Abstract
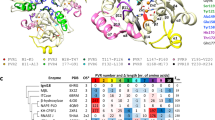
We report the crystal structure of a class D β-lactamase, the broad spectrum enzyme OXA-10 from Pseudomonas aeruginosa at 2.0 Å resolution. There are significant differences between the overall fold observed in this structure and those of the evolutionarily related class A and class C β-lactamases. Furthermore, the structure suggests the unique, cation mediated formation of a homodimer. Kinetic and hydrodynamic data shows that the dimer is a relevant species in solution and is the more active form of the enzyme. Comparison of the molecular details of the active sites of the class A and class C enzymes with the OXA-10 structure reveals that there is no counterpart in OXA-10 to the residues proposed to act as general bases in either of these enzymes (Glu 166 and Tyr 150, respectively). Our structures of the native and chloride inhibited forms of OXA-10 suggest that the class D enzymes have evolved a distinct catalytic mechanism for β-lactam hydrolysis. Clinical variants of OXA-10 are also discussed in light of the structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Frère, J.M. Beta-lactamases and bacterial resistance to antibiotics. Mol. Microbiol. 16, 385–395 (1995).
Joris, B., et al. Comparison of the sequences of class A beta-lactamases and of the secondary structure elements of penicillin-recognizing proteins. Antimicrob. Agents Chemother. 35, 2294–2301 (1991).
Frère, J.M., Dubus, A., Galleni, M., Matagne, A. & Amicosante, G. Mechanistic diversity of beta-lactamases. Biochem. Soc. Trans. 27, 58–63 (1999).
Bush, K. Metallo-beta-lactamases: a class apart. Clin. Infect. Dis. 27 (Suppl 1), S48–53 (1998).
Knox, J.R., Moews, P.C. & Frère, J.M. Molecular evolution of bacterial beta-lactam resistance. Chem. Biol. 3, 937–947 (1996).
Gordon, E., Mouz, N., Duée, E. & Dideberg, O. The crystal structure of the penicillin-binding protein 2x from Streptoccus pneumoniae and its acyl-enzyme form: implication in drug resistance. J. Mol. Biol. 299, 477–485 (2000).
Nass, T. & Nordmann, P. OXA-type beta-lactamases. Curr. Pharm. Design 5, 865–879 (1999).
Mugnier, P., Casin, I., Bouthers, A.T. & Collatz, E. Novel OXA-10-derived extended-spectrum beta-lactamases selected in vivo or in vitro. Antimicrob. Agents Chemother. 42, 3113–3116 (1998).
Bush, K. The evolution of beta-lactamases. Ciba Found. Symp. 207, 152–163 (1997).
Strynadka, N.C.J., et al. Molecular structure of the acyl-enzyme intermediate in beta-lactam hydrolysis at 1.7 Å resolution. Nature 359, 700–705 (1992).
Jelsch, C., Mourey, L., Masson, J-M. & Samama, J-P. Crystal structure of Escherichia coli TEM1 beta-lactamase at 1.8 Å resolution. Proteins 16, 364–383 (1993).
Oefner, C., et al. Refined crystal structure of beta-lactamase from Citrobacter freundii indicates a mechanism for beta-lactam hydrolysis. Nature 343, 284–289 (1990).
Monaghan, C., Holland, S. & Dale, J.W. The interaction of anthraquinone dyes with the plasmid-mediated OXA-2 beta-lactamase. Biochem. J. 205, 413–417 (1982).
Sanschagrin, F., Couture, F. & Levesque, R.C. Primary structure of OXA-3 and phylogeny of oxacillin-hydrolyzing class D beta-lactamases. Antimicrob. Agents Chemother. 39, 887–893 (1995).
Glusker, J.P. Structural aspects of metal liganding to functional groups in proteins. Adv. Protein Chem. 42, 1–76 (1991).
Dale J.W. & Smith, J.T. The dimeric nature of an R-factor mediated beta-lactamase. Biochem. Biophys. Res. Commun. 68, 1000–1005 (1976).
Danel, F., Hall, L.M., Gur, D. & Livermore, D.M. OXA-16, a further extended-spectrum variant of OXA-10 beta-lactamase, from two Pseudomonas aeruginosa isolates. Antimicrob. Agents Chemother. 42, 3117–3122 (1998).
Lobkovsky, E., et al. Crystallographic structure of a phosphonate derivative of the Enterobacter cloacae P99 cephalosporinase: mechanistic interpretation of a beta-lactamase transition-state analog. Biochemistry 33, 6762–6772 (1994).
Maveyraud, L., Pratt, R.F. & Samama, J.P. Crystal structure of an acylation transition-state analog of the TEM-1 beta-lactamase. Mechanistic implications for class A beta-lactamases. Biochemistry 37, 2622–2628 (1998).
Zhu, Y., et al. Structure, function, and fate of the BlaR signal transducer involved in induction of beta-lactamase in Bacillus licheniformis. J. Bacteriol. 174, 6171–6178 (1992).
Jacob-Dubuisson, F., Lamotte-Brasseur J., Dideberg, O., Joris, B. & Frere, J.M. Arginine 220 is a critical residue for the catalytic mechanism of the Streptomyces albus G beta-lactamase. Protein Eng. 4, 811–819 (1991).
Dubus, A., Normark, S., Kania, K. & Page, M.G.P. The role of tyrosine 150 in catalysis of β-lactam hydrolysis by AmpC β-lactamase from Escherichia coli investigated by site-directed mutagenesis. Biochemistry 33, 8577–8586 (1994).
Ledent, P. Raquet, X., Joris, B., Van Beeumen, J. & Frere J.M. A comparative study of class-D beta-lactamases. Biochem. J. 292, 555–562 (1993).
Little, J.W., et al. Cleavage of LexA repressor. Methods Enzymol. 244, 266–284 (1994).
Paetzel, M., et al. Use of site-directed chemical modification to study an essential lysine in Escherichia coli leader peptidase. J. Biol. Chem. 272, 9994–10003 (1997).
Patricelli, M.P. & Cravatt, B.F. Fatty acid amide hydrolase competitively degrades bioactive amides and esters through a nonconventional catalytic mechanism. Biochemistry 38, 14125–14130 (1999).
Birghan, C., Mundt, E. & Gorbalenya, A.E. A non-canonical lon proteinase lacking the ATPase domain employs the Ser-Lys catalytic dyad to exercise broad control over the life cycle of a double-stranded RNA virus. EMBO J. 19, 114–123 (2000).
Keiler, K.C. & Sauer, R.T. Identification of active site residues of the Tsp protease. J. Biol. Chem. 270, 28864–28868 (1995).
Haase, J. & Lanka, E. A specific protease encoded by the conjugative DNA transfer systems of IncP and Ti plasmids is essential for pilus synthesis. J. Bacteriol. 179, 5728–5735 (1997).
Lietz, E.J., Truher, H., Kahn, D., Hokenson, M.J. & Fink, A.L. Lysine-73 is involved in the acylation and deacylation of beta-lactamase. Biochemistry 39, 4971–4981 (2000).
Damblon, C., et al. The catalytic mechanism of beta-lactamases: NMR titration of an active-site lysine residue of the TEM-1 enzyme. Proc. Natl. Acad. Sci. USA 93, 1747–1752 (1996).
Philippon, A.M., Paul G. & Jacoby, G.A. Properties of PSE-2 beta-lactamase and genetic basis for its production in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 24, 362–369 (1983).
Bush, K., Jacoby G.A., Medeiros, A.A. A functional classification scheme for beta-lactamases and its correlation with molecular structure. Antimicrob. Agents Chemother. 39, 1211–1233 (1995).
Bilwes, A., Rees, B., Moras, D., Menez, R. & Menez, A. X-ray structure at 1.55 Å of toxin gamma, a cardiotoxin from Naja nigricollis venom. Crystal packing reveals a model for insertion into membranes. J. Mol. Biol. 239,122–136 (1994).
Otwinowski, Z. In Denzo (eds, Sawyer, L., Isaacs, N. & Baily, S.) 56–62 (SERC Daresbury Laboratory, Warrington, UK; 1993).
Terwilliger, T.C. & Berendzen, J. Automated structure solution for MIR and MAD. Acta Crystallogr. D 55, 849–861 (1999).
La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement in the MIR and MAD methods Methods Enzymol. 276, 472–494 (1997).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kieldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, S. & Thornton, J.M. Protein-protein interactions: a review of protein dimer structure. Prog. Biophys. Mol. Biol. 63, 31–165 (1995).
Cohn, E.J. & Edsall, J.T. Density and apparent specific volume of proteins. In Proteins, amino acids and peptides as ions and dipolar ions (ed. E. J. Cohn), 370–381 (Reinhold, New York; 1943).
Schuck, P. Simultaneous radial and wavelength analysis with the Optima XL-A analytical ultracentrifuge. Prog. Colloid Polymer Sci. 94, 1–13 (1994).
Kraulis, P. G. Molscript: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallog. 24, 946–950 (1991).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Couture, F., Lachapelle, J. & Levesque, R.C. Phylogeny of LCR-1 and OXA-5 with class A and class D beta-lactamases. Mol. Microbiol. 6, 1693–1705 (1992).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Acknowledgements
We thank the Medical Research Council of Canada (M.P. is a MRC fellow, N.C.J.S. is a MRC scholar), the Burroughs Wellcome Foundation (New Investigator Award to N.C.J.S.) and Hoffman-La Roche Pharmaceuticals (to F.D. and M.G.P.P.) for support. We thank H. Bellamy of beamline 1-5 at the SSRL for data collection access and R. Sweet for access to beamline X12C at the NSLS (Brookhaven National Laboratory).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Paetzel, M., Danel, F., de Castro, L. et al. Crystal structure of the class D β-lactamase OXA-10. Nat Struct Mol Biol 7, 918–925 (2000). https://doi.org/10.1038/79688
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/79688
This article is cited by
-
Repurposing Amphotericin B: anti-microbial, molecular docking and molecular dynamics simulation studies suggest inhibition potential of Amphotericin B against MRSA
BMC Chemistry (2023)
-
Two non-active site residues W165 and L166 prominently influence the beta-lactam hydrolytic ability of OXA-23 beta-lactamase
The Journal of Antibiotics (2023)
-
Molecular diversity of extended-spectrum β-lactamases and carbapenemases, and antimicrobial resistance
Journal of Intensive Care (2020)
-
New eight genes identified at the clinical multidrug-resistant Acinetobacter baumannii DMS06669 strain in a Vietnam hospital
Annals of Clinical Microbiology and Antimicrobials (2017)
-
In vivo protein interaction network analysis reveals porin-localized antibiotic inactivation in Acinetobacter baumannii strain AB5075
Nature Communications (2016)