplify the interpretation and presentation of our findings. Here we bring back “the others” in Fig. 4. In Fig. 4, rather than laying out the FBA cluster as a ‘bipartite’ network as in Figs. 2 and 3, the network was laid out as four blocks. Besides the three blocks of Firmicutes, Bacteroidetes and Actinobacteria, respectively, the “other phyla” occupied a fourth block (the left-down group, nodes in grey). First, the different layouts from Fig. 1 to Fig. 4 were made to facilitate the visual inspection of various facets of the network graph, and they, of course, influence neither the true cluster structure nor its interpretation. Second, most of the other group members belong to Proteobacteria and Fusobacteria, the two other core phyla Zaura et al. (2009) had already identified24. While the distributions of Firmicutes, Bacteroidetes and Actinobacteria are rather aggregated in the sense that they form strongly connected sub-clusters, the distributions of Proteobacteria and Fusobacteria are rather dispersed in the sense that they are distributed all over the place (all sub-clusters), and they do not dominate in any sub-clusters. Furthermore, “the others” do not seem to have a special or fixed pattern in their interactions with Firmicutes, Bacteroidetes and Actinobacteria. For example, while most interactions are cooperative, competitive relationships also exist. Using an analogy, we characterize “the others” as “nomads” of small “ethnic groups” in the FBA guild. Although further investigation on “the others” could be interesting, we believe the results should not affect the validity of the findings discussed in previous sections.
The relationships between Actinobacteria, Firmicutes, Bacteroidetes and the “other phyla” in the FBA guild. Symbols used: nodes in magenta—the OTUs of Actinobacteria phylum, nodes in yellow—the OTUs of Firmicutes phylum, nodes in cyan—the OTUs of Bacteroidetes phylum, nodes in gray— the OTUs of “other phyla”; edges in green—positive correlations, edges in red—negative correlations.
As a side note, we conducted similar examinations of other clusters detected with MCODE and listed in Table 2, but failed to find similarly interesting structures or interactions. Since majority of the core oral microbes (i.e., Firmicutes, Proteobacteria, Actinobacteria, Bacteroidetes and Fusobacteria) identified by Zaura et al. (2009) are contained in the FBA guild, which is the largest (also the strongest) cluster we detected, the failure should not be surprising24. Hence, the structure and species-interaction mechanism of FBA guild revealed in this article also represent those of core oral microbes.
The mission of FBA guild in the oral microbiome—a new hypothesis
In previous sections, we have showed the structure and inter-species interactions within the FBA. Nevertheless, at this stage, we cannot fully explain the underlying mechanisms leading to this interesting triangular relationship, which requires experimental investigations beyond the scope of this article. Here, we propose a new hypothesis to explain the observed phenomenon, and hope to stimulate the further studies on this obviously rather important phenomenon.
First, both Firmicutes and Bacteroidetes have been the dominant players in the gut microbiome and have attracted extensive attentions in recent years, in particularly, their implications to obesity. The ratio of Firmicutes to Bacteroidetes (F/B) has been suggested as an index of the health of gut microbiome. Both the phyla are the most abundant taxa of gut microbiome, although the inter-individual variations are huge and their dynamics is rather dramatic30,45,46,47. For example, the F/B ratio could decrease from approximately 10.9 in middle-age adults to 0.6 in the elderly45,48. In the oral microbiome, Zaura et al. (2009, 2014, 2015) studies also suggested the dominance of both phyla, and contributed approximately 50% (36% Firmicutes and 12% Bacteroidetes) to the oral microbiome, and were two of the five major phyla in the oral microbes [the other three were: Proteobacteria (22%), Actinobacteria (24%), and Fusobacteria (4%)]23,24,33. Since oral and gut environments are well connected, and bacteria may freely disperse but environment would select who can stay and who can only be by-passers or nomads49,50. Therefore, it can be expected that the oral and gut microbiomes should be of certain level of similarity. Therefore, the dominance of F & B in the oral environment can be expected, but we are puzzled by the fact that there were not any negative interactions between F & B (Table 5, Fig. 4). It might be just that the gut environment allows for the competition between the both because both F & B may be competing for the fermentation niche, one of the three metabolic niches (the other twos are sulfate reduction and methanogenesis) gut microbes compete for in the gut ecosystem51. Existing literature reveals that majority of species in F & B are involved in fermentation51. However, healthy oral environment is not a fermentation habitat in general, and therefore F & B lose the battle ground for competing, instead they may turn to cooperation (positive correlations), possibly forming an alliance against Actinobacteria as we discovered previously. But this leads to another question, which we try to answer below, why do F & B both do not like Actinobacteria?
Second, note that in the Actinobacteria-dominant sub-cluster, Cornebacterium and Actinomyces are the two primary genera. Existing literatures suggest that these two genera include some of the notorious pathogens, especially opportunistic pathogens. For example, C. diphtheriae causes diphtheria. Other pathogenic species in humans include: C. amicolatum, C. striatum, C. jeikeium, C. urealyticum, and C. xerosis5,52,53,54,55. Certain species of Actinomyces are known to be opportunistic pathogens, particularly, when the immune system of host is weak56,57,58,59,60,61. Of course, there are innocuous species in these genera, given that oral microbiome and its environment (host) usually live harmoniously and their interactions are cooperative in large62,63,64,65,66. We conjecture that the potentially suppression of F-B alliance against A exhibited by the FBA guild, as the primary component of core oral microbiome, should be important for maintaining a healthy oral microbiome and protect humans from many opportunistic infections.
A major limitation of this study is that the dataset used to reconstruct the oral microbiome network was published a decade ago, and the 29 samples were collected from three healthy individuals only (Zaura et al. 2009). Therefore, the findings from our reanalysis of the datasets should be validated with more extensive datasets in future. A primary motivation for us to publish our results was to demonstrate the potentially important application of the concept of ecological guild in microbiome studies. The dataset originally reported by Zaura et al. (2009), which we reanalyzed, offered us an excellent opportunity to pursue our objective because of its multi-site nature and high-quality sequencing experiments.
Data availability
The raw sequencing datasets were originally collected and published by Zaura et al. (2009). Detailed access information was available in “Zaura E, Keijser BJF, Huse SM, et al. (2009). Defining the healthy “core microbiome” of oral microbial communities. BMC Microbiology, 9: 259”.
References
Grenier, D. & Mayrand, D. Adult periodontitis: an ecological perspective of mixed infections. Trends in Microbiology 3(4), 148 (1995).
Kroes, I., Lepp, P. W. & Relman, D. A. Bacterial diversity within the human subgingival crevice. Proc. Natl. Acad. Sci. USA 96(25), 14547–52 (1999).
Marsh, P. D. Microbiology of dental plaque biofilms and their role in oral health and caries. Dental. clin. North Am. 54, 441–454 (2010).
Belda-Ferre, P., Alcaraz, L. D., Raúl Cabrera-Rubio, H. R. & Mira, A. The oral metagenome in health and disease. The ISME Journal 6, 46–56 (2012).
Zhou, M. et al. DongInvestigation of the effect of type 2 diabetes mellitus on subgingival plaque microbiota by high-throughput 16S rDNA pyrosequencing. Plos One 8, 1–8 (2013).
Shaw, L. et al. Distinguishing the Signals of Gingivitis and Periodontitis in Supragingival Plaque: a Cross-Sectional Cohort Study in Malawi. Appl. Environ. Microbiol. 82(19), 6057–67 (2016).
Kirst, M. E. et al. Dysbiosis and Alterations in Predicted Functions of the Subgingival Microbiome in Chronic Periodontitis. Applied and Environmental Microbiology 81(2), 783–93 (2015).
Szafranski, S. P. et al. High-Resolution Taxonomic Profiling of the Subgingival Microbiome for Biomarker Discovery and Periodontitis Diagnosis. Appl. Environ. Microbiol., 81(3), 1047–58 (2015).
Meuric, V. et al. Signature of Microbial Dysbiosis in Periodontitis. 83(14), e00462–17 (2017).
Koren, O. et al. Human oral, gut, and plaque microbiota in patients with atherosclerosis. Proc. Natl. Acad. Sci. USA 108(Suppl), 4592–4598 (2011).
Ahn, J., Chen, C. Y. & Hayes, R. B. Oral microbiome and oral and gastrointestinal cancer risk. Cancer Causes Control 23, 399–404 (2012).
Atarashi, K. et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science 358, 359–365 (2017).
Long, J. et al. Association of oral microbiome with type 2 diabetes risk. J. Periodont Res. 52, 636–643 (2017).
Gomez, A., Espinoza, J. L. & Harkins, D. M. Host Genetic Control of the Oral Microbiome in Health and Disease. Cell Host & Microbe. 22, 269–278 (2017).
Belstrom, D. et al. Bacterial profiles of saliva in relation to diet, lifestyle factors, and socioeconomic status. J Oral Microbiol 4(6), 1–9 (2014).
Adler, C. J. et al. Sequencing ancient calcified dental plaque shows changes in oral microbiota with dietary shifts of the Neolithic and Industrial revolutions. Nature Genet 45(4), 450–5 (2013).
Wu, J. et al. Cigarette smoking and the oral microbiome in a large study of American adults. The ISME Journal 10(10), 2435–46 (2016).
Ganesan, S. M. et al. A tale of two risks: smoking, diabetes and the subgingival microbiome. The ISME Journal 11, 2075–2089 (2017).
Zhou, J. et al. Influences of pH and Iron Concentration on the Salivary Microbiome in Individual Humans with and without Caries. Appl. Environ. Microbiol. 83(4), e02412 (2018).
He, X., Mclean, J. S., Guo, L., Lux, R. & Shi, W. The social structure of microbial community involved in colonization resistance. ISME J. 8, 564–574 (2014).
Belstrøm, D. et al. Temporal stability of the salivary microbiota in oral health. PLoS One 11(1), e0147472 (2016).
Zaura, E., Nicu, E. A., Krom, B. P. & Keijser, B. J. F. Acquiring and maintaining a normal oral microbiome current perspective. Frontiers in Cellular and Infection Microbiology 4, 85–92 (2014).
Human Microbiome Project C. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Zaura, E., Keijser, B. J., Huse, S. M. & Crielaard, W. Defining the healthy “core microbiome” of oral microbial communities. BMC Microbiology 9, 259 (2009).
Griffen, A. L. et al. CORE: A Phylogenetically-Curated 16S rDNA Database of the Core Oral Microbiome. PloS One 6(4), e19051 (2011).
Wang, Y. et al. Profiling of Oral Microbiota in Early Childhood Caries Using Single-Molecule Real-Time Sequencing. Frontiers in Microbiology 8, 2244 (2017).
Ling, Z. et al. Pyosequencing analysis of the human microbiota of healthy Chinese undergraduates. BMC Genomics 14, 390 (2013).
Li, K., Bihan, M. & Methe, B. A. Analyses of the stability and core taxonomic memberships of the human microbiome. PLoS One 8, e63139 (2013).
Jiang, W. X. et al. The impact of various time intervals on the supragingival plaque dynamic core microbiome. PloS One 10(5), e0124631 (2015).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473(7346), 174–80 (2011).
Li, W. & Ma, Z. S. Diversity scaling of human vaginal microbial communities. Zoological Research 40(6), 552–559 (2019).
Ma, Z. S. & Li, L. Quantifying the human vaginal community state types (CSTs) with the species specificity index. PeerJ. 5, e3366 (2017).
Zaura, E. & Mira, A. Editorial: The oral microbiome in an ecological perspective. Frontiers in Cellular and Infection Microbiology 5, 39 (2015).
Takahashi, N. Oral Microbiome Metabolism: From “Who Are They?” to “What Are They Doing”. J. Dent. Res. 94(12), 1628–37 (2015).
Marsh, P. D. & Zaura, E. Dental biofilm: ecological interactions in health and disease. J. Clin. Periodontol. 44(Supple 18), S12–S22 (2017).
Junker, B. H. & Schreiber, F. Analysis of Biological Networks. Wiley, NJ, USA (2008).
Ma, Z. S. et al. Network analysis suggests a potentially ‘evil’ alliance of opportunistic pathogens inhibited by a cooperative network in human milk bacterial communities. Scientific Reports 5, 8275 (2015).
Ma, Z. S., Li, L., Li, W., Li, J. & Chen, H. Integrated network-diversity analyses suggest suppressive effect of Hodgkin’s lymphoma and slightly relieving effect of chemotherapy on human milk microbiome. Scientific Reports 6, 28048 (2016).
Ma, Z. S. & Ye, D. D. Trios—promising in silico biomarkers for differentiating the effect of disease on the human microbiome network. Scientific Reports 7(1), 13259 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 4(1), 2 (2003).
Csardi, G. & Nepusz, T. The iGraph software package for complex network research. I. J. Complex Systems, No. 5 (2005).
Ma, Z. S. The P/N (Positive-to-Negative Links) Ratio in Complex Networks—A Promising In Silico Biomarker for Detecting Changes Occurring in the Human Microbiome. Microbial. Ecology 75(4), 1063–1073 (2017).
Simberloff, D. & Dayan, T. The Guild Concept and the Structure of Ecological Communities. Annual Review of Ecology and Systematics 22, 115–43 (1991).
Marx, T. Immunoprotective Effects of Probiotics in the Elderly, p 363–372. In Wastson, R. R. (ed), Foods and Dietary Supplements in the Prevention and Treatment of Disease in Older Adults, Academic Press (2015).
Rajilic-Stojanovic, M. & de Vos, W. S. The first 1000 cultured species of the human gastrointestinal microbota. FEMS Microbiol. Rev. 38, 996–1047 (2014).
Sankar, S. A., Lagier, J., Pontarotti, P., Raoult, D. & Fournier, P. The human gut microbiome, a taxonomic conundrum. Systematic and Applied Microbiology 38, 276–286 (2015).
Mariat, D. et al. The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiology 9, 123 (2009).
van der Gast, C. J. Microbial biogeography and what Baas Becking should have said. Microbiology Today 40, 108–111 (2013).
van der Gast, C. J. Microbial biogeography: the end of the ubiquitous dispersal hypothesis? Environmental Microbiology 17(3), 544–6 (2015).
Gillilland, M. G., Young, V. B. & Huffnagle, G. B. Gastrointestinal Microbial Ecology with Perspectives on Health and Disease, p 1119–1134. In Said, H. M. (ed), Physiology of the Gastrointestinal Tract, 5th ed., Academic Press (2012).
Takazoe, I. & Itoyama, T. Analytical electron microscopy of Bacterionema matruchotii calcification. J. Dent. Res. 59, 1090–1094 (1980).
Tsuzukibashi, O. et al. A selective medium for the isolation of Corynebacterium species in oral cavities. J. Microbiol. Methods 104, 67–71 (2014).
Moon, J. H., Lee, J. H. & Lee, J. Y. Subgingival microbiome in smokers and non-smokers in Korean chronic periodontitis patients. Mol. Oral. Microbiol 30(3), 227–41 (2014).
Ribeiro, A. A. et al. The oral bacterial microbiome of occlusal surfaces in children and its association with diet and caries. PLoS One 12(7), e0180621 (2017).
Feder, H. M. Actinomycosis manifesting as an acute painless lump of the jaw. Pediatrics 85, 858–864 (1990).
Smego, R. A. & Foglia, G. Actinomycosis. Clin. Infect. Dis. 26, 1255–1263 (1998).
Dige, I., Raarup, M. K., Nyengaard, J. R., Kilian, M. & Nyvad, B. Actinomyces naeslundii in initial dental biofilm formation. Microbiology 155(Pt 7), 2116–2126 (2009).
Shimada, E., Kataoka, H., Miyazawa, Y., Yamamoto, M. & Igarashi, T. Lipoproteins of Actinomyces viscosus induce inflammatory responses through TLR2 in human gingival epithelial cells and macrophages. Microbes. Infect 14(11), 916–921 (2012).
Sato, T. et al. Peptidoglycan of Actinomyces naeslundii induces inflammatory cytokine production and stimulates osteoclastogenesis in alveolar bone resorption. Arch. Oral. Biol. 57(11), 1522–1528 (2012).
Valour, F. et al. Actinomycosis: etiology, clinical features, diagnosis, treatment, and management. Infect. Drug. Resist. 7, 183–197 (2014).
Lagrou, K., Verhaegen, J., Janssens, M., Wauters, G. & Verbist, L. Prospective Study of Catalase-positive Coryneform Organisms in Clinical Specimens: Identification, Clinical Relevance, and Antibiotic Susceptibility. Diagnostic Microbiology and Infectious Disease 30(1), 7–15 (1998).
Boc, S. F. & Martone, J. D. Osteomyelitis caused by Corynebacterium jeikeium. Journal of the American Podiatric Medical Association 85(6), 338–9 (1995).
Kono, M., Sasatsu, M. & Aoki, T. R Plasmids in Corynebacterium xerosis Strains. Antimicrobial Agents and Chemotherapy 23(3), 506–8 (1983).
Pitcher, D. G. Deoxyribonucleic acid base composition of Corynebacterium diphtheriaeand other corynebacteria with cell wall type IV. FEMS Microbiology Letters 16(2–3), 291–5 (1983).
Mark Welch, J. L., Rossetti, B. J., Rieken, C. W., Dewhirst, F. E. & Borisy, G. G. Biogeography of a human oral microbiome at the micron scale. Proc. Natl. Acad. Sci. USA 113(6), E791–800 (2016).
Author information
Authors and Affiliations
Contributions
Z.M. designed the study and wrote the paper. W.L. performed the data analysis and interpretations. All authors approved the submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, W., Ma, Z.(. FBA Ecological Guild: Trio of Firmicutes-Bacteroidetes Alliance against Actinobacteria in Human Oral Microbiome. Sci Rep 10, 287 (2020). https://doi.org/10.1038/s41598-019-56561-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56561-1
This article is cited by
-
The association of tonsillar microbiota with biochemical indices based on obesity and tonsillar hypertrophy in children
Scientific Reports (2023)
-
The Upper Respiratory Tract Microbiome Network Impacted by SARS-CoV-2
Microbial Ecology (2023)
-
Brief overview of dietary intake, some types of gut microbiota, metabolic markers and research opportunities in sample of Egyptian women
Scientific Reports (2022)
-
Differences in the gut Firmicutes to Bacteroidetes ratio across age groups in healthy Ukrainian population
BMC Microbiology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.