Abstract
Arising from: G. P. Wagner et al. Nature 452, 470–472 (2008)10.1038/nature06756; Wagner et al. reply
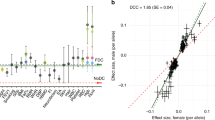
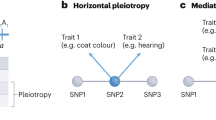
Wagner et al.1 have recently introduced much-needed data to the debate on how complexity of the genotype–phenotype map affects the distribution of mutational effects. They used quantitative trait loci (QTLs) mapping analysis of 70 skeletal characters in mice2 and regressed the total QTL effect on the number of traits affected (level of pleiotropy). From their results they suggest that mutations with higher pleiotropy have a larger effect, on average, on each of the affected traits—a surprising finding that contradicts previous models3,4,5,6,7. We argue that the possibility of some QTL regions containing multiple mutations, which was not considered by the authors, introduces a bias that can explain the discrepancy between one of the previously suggested models and the new data.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Wagner, G. P. et al. Pleiotropic scaling of gene effects and the ‘cost of complexity’. Nature 452, 470–473 (2008)
Kenney-Hunt, J. P. et al. Pleiotropic patterns of quantitative trait loci for 70 murine skeletal traits. Genetics 178, 2275–2288 (2008)
Turelli, M. Effects of pleiotropy on predictions concerning mutation-selection balance for polygenic traits. Genetics 111, 165–195 (1985)
Wagner, G. P. The influence of variation and developmental constraints on the rate of multivariate phenotypic evolution. J. Evol. Biol. 1, 45–66 (1988)
Orr, H. A. Adaptation and the cost of complexity. Evolution Int. J. Org. Evolution 54, 13–20 (2000)
Wingreen, N. S., Miller, J. & Cox, E. C. Scaling of mutational effects in models of pleiotropy. Genetics 164, 1221–1228 (2003)
Martin, G. & Lenormand, T. A general multivariate extension of Fisher’s geometrical model and the distribution of mutational effects across species. Evolution Int. J. Org. Evolution 60, 893–907 (2006)
Orgogozo, V., Broman, K. W. & Stern, D. L. High-resolution quantitative trait locus mapping reveals sign epistasis controlling ovariole number between two Drosophila species. Genetics 173, 197–205 (2006)
McGregor, A. P. et al. Morphological evolution through multiple cis-regulatory mutations at a single gene. Nature 448, 587–590 (2007)
Long, A. D., Lyman, R. F., Langley, C. H. & Mackay, T. F. C. Two sites in the Delta gene region contribute to naturally occurring variation in bristle number in Drosophila melanogaster . Genetics 149, 999–1017 (2000)
Author information
Authors and Affiliations
PowerPoint slides
Rights and permissions
About this article
Cite this article
Hermisson, J., McGregor, A. Pleiotropic scaling and QTL data. Nature 456, E3 (2008). https://doi.org/10.1038/nature07452
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07452
This article is cited by
-
Intermediate degrees of synergistic pleiotropy drive adaptive evolution in ecological time
Nature Ecology & Evolution (2017)
-
The pleiotropic structure of the genotype–phenotype map: the evolvability of complex organisms
Nature Reviews Genetics (2011)
-
Sexual Dimorphism Increases Evolvability in a Genetic Regulatory Network
Evolutionary Biology (2011)
-
The Developmental Basis of Variational Modularity: Insights from Quantitative Genetics, Morphometrics, and Developmental Biology
Evolutionary Biology (2009)
-
Wagner et al. reply
Nature (2008)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.