Abstract
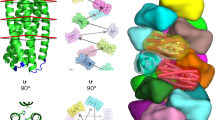
Chemists have long sought the ability to modify molecules precisely when presented with several sites of similar reactivity. We reasoned that the confinement of substrates within nanostructures might permit site-selective reactions unachievable in bulk solution, even with sophisticated reagents. In particular, the stretching and alignment of polymers within nanotubes might allow site-specific cleavage or modification. To explore this proposition, macromolecular disulfide substrates were elongated within members of a collection of tubular protein nanoreactors, which contained cysteine residues positioned at different locations along the length of each tube. For each nanoreactor, we defined the reactive location by using a set of polymer substrates (site-selectivity) and which of the two sulfur atoms was attacked (regioselectivity), and found that disulfide interchange occurs with atomic precision. Our strategy has potential for the selective processing of a wide variety of biomacromolecules, and the chemistry and substrates might be generalized yet further by using alternative nanotubes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the plots within this paper and other findings of this study are available from the corresponding author upon reasonable request.
References
Davis, H. J. & Phipps, R. J. Harnessing non-covalent interactions to exert control over regioselectivity and site-selectivity in catalytic reactions. Chem. Sci. 8, 864–877 (2017).
Mahatthananchai, J., Dumas, A. M. & Bode, J. W. Catalytic selective synthesis. Angew. Chem. Int. Ed. 51, 10954–10990 (2012).
Rousseau, G. & Breit, B. Removable directing groups in organic synthesis and catalysis. Angew. Chem. Int. Ed. 50, 2450–2494 (2011).
Wang, J., Li, G. & Reetz, M. T. Enzymatic site-selectivity enabled by structure-guided directed evolution. Chem. Commun. 53, 3916–3928 (2017).
Dodani, S. C. et al. Discovery of a regioselectivity switch in nitrating P450s guided by molecular dynamics simulations and Markov models. Nat. Chem. 8, 419–425 (2016).
Su, D. & Jaeger, D. A. High regioselectivity in the Diels–Alder reaction of a surfactant 1,3-diene with a surfactant dienophile resulting from a short tether between their functional groups and head groups. Tetrahedron Lett. 40, 7871–7874 (1999).
Yoshizawa, M., Tamura, M. & Fujita, M. Diels–Alder in aqueous molecular hosts: unusual regioselectivity and efficient catalysis. Science 312, 251–254 (2006).
Nakanishi, Y. et al. Template synthesis of linear-chain nanodiamonds inside carbon nanotubes from bridgehead-halogenated diamantane precursors. Angew. Chem. Int. Ed. 54, 10802–10806 (2015).
Miners, S. A., Rance, G. A. & Khlobystov, A. N. Regioselective control of aromatic halogenation reactions in carbon nanotube nanoreactors. Chem. Commun. 49, 5586 (2013).
Bayley, H., Luchian, T., Shin, S.-H. & Steffensen, M. B. in Single Molecules and Nanotechnology Springer Series in Biophysics 12 (eds Rigler, R. & Vogel, H.) 251–277 (Springer, 2008).
Alegre-Cebollada, J., Kosuri, P., Rivas-Pardo, J. A. & Fernández, J. M. Direct observation of disulfide isomerization in a single protein. Nat. Chem. 3, 882–887 (2011).
Beedle, A. E. M. et al. Forcing the reversibility of a mechanochemical reaction. Nat. Commun. 9, 3155 (2018).
Beedle, A. E. M., Mora, M., Lynham, S., Stirnemann, G. & Garcia-Manyes, S. Tailoring protein nanomechanics with chemical reactivity. Nat. Commun. 8, 15658 (2017).
Beedle, A. E. M., Lynham, S. & Garcia-Manyes, S. Protein S-sulfenylation is a fleeting molecular switch that regulates non-enzymatic oxidative folding. Nat. Commun. 7, 1–10 (2016).
Garcia-Manyes, S. & Beedle, A. E. M. Steering chemical reactions with force. Nat. Rev. Chem. 1, 0083 (2017).
Chivers, C. E. et al. A streptavidin variant with slower biotin dissociation and increased mechanostability. Nat. Methods 7, 391–393 (2010).
Murzin, A. G., Lesk, A. M. & Chothia, C. Principles determining the structure of beta-sheet barrels in proteins. J. Mol. Biol. 236, 1382–1400 (1994).
Song, L. et al. Structure of staphylococcal α-hemolysin, a heptameric transmembrane pore. Science 274, 1859–1866 (1996).
Stoddart, D., Heron, A. J., Mikhailova, E., Maglia, G. & Bayley, H. Single-nucleotide discrimination in immobilized DNA oligonucleotides with a biological nanopore. Proc. Natl Acad. Sci. USA 106, 7702–7707 (2009).
Fernandes, P. A. & Ramos, M. J. Theoretical insights into the mechanism for thiol/disulfide exchange. Chem. Eur. J. 10, 257–266 (2004).
Bach, R. D., Dmitrenko, O. & Thorpe, C. Mechanism of thiolate-disulfide interchange reactions in biochemistry. J. Org. Chem. 73, 12–21 (2008).
Howorka, S. & Bayley, H. Probing distance and electrical potential within a protein pore with tethered DNA. Biophys. J. 83, 3202–3210 (2002).
Bosco, A., Camunas-Soler, J. & Ritort, F. Elastic properties and secondary structure formation of single-stranded DNA at monovalent and divalent salt conditions. Nucleic Acids Res. 42, 2064–2074 (2014).
Whitesides, G. M., Lilburn, J. E. & Szajewski, R. P. Rates of thiol–disulfide interchange reactions between mono and dithiols and Ellman’s reagent. J. Org. Chem. 42, 332–338 (1977).
Qin, F., Auerbach, A. & Sachs, F. Estimating single-channel kinetic parameters from idealized patch-clamp data containing missed events. Biophys. J. 70, 264–280 (1996).
Pace, C. N., Grimsley, G. R. & Scholtz, J. M. Protein ionizable groups: pK values and their contribution to protein stability and solubility. J. Biol. Chem. 284, 13285–13289 (2009).
Depuydt, M., Messens, J. & Collet, J.-F. How proteins form disulfide bonds. Antioxid. Redox Signal. 15, 49–66 (2011).
Ukuwela, A. A., Bush, A. I., Wedd, A. G. & Xiao, Z. Glutaredoxins employ parallel monothiol–dithiol mechanisms to catalyze thiol–disulfide exchanges with protein disulfides. Chem. Sci. 9, 1173–1183 (2018).
Luchian, T., Shin, S.-H. & Bayley, H. Single-molecule covalent chemistry with spatially separated reactants. Angew. Chem. Int. Ed. 42, 3766–3771 (2003).
Qing, Y., Ionescu, S. A., Pulcu, G. S. & Bayley, H. Directional control of a processive molecular hopper. Science 361, 908–912 (2018).
Lu, B., Fleming, S., Szalay, T. & Golovchenko, J. Thermal motion of DNA in an MspA pore. Biophys. J. 109, 1439–1445 (2015).
Bayley, H. Nanopore sequencing: from imagination to reality. Clin. Chem. 61, 25–31 (2015).
Huxley, M. T. et al. Protecting-group-free site-selective reactions in a metal–organic framework reaction vessel. J. Am. Chem. Soc. 140, 6416–6425 (2018).
Acknowledgements
We thank L. Taemaitree for her help with LC–MS characterization. We also thank M. Booth for useful discussions and M. Howarth for providing a sample of traptavidin for preliminary tests. This research was supported by a European Research Council Advanced Grant (COSIMO). Y.Q. was supported by a China Scholarship Council-University of Oxford Scholarship. S.A.I. was funded by an Amelia Jackson Studentship, Exeter College, Oxford.
Author information
Authors and Affiliations
Contributions
Y.Q. and H.B. conceived the research. Y.Q. performed the experiments. H.T.-A. contributed to the solid-phase peptide synthesis. S.A.I. and M.D.L. contributed to the electrical measurements. Y.Q., S.A.I. and H.B. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
Y.Q. and H.B. have filed patents describing the molecular hopper and applications thereof. The hopper is ratcheted by regioselective thiol–disulfide interchange.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary notes, Table 1 and 2, Figs. 1–9 and refs. 1–10.
Rights and permissions
About this article
Cite this article
Qing, Y., Tamagaki-Asahina, H., Ionescu, S.A. et al. Catalytic site-selective substrate processing within a tubular nanoreactor. Nat. Nanotechnol. 14, 1135–1142 (2019). https://doi.org/10.1038/s41565-019-0579-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-019-0579-7
This article is cited by
-
Highly shape- and size-tunable membrane nanopores made with DNA
Nature Nanotechnology (2022)
-
A reversibly gated protein-transporting membrane channel made of DNA
Nature Communications (2022)
-
Nanopore-based technologies beyond DNA sequencing
Nature Nanotechnology (2022)
-
Remodeling nanodroplets into hierarchical mesoporous silica nanoreactors with multiple chambers
Nature Communications (2022)
-
Towards explicit regulating-ion-transport: nanochannels with only function-elements at outer-surface
Nature Communications (2021)