Abstract
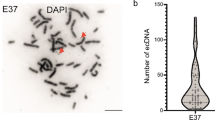
To elucidate the reasons why mRNA expression of the LGI1 candidate tumor-suppressor gene was severely reduced in the glioma-derived cell line H4, as demonstrated in a previous study, we performed a cytogenetic analysis of this cell line using conventional methods and fluorescence in situ hybridization (FISH) techniques [spectral karyotyping (SKY), interphase- and chromosome FISH of metaphases (I- and C-FISH)]. Cell line H4 is monoclonal and near triploid (±3n). SKY enabled us to detect 24 structural aberrations: unbalanced translocations, n = 12; deletions, n = 10; insertion, n = 1; duplication, n = 1. The results were confirmed by I- and C-FISH analysis using chromosome-specific paints, centromer-specific probes and locus-specific probes for p53, PTEN/MMAC1, LGI1, Cyclin D1, EGR1, ETV6/TEL, AML1, and the genomic region 13q14.3 containing the Rb locus. We found loss of one copy of p53 as well as of one copy of Rb. Complete loss of PTEN/MMAC1 was detected, while all copies of LGI1 and Cyclin D1 were preserved. Interestingly, there was a gain of ETV6/TEL and EGR1, which were each present in quadruplicate. Additionally, the AML1 locus revealed mosaicism of cells with three and four copies, respectively. Additionally, a 5q-chromosome [del(5)(q13q33)] was found, which is one of the common features in hematological malignancies, and der(12)t(1;12) was found, suggesting that there might be an additional ETV6/TEL fusion protein.
The combination of SKY, I- and C-FISH demonstrates that the neuroglioma cell line H4 harbors cytogenetic aberrations that are reported to occur in glioma-derived cell lines and additional chromosomal aberrations that have so far not been reported to occur in these cell lines. The complex aberrant karyotype and possibly generation of transcription factors by fusion proteins might be reasons for the impaired mRNA expression of the LGI1 candidate tumor-suppressor gene in cell line H4.
Similar content being viewed by others
References
Davis FG, Freels S, Grutsch J, Barlas S, Brem S: Survival rates in patients with primary malignant brain tumors stratified by patient age and tumor histological type: an analysis based on Surveillance, Epidemiology, and End Results (SEER) data, 1973-1991. J Neurosurg 88: 1–10, 1998
Jenkins RB, Kimmel DW, Moertel CA, Schultz CG, Scheithauer BW, Kelly PJ, Dewald GW: A cytogenetic study of 53 human gliomas. Canc Genet Cytogenet 39: 253–279, 1989
Schrock E, Thiel G, Lozanova T, du Manoir S, Meffert MC, Jauch A, Speicher MR, Nurnberg P, Vogel S, Janisch W et al.: Comparative genomic hybridization of human malignant gliomas reveals multiple amplification sites and nonrandom chromosomal gains and losses. Am J Pathol 144: 1203–1218, 1994
von Deimling A, Louis DN, Wiestler OD: Molecular pathways in the formation of gliomas. Glia 15: 328–338, 1995
Duerr EM, Rollbrocker B, Hayashi Y, Peters N, Meyer-Puttlitz B, Louis DN, Schramm J, Wiestler OD, Parsons R, Eng C, von Deimling A: PTEN mutations in gliomas and glioneuronal tumors. Oncogene 16: 2259–2264, 1998
Lin H, Bondy ML, Langford LA, Hess KR, Delclos GL, Wu X, Chan W, Pershouse MA, Yung WK, Steck PA: Allelic deletion analyses of MMAC/PTEN and DMBT1 loci in gliomas: relationship to prognostic significance. Clin Cancer Res 4: 2447–2454, 1998
Chernova OB, Somerville RP, Cowell JK: A novel gene, LGI1, from 10q24 is rearranged and downregulated in malignant brain tumors. Oncogene 17: 2873–2881, 1998
Arnstein P, Taylor DO, Nelson-Rees WA, Huebner RJ, Lennette EH: Propagation of human tumors in antithymocyte serum-treated mice. J Natl Cancer Inst 52: 71–84, 1974
Schrock E, du Manoir S, Veldman T, Schoell B, Wienberg J, Ferguson-Smith MA, Ning Y, Ledbetter DH, Bar-Am I, Soenksen D, Garini Y, Ried T: Multicolor spectral karyotyping of human chromosomes. Science 273: 494–497, 1996
Speicher M, Ward D: The coloring of cytogenetics. Nat Med 2: 1046–1048, 1996
Bigner DD, Bigner SH, Ponten J, Westermark B, Mahaley MS, Ruoslahti E, Herschman H, Eng LF, Wikstrand CJ: Heterogeneity of Genotypic and phenotypic characteristics of fifteen permanent cell lines derived from human gliomas. J Neuropathol Exp Neurol 40: 201–229, 1981
Takeshita I, Sawa H, Nakamura T, Kuramitsu M, Kitamura K, Fukui M: Contrary effect of lactic acid on expression of neuron-specific enolase and glial fibrillary acidic protein in human glioma cells. Acta Neuropathol 79: 506–512, 1990
Bocchini V, Casalone R, Collini P, Rebel G, Lo Curto F: Changes in glial fibrillary acidic protein and karyotype during culturing of two cell lines established from human glioblastoma multiforme. Cell Tiss Res 265: 73–81, 1991
Smith TW, Nikulasson S, De Girolami U, De Gennaro LJ: Immunohistochemistry of synapsin I and synaptophysin in human nervous system and neuroendocrine tumors. Applications in diagnostic neuro-oncology. Clin Neuropathol 12: 335–342, 1993
Ekstrand AJ, Longo N, Hamid ML, Olson JJ, Liu L, Collins VP, James CD: Functional characterization of an EGF receptor with a truncated extracellular domain expressed in glioblastomas with EGFR gene amplification. Oncogene 9: 2313–2320, 1994
Frederick L, Wang XY, Eley G, James CD: Diversity and frequency of epidermal growth factor receptor mutations in human glioblastomas. Cancer Res 60: 1383–1387, 2000
Mollenhauer J, Wiemann S, Scheurlen W, Korn B, Hayashi Y, Wilgenbus KK, von Deimling A, Poustka A: DMBT1, a new member of the SRCR superfamily, on chromosome 10q25.3-26.1 is deleted in malignant brain tumours. Nat Genet 17: 32–39, 1997
Steck PA, Pershouse MA, Jasser SA, Yung WK, Lin H, Ligon AH, Langford LA, Baumgard ML, Hattier T, Davis T, Frye C, Hu R, Swedlund B, Teng DH, Tavtigian SV: Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced Cancer. Nat Genet 15: 356–362, 1997
Nakamura H, Yoshida M, Tsuiki H, Ito K, Ueno M, Nakao M, Oka K, Tada M, Kochi M, Kuratsu J, Ushio Y, Saya H: Identification of a human homolog of the Drosophila neuralized gene within the 10q25.1 malignant astrocytoma deletion region. Oncogene 16: 1009–1019, 1998
Wechsler DS, Shelly CA, Petroff CA, Dang CV: MXI1, a putative tumor suppressor gene, suppresses growth of human glioblastoma cells. Cancer Res 57: 4905–4912, 1997
Zhou XP, Li YJ, Hoang-Xuan K, Laurent-Puig P, Mokhtari K, Longy M, Sanson M, Delattre JY, Thomas G, Hamelin R: Mutational analysis of the PTEN gene in gliomas: molecular and pathological correlations. Int J Cancer 84: 150–154, 1999
James CD, Galanis E, Frederick L, Kimmel DW, Cunningham JM, Atherton-Skaff PJ, O'Fallon JR, Jenkins RB, Buckner JC, Hunter SB, Olson JJ, Scheithauer BW: Tumor suppressor gene alterations in malignant gliomas: histopathological associations and prognostic evaluation. Int J Oncol 15: 547–553, 1999
Henson JW, Schnitker BL, Correa KM, von Deimling A, Fassbender F, Xu HJ, Benedict WF, Yandell DW, Louis DN: The retinoblastoma gene is involved in malignant progression of astrocytomas. Ann Neurol 36: 714–721, 1994
Bello MJ, de Campos JM, Kusak ME, Vaquero J, Sarasa JL, Pestana A, Rey JA: Molecular analysis of genomic abnormalities in human gliomas. Cancer Genet Cytogenet 73: 122–129, 1994
Hannon GJ, Beach D: p15(INK4B) is a potential effector of TGF-beta-induced cell cycle arrest. Nature 371: 257–261, 1994
Kamb A, Gruis NA, Weaver-Feldhaus J, Liu Q, Harshman K, Tavtigian SV, Stockert E, Day RSI, Johnson BE, Skolnick MH: A cell cycle regulator potentially involved in genesis of many tumor types. Science 264: 436–440, 1994
Sallinen SL, Sallinen PK, Kononen JT, Syrjakoski KM, Nupponen NN, Rantala IS, Helen PT, Helin HJ, Haapasalo HK: Cyclin D1 expression in astrocytomas is associated with cell proliferation activity and patient prognosis. J Pathol 188: 289–293, 1999
Bigner SH, Mark J, Bigner DD: Chromosomal composition of four permanent cultured cell lines derived from human gliomas. Cancer Genet Cytogenet 10: 335–349, 1983
Patel A, van Meyel DJ, Mohapatra G, Bollen A, Wrensch M, Cairncross JG, Feuerstein BG: Gliomas in families: chromosomal analysis by comparative genomic hybridization. Cancer Genet Cytogenet 100: 77–83, 1998
Jung V, Romeike BF, Henn W, Feiden W, Moringlane JR, Zang KD, Urbschat S: Evidence of focal genetic microheterogeneity in glioblastoma multiforme by area-specific CGH on microdissected tumor cells. J Neuropathol Exp Neurol 58: 993–999, 1999
Heim S, Mitelmann F: Cancer Cytogenetics 2nd edn., Wiley-Liss., New York, 1995, pp 141–165
Buijs A, Sherr S, van Baal S, van Bezouw S, van der Plas D, Geurts van Kessel A, Riegman P, Lekanne Deprez R, Zwarthoff E, Hagemeijer A, Grosveld G: Translocation (12;22) (p13;q11) in myeloproliferative disorders results in fusion of the ETS-like TEL gene on 12p13 to the MN1 gene on 22q11. Oncogene 10: 1511–1519, 1995
Wlodarska I, Selleri L, La Starza R, Paternotte C, Evans GA, Boogaerts M, Van den Berghe H, Mecucci C: Molecular cytogenetics localizes two new breakpoints on 11q23.3 and 21q11.2 in myelodysplastic syndrome with t(11;21) translocation. Gen Chrom Canc 24: 199–206, 1999
Golub TR, Barker GF, Bohlander SK, Hiebert SW, Ward DC, Bray-Ward P, Morgan E, Raimondi SC, Rowley JD, Gilliland DG: Fusion of theTELgene on 12p13 to theAML1 gene on 21q22 in acute lymphoblastic leukemia. Proc Natl Acad Sci USA 92: 4917–4921, 1995
Sehgal A: Molecular changes during the genesis of human gliomas. Semin Surg Oncol 14: 3–12, 1998
Iijima Y, Ito T, Oikawa T, Eguchi M, Eguchi-Ishimae M, Kamada N, Kishi K, Asano S, Sakaki Y, Sato Y: A new ETV6/TEL partner gene, ARG (ABL-related gene or ABL2), identified in an AML-M3 cell line with a t(1;12)(q25;p13) translocation. Blood 95: 2126–2131, 2000
Hecht BK, Turc-Carel C, Chatel M, Grellier P, Gioanni J, Attias R, Gaudray P, Hecht F: Cytogenetics of malignant gliomas: I. The autosomes with reference to rearrangements. Canc Genet Cytogenet 84: 1–8, 1995
Rey JA, Bello MJ, de Campos JM, Kusak ME, Moreno S: Cytogenetic follow-up from direct preparation to advanced in vitro passages of a human malignant glioma. Canc Genet Cytogenet 41: 175–183, 1989
Bigner SH, Mark J, Bigner DD: Chromosomal progression of malignant human gliomas from biopsy to establishment as permanent lines in vitro. Canc Genet Cytogenet 24: 163–176, 1987
Harada K, Nishizaki T, Ozaki S, Kubota H, Ito H, Sasaki K: Intratumoral cytogenetic heterogeneity detected by comparative genomic hybridization and laser scanning cytometry in human gliomas. Canc Res 58: 4694–4700, 1998
Macville M, Schrock E, Padilla-Nash H, Keck C, Ghadimi BM, Zimonjic D, Popescu N, Ried T: Comprehensive and definitive molecular cytogenetic characterization of HeLa cells by spectral karyotyping. Canc Res 59: 141–150, 1999
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Krex, D., Mohr, B., Hauses, M. et al. Identification of Uncommon Chromosomal Aberrations in the Neuroglioma Cell Line H4 by Spectral Karyotyping. J Neurooncol 52, 119–128 (2001). https://doi.org/10.1023/A:1010680920087
Issue Date:
DOI: https://doi.org/10.1023/A:1010680920087