Abstract
Interstitial deletion of the proximal short arm of chromosome 10 represents a rare genetic alteration. Literature review revealed that only 10 postnatal diagnosed clinical cases with deletions overlapping 10p12p11 were published until present. We report the first prenatal diagnosis and postnatal findings in a male fetus with a 10.6 Mb interstitial deletion of the short arm of chromosome 10 (10p11.22-p12.31).
Similar content being viewed by others
Introduction
Although array comparative genomic hybridization (CGH) is nowadays an important diagnostic tool in postnatal genetic pathologies that associate congenital malformations and mental retardation, it has not been widely used in the field of prenatal genetic diagnostics.1 Here we report the prenatal diagnosis of an interstitial deletion located on the short arm of chromosome 10 in a male fetus with sonographic abnormal genetic features and abnormal maternal serum biochemistry. To our knowledge this is the first 10p interstitial deletion case identified prenatally, with amniotic fluid as the analyzed biologic specimen. All previously reported cases were diagnosed and characterized after birth. Prenatal diagnosis is important, as the developmental delay, craniofacial abnormalities, cardiac malformation, immune deficiencies and cryptorchidism are the main phenotypic features described in patients with deletions overlapping the proximal 10p11–p12 region on the short arm of chromosome 10.2, 3, 4
Materials and methods
Amniocentesis was performed on a 31-year-old woman, at 16 weeks of gestation, after second trimester genetic risk calculation revealed low MOMs (multiple of median) of alpha fetoprotein (AFP) (0.43) and human chorionic gonadotropin beta (hCGb) (0.57), 1.26 MOMs for unconjugated Estriol 3 (uE3) and increased risk for trisomy 21 (1/220). A previous scan at 12 gestational weeks (GW) had revealed increased fetal nuchal translucency (4.92 mm) and agenesis of ductus venosus (Figure 1). Invasive studies through chorionic villus sampling showed no numerical abnormalities of 13, 18, 21, X and Y chromosomes by quantitative fluorescence - polymerase chain reaction (QF-PCR). Both parents were healthy, with no family history of genetic diseases or congenital malformations.
First (a–d) and second trimester (e–i) ultrasound genetic markers are 12 and 15 gestational weeks scan, respectively. (a) Increased nuchal translucency; (b) present nasal bone; (c) normal tricuspid flow; (d) agenesis of ductus venosus in four-dimensional STIC power Doppler reconstruction; (e, f) abnormal facial sagittal plane, with micrognathiain two- and three-dimensional reconstruction of the fetal face; (g) echogenic intracardiac focus; (h) absence of the ductus venosus; and (i) clinodactily. H, heart; HV, hepatic veins; IVC, inferior vena cava; UA, umbilical arteries; UC, umbilical cord; UV, umbilical vein. A full color version of this figure is available at the Journal of Human Genetics journal online.
A detailed morphological evaluation was achieved during an early second trimester ultrasound scan at 15 GW. Sonographic soft markers for genetic abnormalities were present (Figure 1): agenesis of ductus venosus, echogenic intracardiac focus, clinodactily and micrognathia.
The 16 GW cytogenetic analyses suggested a deletion on the short arm of chromosome 10 (46, XY, 10p). At 22 GW, the morphological ultrasound evaluation revealed no major malformations but abnormal ultrasound genetic markers were again present (Figure 2): nuchal and prefrontal edema, ‘flat face’, short femur and humerus and fetal growth restriction. Also, relative hypertelorism, persistent small stomach, persistent flexion of fingers and short feet were detected.
Supplementary ultrasound findings at the 22 GW scan (a–e) and postnatal features. (a) prefrontal edema; (b) hypertelorism; (c) increased nuchal fold; (d) persistent small stomach (arrow); (e) small and wide feet and abnormal persistent flexion of the hand fingers; (f, g) anterior and lateral aspect of the newborn. A full color version of this figure is available at the Journal of Human Genetics journal online.
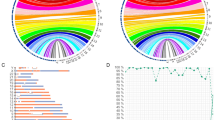
Another amniocentesis was performed at 22 GW. The routine G-band analysis of cultured amniocytes depicted a 10p deletion. The result was further validated through array CGH (Figure 3), which accurately detected the genomic size and gene content of the deleted region.
Array CGH result on chromosome 10. Image generated with Nexus 6.1 software (Nexus BioDiscovery, El Segundo, CA, USA). Data extraction (signal intensities) was performed with DEVA software (Roche Nimblegen). A full color version of this figure is available at the Journal of Human Genetics journal online.
After genetic counseling, the parents elected to continue the pregnancy. A late premature boy, with intrauterine growth restriction, was delivered by cesarean section at 36 GW. Apgar scores were 6 and 7 at 1 and 5 min, respectively. The neonatal evaluation revealed dysmorphic features including frontal and parietal bossing with hypoplastic facial bones leading to a triangular face, aged face, low and flat nasal bridge, short nose with a bulbous nasal tip, hypertelorism, deeply set eyes, low-set ears with dysplastic helices, thin upper lip, trismus, short trunk with a grade of contractures of thoracic cage, reflected in reduced amplitude of breathing movements, short broad neck, hands in ulnar deviation, camptodactyly and cryptorchidism (Figure 2). The newborn presented fever syndrome of unknown etiology, intermittent hypoglycemic, feeding difficulties and respiratory problems related to superior airways hypoplasia. The growth curve was slow ascendant, 380 g gain in 8 weeks. The neonatal death occurred at the age of 8 weeks, due to an acute respiratory failure exacerbated by a respiratory tract infection. The autopsy ascertained growth restriction and did not reveal other congenital malformations.
Results
The array CGH analysis, using a whole-genome 135 K oligonucleotide microarray platform (Roche NimbleGen, Madison, WI, USA), detected a loss in 10p11.22—p12.31, encompassing a 10.6 Mb region (chr10:21,461,075–32,075,988, hg19) within the proximal short arm of chromosome 10.
Discussion
The above mentioned panel of dysmorphic features detected by ultrasound might be considered as a pattern for the prenatal diagnosis of interstitial deletion of chromosome 10, but further cases with similar prenatal findings are expected to sustain our hypothesis. Craniofacial dysmorphic features like deep-set eyes or bulbous nose were shared by our patient and seven of the previously diagnosed cases.3, 4, 5 Comparative analysis of the array CGH results in these eight patients revealed that the overlapping region has a length of 401 kb and among the contained genes are BAMBI or WAC, suggested to be involved in the genetic etiology of developmental delay and craniofacial dysmorphic features like deeply set eyes, synophrys or bulbous nasal tip.2, 4 Cardiac abnormalities like conotruncal heart defects described in 7 out of 10 published cases with 10p interstitial deletion have not been found in our patient despite the fact that he shared the deletion of LYZL1 and SVIL genes.
Cryptorchidism reported in three out of five reported males with interstitial 10p deletion was also detected in our patient that shared the deletion of MKX gene found in the previous three cases, too.2, 4 These findings suggest that MKX gene haploinsufficiency could be involved in genetic susceptibility to cryptorchidism development.
Comparative analyses of array CGH coupled with fine mapping of deleted regions, showed that our case and other 9 patients share an overlapping 711 kb deletion (chr10: 28802788-28091602—hg18). The deleted region contains two protein-encoding genes: ARMC4 (armadillo repeat containing 4) and MPP7 (membrane protein, palmitoylated 7). It is therefore possible that defects of these genes be responsible for craniofacial dysmorphism associated or not with developmental disorders and mental impairment.
We report, to our knowledge, the first prenatal case with a deletion at 10p12p11 diagnosed through integrated cytogenetic and genomic analyses. Our case together with the 10 previously published cases shared a similar clinical appearance characterized by developmental delay and craniofacial dysmorphic features, prompting us to define a genotype–phenotype correlation in order to sustain the statement that proximal 10p deletion is a contiguous gene syndrome.
References
Kleeman, L., Bianchi, D. W., Shaffer, L. G., Rorem, E., Cowan, J., Craigo, S. D. et al. Use of array comparative genomic hybridization for prenatal diagnosis of fetuses with sonographic anomalies and normal metaphase karyotype. Prenat. Diagn. 29, 1213–1217 (2009).
Mroczkowski, H. J., Arnold, G., Schneck, F. X., Rajkovic, A. & Yatsenko, S. A. Interstitial 10p11.23-p12.1 microdeletions associated with developmental delay, craniofacial abnormalities, and cryptorchidism. Am. J. Med. Genet. A 164A, 2623–2626 (2014).
Okamoto, N., Hayashi, S., Masui, A., Kosaki, R., Oguri, I., Hasegawa, T. et al. Deletion at chromosome 10p11.23-p12.1 defines characteristic phenotypes with marked midface retrusion. J. Hum. Genet. 57, 191–196 (2012).
Wentzel, C., Rajcan-Separovic, E., Ruivenkamp, C. A., Chantot-Bastaraud, S., Metay, C., Andrieux, J. et al. Genomic and clinical characteristics of six patients with partially overlapping interstitial deletions at 10p12p11. Eur. J. Hum. Genet. 19, 959–964 (2011).
Shahdadpuri, R., de Vries, B., Pfundt, R., de Leeuw, N. & Reardon, W. Pseudoarthrosis of the clavicle and copper beaten skull associated with chromosome 10p11.21p12.1 microdeletion. Am. J. Med. Genet. A 146A, 233–237 (2008).
Acknowledgements
For this paper IS was supported within the frame of European Social Found, Human Resources Development Operational Program 2007–2013, project no. POSDRU/159/1.5/S/136893.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Sosoi, S., Streata, I., Tudorache, S. et al. Prenatal and postnatal findings in a 10.6 Mb interstitial deletion at 10p11.22-p12.31. J Hum Genet 60, 183–185 (2015). https://doi.org/10.1038/jhg.2015.4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.4