Abstract
We developed EASI-tag (easily abstractable sulfoxide-based isobaric-tag), a new type of amine-derivatizing and sulfoxide-containing isobaric labeling reagents for highly accurate quantitative proteomics analysis using mass spectrometry. We observed that EASI-tag labels dissociate at low collision energy and generate peptide-coupled, interference-free reporter ions with high yield. Efficient isolation of 12C precursors and quantification at the MS2 level allowed accurate determination of quantitative differences between up to six multiplexed samples.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Larance, M. & Lamond, A. I. Nat. Rev. Mol. Cell Biol. 16, 269–280 (2015).
Lössl, P., van de Waterbeemd, M. & Heck, A. J. EMBO J. 35, 2634–2657 (2016).
Aebersold, R. & Mann, M. Nature 537, 347–355 (2016).
Rauniyar, N. & Yates, J. R.III. J. Proteome Res. 13, 5293–5309 (2014).
Richards, A. L., Merrill, A. E. & Coon, J. J. Curr. Opin. Chem. Biol. 24, 11–17 (2015).
Bantscheff, M., Lemeer, S., Savitski, M. M. & Kuster, B. Anal. Bioanal. Chem. 404, 939–965 (2012).
Thompson, A. et al. Anal. Chem. 75, 1895–1904 (2003).
Ross, P. L. et al. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Bantscheff, M. et al. Mol. Cell. Proteomics 7, 1702–1713 (2008).
Karp, N. A. et al. Mol. Cell. Proteomics 9, 1885–1897 (2010).
Ting, L., Rad, R., Gygi, S. P. & Haas, W. Nat. Methods 8, 937–940 (2011).
Wenger, C. D. et al. Nat. Methods 8, 933–935 (2011).
Pichler, P. et al. Anal. Chem. 82, 6549–6558 (2010).
Wühr, M. et al. Anal. Chem. 84, 9214–9221 (2012).
Sonnett, M., Yeung, E. & Wühr, M. Anal. Chem. 90, 5032–5039 (2018).
Kao, A. et al. Mol. Cell. Proteomics 10, M110.002212 (2011).
Stadlmeier, M., Bogena, J., Wallner, M., Wühr, M. & Carell, T. Angew. Chem. Int. Edn. Engl. 57, 2958–2962 (2018).
Cox, J. & Mann, M. Nat. Biotechnol. 26, 1367–1372 (2008).
Erickson, B. K. et al. Mol. Cell 65, 361–370 (2017).
Meissner, F., Scheltema, R. A., Mollenkopf, H. J. & Mann, M. Science 340, 475–478 (2013).
Kulak, N. A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Nat. Methods 11, 319–324 (2014).
Werner, T. et al. Anal. Chem. 86, 3594–3601 (2014).
Rappsilber, J., Mann, M. & Ishihama, Y. Nat. Protoc. 2, 1896–1906 (2007).
Kulak, N. A., Geyer, P. E. & Mann, M. Mol. Cell. Proteomics 16, 694–705 (2017).
Tyanova, S. et al. Nat. Methods 13, 731–740 (2016).
R Development Core Team. R: A Language and Environment for Statistical Computing. (The R Foundation, Vienna, Austria, 2008).
Vizcaíno, J. A. et al. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
We acknowledge all members of the Department of Proteomics and Signal Transduction for help and valuable discussions. We thank P. Geyer, S. Doll, and A. Strasser for assistance with the experiments, and G. Sowa, I. Paron, and K. Mayr for technical support. This study was partially supported by funding from the German Research Foundation (DFG/Gottfried Wilhelm Leibniz Prize) (F. Meier, M.M.), European Union’s Horizon 2020 research and innovation program (grant agreement no. 686547; MSmed Project (S.V.W., F. Meier, C.W., J.C., M.M., F. Meissner), and the Max-Planck Society for the Advancement of Sciences (S.V.W., F. Meier, C.W., J.C., M.M., F. Meissner).
Author information
Authors and Affiliations
Contributions
F. Meissner and M.M. conceived the study. F. Meissner and S.V.W. conceptualized and designed the EASI-tag. F. Meier, F. Meissner, and M.M. conceptualized the acquisition method, and C.W. contributed to its implementation. S.V.W., F. Meier, and C.W. performed the experiments. J.C. developed and implemented data analysis software. S.V.W., F. Meier, F. Meissner, and M.M. analyzed the data. S.V.W., F. Meier, F. Meissner, and M.M. wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
S.V.W., F. Meier, M.M., and F. Meissner are inventors on a patent application related to the use of EASI-tag in quantitative proteomics (application number PCT/EP2017/079211).
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Fragmentation of asymmetric and symmetric sulfoxide moieties.
Fragments generated upon HCD-induced cleavage of an asymmetric sulfoxide site as it is present in EASI-tag (left panel) and of a symmetric sulfoxide site as it is present in MS-cleavable crosslinkers (right panel). Here, ‘asymmetry’ refers to the different chemical constitution of the residues on either side of the sulfoxide.
Supplementary Figure 2 Charge state distribution of unlabeled and EASI-tag-labeled peptides.
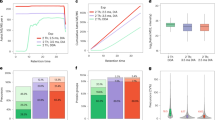
Distribution of charge states for all features detected by MaxQuant for unlabeled HeLa peptides and EASI-tag-labeled peptides from the one and two proteome experiments for (N=9 for unlabeled and N=3 for EASI-tag-labeled). The center bars indicate the mean values.
Supplementary Figure 3 Optimization of the asymmetric isolation window by analyses of tryptic HeLa digests.
(a) Abundance of the M+1 isotopic peak in MS/MS scans relative to the monoisotopic precursor ion as a function of the isolation window offset for an isolation width of 0.4 Th and only doubly charged precursor ions (N=4,413) (b) As above but only for triply charged precursor ions (N=2,182). (c) Abundance of the isolated precursor ion in MS/MS scans relative to its abundance when isolated without offset and an 1.4 Th window width (N=5468) (d) RawOvFtT in MS/MS scans relative to the RawOvFtT values with an isolation width of 1.4 Th and without an offset (N=5468) (e) Number of identified MS/MS scans relative to the number of identified MS/MS scans with an 1.4 Th isolation window width and without an offset (N=3). The center bars indicate the mean values an error bars show the standard deviation. Boxplot elements: center line = median, lower and upper box limits = first and third quartiles, whiskers = maximum 1.5x inter-quartile range.
Supplementary Figure 4 Comparison of label fragmentation versus peptide backbone fragmentation.
(a) EASI- and TMT-labeled HeLa peptides were fragmented with normalized collision energies between 10 and 34. ΔNCE was calculated as the difference of label fragmentation versus peptide backbone fragmentation (N = 10,321 precursors for EASI and 12,875 for TMT). Boxplot elements: center line = median, lower and upper box limits = first and third quartiles, whiskers = maximum 1.5x inter-quartile range.
Supplementary Figure 5 Analysis of in-source fragmentation of EASI-tag-labeled HeLa digests.
(a) EASI-tag-zero labeled HeLa peptides were fractioned by high pH reversed-phase and analyzed by LC-MS/MS. The relative number of features with a detectable in-source fragmentation resulting in fragmentation of EASI-tag was determined for each fraction (N=8 raw files and a total of 917,235 features). (b) The intensities of the in-source-fragmented features were determined for each fraction in relation to their unfragmented counterpart (N = 8 raw files and a total of 8,839 features).
Supplementary Figure 6 Distribution of ratios and coefficients of variation for EASI-tagged peptides.
(a) Distribution of peptide-coupled reporter ion ratios (log10) in MS/MS scans of yeast (mixing ratio 1:3:10:10:3:1) and human (mixing ratio 1:1:1:1:1:1) peptides (N=369,591, 375,384, 377,496, 377,839 and 378,101 for human channels 0 to 4 and 224,098, 209,244, 146,253, 152,255 and 215,707 for yeast channels 0 to 4). (b) Distribution of the coefficients of variation for EASI-tag quantified yeast and human peptides in three replicate injections (N= 39,400, 39,827, 39,898, 39,930 and 39,882 unique sequences for human channels 0 to 4 and 20,738, 19,947, 15,657, 16,097 and 20,230 unique sequences for yeast channels 0 to 4).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1 and Supplementary Note 1
Supplementary Data 1
Proteomics data supporting Fig. 1e and Supplementary Fig. 4
Supplementary Data 2
Proteomics data supporting Fig. 2a–c
Supplementary Data 3
Proteomics data supporting Fig. 2d–h
Supplementary Data 4
Proteomics data supporting Supplementary Figs. 2 and 3e
Supplementary Data 5
Proteomics data supporting Supplementary Fig. 5
Supplementary Data 6
Proteomics data supporting Supplementary Fig. 6
Rights and permissions
About this article
Cite this article
Virreira Winter, S., Meier, F., Wichmann, C. et al. EASI-tag enables accurate multiplexed and interference-free MS2-based proteome quantification. Nat Methods 15, 527–530 (2018). https://doi.org/10.1038/s41592-018-0037-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0037-8
This article is cited by
-
Detecting diagnostic features in MS/MS spectra of post-translationally modified peptides
Nature Communications (2023)
-
Deep and fast label-free Dynamic Organellar Mapping
Nature Communications (2023)
-
Development of novel isobaric tags enables accurate and sensitive multiplexed proteomics using complementary ions
Analytical and Bioanalytical Chemistry (2023)
-
The emerging role of mass spectrometry-based proteomics in drug discovery
Nature Reviews Drug Discovery (2022)
-
Light contamination in stable isotope-labelled internal peptide standards is frequent and a potential source of false discovery and quantitation error in proteomics
Analytical and Bioanalytical Chemistry (2022)