Abstract
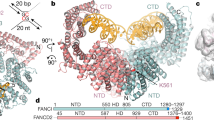
Vertebrate DNA crosslink repair excises toxic replication-blocking DNA crosslinks. Numerous factors involved in crosslink repair have been identified, and mutations in their corresponding genes cause Fanconi anemia (FA). A key step in crosslink repair is monoubiquitination of the FANCD2–FANCI heterodimer, which then recruits nucleases to remove the DNA lesion. Here, we use cryo-EM to determine the structures of recombinant chicken FANCD2 and FANCI complexes. FANCD2–FANCI adopts a closed conformation when the FANCD2 subunit is monoubiquitinated, creating a channel that encloses double-stranded DNA (dsDNA). Ubiquitin is positioned at the interface of FANCD2 and FANCI, where it acts as a covalent molecular pin to trap the complex on DNA. In contrast, isolated FANCD2 is a homodimer that is unable to bind DNA, suggestive of an autoinhibitory mechanism that prevents premature activation. Together, our work suggests that FANCD2–FANCI is a clamp that is locked onto DNA by ubiquitin, with distinct interfaces that may recruit other DNA repair factors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Cryo-EM maps generated during this study have been deposited in the Electron Microscopy Data Bank (EMDB) with accession codes EMD-10532 (D2–I), EMD-10531 (ubD2–I), and EMD-10534 (D2–D2). Models generated during this study have been deposited in the protein databank (PDB) with accession codes PDB 6TNG (D2–I), PDB 6TNF (ubD2–I) and PDB 6TNI (D2–D2). MS data have been deposited in the PRIDE database with accession code PXD017020. Original gels and blots in Figs. 1b, 4 and 5b and Extended Data Figs. 1b, 3b,c, 4a,b and 5 are provided in Supplementary Fig. 1. Data for quantifications in Fig. 4 and Extended Data Fig. 4c are available as Source data with the paper online.
References
Kottemann, M. C. & Smogorzewska, A. Fanconi anaemia and the repair of Watson and Crick DNA crosslinks. Nature 493, 356–363 (2013).
Crossan, G. P. & Patel, K. J. The Fanconi anaemia pathway orchestrates incisions at sites of crosslinked DNA. J. Pathol. 226, 326–337 (2012).
Walden, H. & Deans, A. J. The Fanconi anemia DNA repair pathway: structural and functional insights into a complex disorder. Annu. Rev. Biophys. 43, 257–278 (2014).
Knipscheer, P., Raschle, M., Scharer, O. D. & Walter, J. C. Replication-coupled, DNA interstrand cross-link repair in Xenopus egg extracts. Methods Mol. Biol. 920, 221–243 (2012).
Alpi, A. et al. UBE2T, the Fanconi anemia core complex, and FANCD2 are recruited independently to chromatin: a basis for the regulation of FANCD2 monoubiquitination. Mol. Cell Biol. 27, 8421–8430 (2007).
Garcia-Higuera, I. et al. Interaction of the Fanconi anemia proteins and BRCA1 in a common pathway. Mol. Cell 7, 249–262 (2001).
Meetei, A. R. et al. A multiprotein nuclear complex connects Fanconi anemia and Bloom syndrome. Mol. Cell. Biol. 23, 3417–3426 (2003).
Rajendra, E. et al. The genetic and biochemical basis of FANCD2 monoubiquitination. Mol. Cell 54, 858–869 (2014).
van Twest, S. et al. Mechanism of ubiquitination and deubiquitination in the Fanconi anemia pathway. Mol. Cell 65, 247–259 (2017).
Knipscheer, P. et al. The Fanconi anemia pathway promotes replication-dependent DNA interstrand cross-link repair. Science 326, 1698–1701 (2009).
Sims, A. E. et al. FANCI is a second monoubiquitinated member of the Fanconi anemia pathway. Nat. Struct. Mol. Biol. 14, 564–567 (2007).
Smogorzewska, A. et al. Identification of the FANCI protein, a monoubiquitinated FANCD2 paralog required for DNA repair. Cell 129, 289–301 (2007).
Montes de Oca, R. et al. Regulated interaction of the Fanconi anemia protein, FANCD2, with chromatin. Blood 105, 1003–1009 (2005).
MacKay, C. et al. Identification of KIAA1018/FAN1, a DNA repair nuclease recruited to DNA damage by monoubiquitinated FANCD2. Cell 142, 65–76 (2010).
Liu, T., Ghosal, G., Yuan, J., Chen, J. & Huang, J. FAN1 acts with FANCI-FANCD2 to promote DNA interstrand cross-link repair. Science 329, 693–696 (2010).
Smogorzewska, A. et al. A genetic screen identifies FAN1, a Fanconi anemia-associated nuclease necessary for DNA interstrand crosslink repair. Mol. Cell 39, 36–47 (2010).
Kratz, K. et al. Deficiency of FANCD2-associated nuclease KIAA1018/FAN1 sensitizes cells to interstrand crosslinking agents. Cell 142, 77–88 (2010).
Klein Douwel, D. et al. XPF-ERCC1 acts in unhooking DNA interstrand crosslinks in cooperation with FANCD2 and FANCP/SLX4. Mol. Cell 54, 460–471 (2014).
Hodskinson, M. R. et al. Mouse SLX4 is a tumor suppressor that stimulates the activity of the nuclease XPF-ERCC1 in DNA crosslink repair. Mol. Cell 54, 472–484 (2014).
Yamamoto, K. N. et al. Involvement of SLX4 in interstrand cross-link repair is regulated by the Fanconi anemia pathway. Proc. Natl Acad. Sci. USA 108, 6492–6496 (2011).
Oestergaard, V. H. et al. Deubiquitination of FANCD2 is required for DNA crosslink repair. Mol. Cell 28, 798–809 (2007).
Kim, J. M. et al. Inactivation of murine Usp1 results in genomic instability and a Fanconi anemia phenotype. Dev. Cell 16, 314–320 (2009).
Nijman, S. M. et al. The deubiquitinating enzyme USP1 regulates the Fanconi anemia pathway. Mol. Cell 17, 331–339 (2005).
Tan, W. & Deans, A. J. A defined role for multiple Fanconi anemia gene products in DNA-damage-associated ubiquitination. Exp. Hematol. 50, 27–32 (2017).
Ishiai, M. et al. FANCI phosphorylation functions as a molecular switch to turn on the Fanconi anemia pathway. Nat. Struct. Mol. Biol. 15, 1138–1146 (2008).
Cheung, R. S. et al. Ubiquitination-linked phosphorylation of the FANCI S/TQ cluster contributes to activation of the Fanconi anemia I/D2 complex. Cell Reports 19, 2432–2440 (2017).
Lopez-Martinez, D. et al. Phosphorylation of FANCD2 inhibits the FANCD2/FANCI complex and suppresses the Fanconi anemia pathway in the absence of DNA damage. Cell Reports 27, 2990–3005.e5 (2019).
Sato, K., Toda, K., Ishiai, M., Takata, M. & Kurumizaka, H. DNA robustly stimulates FANCD2 monoubiquitylation in the complex with FANCI. Nucleic Acids Res. 40, 4553–4561 (2012).
Shakeel, S. et al. Structure of the Fanconi anaemia monoubiquitin ligase complex. Nature 575, 234–237 (2019).
Liang, C. C. & Cohn, M. A. UHRF1 is a sensor for DNA interstrand crosslinks. Oncotarget 7, 3–4 (2016).
Joo, W. et al. Structure of the FANCI-FANCD2 complex: insights into the Fanconi anemia DNA repair pathway. Science 333, 312–316 (2011).
Swuec, P. et al. The FA core complex contains a homo-dimeric catalytic module for the symmetric mono-ubiquitination of FANCI-FANCD2. Cell Reports 18, 611–623 (2017).
Liang, C. C. et al. The FANCD2–FANCI complex is recruited to DNA interstrand crosslinks before monoubiquitination of FANCD2. Nat. Commun. 7, 12124 (2016).
Chaugule, V. K., Arkinson, C., Toth, R. & Walden, H. Enzymatic preparation of monoubiquitinated FANCD2 and FANCI proteins. Methods Enzymol. 618, 73–104 (2019).
Zhi, G. et al. Purification of FANCD2 sub-complexes. Br. J. Haematol. 150, 88–92 (2010).
Thompson, E. L. et al. FANCI and FANCD2 have common as well as independent functions during the cellular replication stress response. Nucleic Acids Res. 45, 11837–11857 (2017).
Dubois, E. L. et al. A Fanci knockout mouse model reveals common and distinct functions for FANCI and FANCD2. Nucleic Acids Res. 47, 7532–7547 (2019).
Longerich, S. et al. Regulation of FANCD2 and FANCI monoubiquitination by their interaction and by DNA. Nucleic Acids Res. 42, 5657–5670 (2014).
Raschle, M. et al. Mechanism of replication-coupled DNA interstrand crosslink repair. Cell 134, 969–980 (2008).
Crossan, G. P. et al. Disruption of mouse Slx4, a regulator of structure-specific nucleases, phenocopies Fanconi anemia. Nat. Genet. 43, 147–152 (2011).
Parmar, K. et al. Hematopoietic stem cell defects in mice with deficiency of Fancd2 or Usp1. Stem Cells 28, 1186–1195 (2010).
Weissmann, F. et al. biGBac enables rapid gene assembly for the expression of large multisubunit protein complexes. Proc. Natl Acad. Sci. USA 113, E2564–E2569 (2016).
Hill, C. H. et al. Activation of the Endonuclease that Defines mRNA 3′ Ends Requires Incorporation into an 8-Subunit Core Cleavage and Polyadenylation Factor Complex. Mol. Cell 73, 1217–1231.e11 (2019).
Rueden, C. T. et al. ImageJ2: ImageJ for the next generation of scientific image data. BMC Bioinf. 18, 529 (2017).
Russo, C. J. & Passmore, L. A. Electron microscopy: ultrastable gold substrates for electron cryomicroscopy. Science 346, 1377–1380 (2014).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Nakane, T., Kimanius, D., Lindahl, E. & Scheres, S. H. Characterisation of molecular motions in cryo-EM single-particle data by multi-body refinement in RELION. Elife 7, e36861 (2018).
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Vijay-Kumar, S., Bugg, C. E. & Cook, W. J. Structure of ubiquitin refined at 1.8 Å resolution. J. Mol. Biol. 194, 531–544 (1987).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal. Chem. 68, 850–858 (1996).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and Go Extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Kolbowski, L., Mendes, M. L. & Rappsilber, J. Optimizing the parameters governing the fragmentation of cross-linked peptides in a Tribrid mass spectrometer. Anal. Chem. 89, 5311–5318 (2017).
Mendes, M. L. et al. An integrated workflow for crosslinking mass spectrometry. Mol. Syst. Biol. 15, e8994 (2019).
Lenz, S., Giese, S. H., Fischer, L. & Rappsilber, J. In-search assignment of monoisotopic peaks improves the identification of cross-linked peptides. J. Proteome Res. 17, 3923–3931 (2018).
Perez-Riverol, Y. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D Biol Crystallogr. 60, 2256–2268 (2004).
Naydenova, K. & Russo, C. J. Measuring the effects of particle orientation to improve the efficiency of electron cryomicroscopy. Nat. Commun. 8, 629 (2017).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Acknowledgements
We are grateful to T. Nakane, S. H. W. Scheres, F. O’Reilly, M. Babu, P. Emsley, J. Pruneda, T. Sijacki, J. A. W. Stowell and members of the Passmore lab for assistance and advice; the LMB EM facility, J. Grimmett and T. Darling (LMB scientific computation), and J. G. Shi (baculovirus) for support. This work was supported by the Medical Research Council, as part of United Kingdom Research and Innovation, MRC file reference numbers MC_U105192715 (L.A.P.) and MC_U105178811 (K.J.P.), and the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) grant no. 329673113 (J.R.). The Wellcome Centre for Cell Biology is supported by core funding from the Wellcome Trust (grant no. 203149; J.R.). P.A. is supported by an EMBO Long-Term Fellowship (ALTF 692–2018). We acknowledge Diamond Light Source for access to eBIC (proposals EM17434 and BI23268) funded by the Wellcome Trust, MRC and Biotechnology and Biological Sciences Research Council.
Author information
Authors and Affiliations
Contributions
P.A. and S.S. designed protein expression and purification schemes, performed ubiquitination and binding assays and performed cryo-EM, 3D reconstruction and modeling; Z.A.C. and J.R. performed crosslinking MS analysis. L.A.P. and K.J.P. supervised the research; all authors contributed to writing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Anke Sparmann and Inês Chen were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification of FANCI, FANCD2 and D2–I, and cryo-EM of D2–I, ubD2–I and D2 dimer.
a, Coomassie gel showing purified His-tagged FANCI, StrepII-tagged FANCD2 and D2–I after gel filtration. b–e, Cryo-EM of D2–I, ubD2–I and D2. b, Representative micrographs. Selected individual particles are marked with green circles. Scale bars, 25 nm. c, Fourier shell correlation (FSC) curves for gold-standard refinements. d, Angular distribution density plots of particles used in 3D reconstructions calculated using cryoEF63. Every point is a particle orientation, and the color scale represents the normalized density of views around this point. The color scale runs from 0 (low, blue) to 0.00026 (high, red). All complexes had a preferred orientation. Note that C2 symmetry was applied for FANCD2. e, Local resolution estimates calculated using ResMap64. Uncropped image for a is available in Supplementary Fig. 1.
Extended Data Fig. 2 Model fitting.
a, Overall fit of model to map for ubD2–I, D2–I and D2 (blue, FANCD2; magenta, FANCI; green, ubiquitin; yellow, DNA). b, Representative fits of model to map for FANCD2, FANCI, DNA and ubiquitin in the ubD2–I structure. c, FSC curves for model versus map. d, FANCD2 and FANCI structures from G. gallus (gg) were aligned with each other and with the M. musculus (mm)31 crystal structures using PDBeFOLD62 (http://www.ebi.ac.uk/msd-srv/ssm/); figures were prepared with PyMOL (The PyMOL Molecular Graphics System, Version 2.0, Schrödinger, LLC; https://pymol.org/2/).
Extended Data Fig. 3 Crosslinking mass spectrometry and analysis of DNA binding by FANCD2 and FANCI.
a, Distribution histogram of Cα−Cα distances between linked residue pairs in the 3D model of ubD2–I (left). Crosslinks with Cα−Cα distances below the theoretical crosslinking limit (30 Å) are shown in green. Overlength crosslinks (>30 Å) are shown in red. The distribution of Cα−Cα distances between random crosslinkable residue pairs in the 3D model is shown in gray. Crosslinks mapped onto the front view of the 3D structure are shown on the right. b, Monoubiquitination assays were assembled in the presence of 5 μM linear dsDNA of differing lengths (10–44 bp). DNA binding was analyzed by EMSA (top) after loading the reactions onto native gels and imaging of the fluorescently labeled DNA. Monoubiquitination efficiency was analyzed by Coomassie blue (middle) and western blotting the His-tagged ubiquitin (bottom). Controls lacking ubiquitin or DNA are indicated. These data are representative of experiments performed three times. c, Monoubiquitination assays were assembled without (left) and with (right) ubiquitin, both in the presence of a 39-bp dsDNA at 100 nM and increasing amounts of D2–I (0–1,000 nM). Assays were analyzed by EMSA (top, imaging for the fluorescently labeled DNA) or western blotting His-tagged ubiquitin (bottom). We cannot exclude that other proteins may interact with DNA in these assays, but the migration positions of the shifted bands are similar to the experiment in Fig. 4, in which ubD2–I was purified away from all other proteins. These data are representative of experiments performed three times. Uncropped images for b,c are available in Supplementary Fig. 1.
Extended Data Fig. 4 FANCD2 and FANCI oligomerization state and DNA binding activity.
a, Size-exclusion chromatogram as shown in Fig. 5a (top). Peak fractions were analyzed via SDS−PAGE (bottom). A single asterisk (*) indicates the migration position for monomers. A double asterisk (**) indicates the migration position for dimers. These results are representative of experiments performed three times. b, DNA binding of FANCI, FANCD2, D2–I and FANCI mixed with FANCD2 was analyzed in EMSAs performed with 20 nM 39-bp dsDNA and 0–140 nM protein. Gels shown are representative gels of experiments performed independently three times. The FANCD2 and D2–I gels are same as in Fig 5b. c, Quantification of mean intensities of free DNA from b. Error bars represent the standard deviation. Individual data points (n = 3 independent experiments) are shown. The means are connected by lines for clarity. Uncropped images for a,b are available in Supplementary Fig. 1. Data for the plot in c are available as Source data.
Extended Data Fig. 5 FANCD2 dimers cannot be ubiquitinated but they exchange with FANCI to form a D2–I heterodimer.
a, Monoubiquitination assays of D2–I and FANCD2 homodimer. The FANCD2 homodimer had a StrepII-tag and was therefore larger than D2 in the D2–I complex. Monoubiquitination efficiency was analyzed by Coomassie blue SDS−PAGE (top) and western blotting the His-tagged ubiquitin (bottom). b, Exchange assay. The FANCD2 homodimer was immobilized on Strep-Tactin resin and incubated with free FANCI. The resin was washed, then bound and unbound fractions were analyzed via by SDS−PAGE. These data are representative of experiments performed twice. Uncropped images are available in Supplementary Fig. 1.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2, Supplementary Tables 1 and 3 and Supplementary Fig. 1.
Supplementary Video 1
Motions detected by multibody refinement of ubD2–I: principal component analysis of multibody refinement results revealed two major motions.
Supplementary Video 2
Morph between D2–I and ubD2–I structures: a morph between the unmodified and modified D2–I complexes shows how FANCD2 and FANCI rotate around a hinge to clamp around DNA. In this video, the complexes are viewed from the back side of the hinge where ubiquitin is located.
Supplementary Video 3
Morph between D2–I and ubD2–I structures: a morph between the unmodified and modified D2–I complexes shows how FANCD2 and FANCI rotate around a hinge to clamp around DNA. In this video, the complexes are viewed from the DNA-binding groove.
Supplementary Video 4
Morph between D2–I and ubD2–I structures: a morph between the unmodified and modified D2–I complexes shows how FANCD2 and FANCI rotate around a hinge to clamp around DNA. In this video, the complexes are viewed from the top.
Supplementary Table 2
Crosslinking mass spectrometry data file.
Source data
Source Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 4
Statistical source data
Rights and permissions
About this article
Cite this article
Alcón, P., Shakeel, S., Chen, Z.A. et al. FANCD2–FANCI is a clamp stabilized on DNA by monoubiquitination of FANCD2 during DNA repair. Nat Struct Mol Biol 27, 240–248 (2020). https://doi.org/10.1038/s41594-020-0380-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0380-1
This article is cited by
-
FANCD2 as a novel prognostic biomarker correlated with immune and drug therapy in Hepatitis B-related hepatocellular carcinoma
European Journal of Medical Research (2023)
-
The structure-specific endonuclease complex SLX4–XPF regulates Tus–Ter-induced homologous recombination
Nature Structural & Molecular Biology (2022)
-
The key to the FANCD2–FANCI lock
Nature Structural & Molecular Biology (2022)
-
The DNA-damage kinase ATR activates the FANCD2-FANCI clamp by priming it for ubiquitination
Nature Structural & Molecular Biology (2022)
-
Screening and identification of novel candidate biomarkers of focal cortical dysplasia type II via bioinformatics analysis
Child's Nervous System (2022)