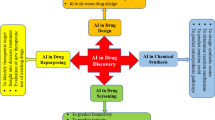
Abstract
Computer-aided drug design (CADD) has emerged as a highly effective and indispensable tool for streamlining the drug discovery process, leading to significant reductions in cost and time. The integration of CADD with machine learning (ML) and deep learning (DL) technologies further enhances its potential and promises novel advancements in the field. In this article, we provide a review of the computational methods employed in the development of novel anesthetics, outlining their respective advantages and limitations. These techniques have demonstrated their utility across various stages of drug discovery, encompassing the exploration of target-ligand interactions, identification and validation of new binding sites, de novo drug design, evaluation and optimization of absorption, distribution, metabolism, excretion, and toxicity (ADMET) properties in lead compounds, as well as prediction of adverse effects. Through an in-depth exploration of computational approaches and their applications, this article aims to help relevant researchers develop safer and more effective anesthetic drugs.
Similar content being viewed by others
1 Introduction
One of the most remarkable milestones in the history of medicine is attributed to William T.G. Morton (1819–1868), who pioneered the use of inhaled ether as a surgical anesthetic [1]. Anesthesia is a temporary and reversible state induced by medications or other interventions that leads to a controlled loss of sensation or consciousness [2]. Depending on the nature of the procedure and individual patient requirements, various types and levels of anesthesia are available, including general anesthesia, regional anesthesia, and local anesthesia. The primary objective of anesthesia is to ensure patient safety and comfort, providing a foundation for further examinations or surgical procedures.
The effects of anesthesia are intricately intertwined with a wide array of physiological processes, encompassing analgesia, sedation, hypnosis, immobility, unconsciousness, amnesia, suppression of autonomic reflexes, and muscle relaxation [3, 4]. In clinical practice, anesthesia involves the administration of a combination of anesthetic drugs rather than relying solely on a single agent. Broadly speaking, anesthetic drugs can be categorized into various types, including general anesthetics, local anesthetics, analgesics, sedatives, and muscle relaxants [5].
The pursuit of an ideal anesthetic encompasses a wide range of characteristics including safety, effectiveness, controllability, rapid onset and offset, minimal side effects, stability, compatibility, reversibility, and cost-effectiveness, yet the realization of such an agent remains elusive [4, 6, 7]. Although commonly used anesthetic drugs possess certain desirable properties, they are also associated with unwanted effects (Table 1). Improper utilization may lead to adverse reactions including respiratory depression, hypotension, adrenal suppression, inflammation, nausea, vomiting, and convulsions in patients [8,9,10,11,12]. Consequently, the expectation among all anesthesia practitioners is the development of a novel, safer, and more efficacious anesthetic drug.
The development of novel drugs is a complex, protracted, high-risk, and costly endeavor, typically spanning more than a decade and involving investments amounting to billions of dollars [13]. Despite the extensive research and financial commitments, only a limited number of drugs successfully navigate the approval process and reach the market [14, 15]. Notably, drugs targeting the central nervous system, including anesthetics, experience the highest rate of failure in development [16]. Numerous factors contribute to the failure of drug development, including but not limited to inadequate potency, unforeseen adverse effects, off-target effects, and significant challenges in synthesis [17].
The advancement of computer-aided methods and artificial intelligence in the medical field has propelled modern anesthesiology into the era of information technology. In this regard, the integration of computer-aided drug design (CADD) and machine learning techniques has yielded diverse applications in the field of drug discovery [18]. These methodologies have proven instrumental in reducing costs, saving time, preemptively eliminating unqualified molecules, and minimizing failures in the final stage of drug development. This paper provides a comprehensive review of commonly utilized computer-aided and machine-learning methods, highlighting their applications in the development of anesthetic drugs.
2 Methods of computer-aided drug design
The concept of CADD can be traced back to the early 1960s when researchers started using computers to simulate and model chemical interactions. However, it was not until the 1980s that the term “computer-aided drug design” was coined, and the field really began to take off in the 1990s [33]. Generally, there are two types of techniques: structure-based drug design and ligand-based drug design (Fig. 1).
2.1 Structure-based drug design
Structure-based drug design (SBDD) has gained significant popularity as a widely embraced strategy for expedited and cost-effective lead identification and optimization. SBDD has emerged as a widely adopted approach for rapid and cheaper lead identification and optimization [34]. It encompasses a range of techniques, such as homology modeling, molecular dynamics, molecular docking, and de novo drug design [35]. The core of SBDD is to explore binding modes in protein-ligand complexes and predict the binding affinity [36, 37]. The initial step in SBDD is to identify drug targets and obtain their three-dimensional structure, which can be experimentally measured or computationally predicted using methods like homology modeling [13].
2.1.1 Molecular docking
Once the 3D structure of proteins is obtained, the subsequent step involves exploring atomic-level interactions between macromolecular structures and small molecules [38]. The most well-known SBDD technique in this process is molecular docking. Docking methods typically consist of two components: the search algorithm and the scoring scheme [39, 40]. The search algorithm is responsible for sampling suitable conformations from high-dimensional spaces, while the scoring scheme is employed to evaluate interaction energies and rank different candidate dockings [41, 42]. Classic scoring schemes can be force-field-based, empirical, or knowledge-based [43], while the introduction of machine learning (ML) approaches has greatly improved the accuracy of scoring functions [44]. Recent developments in molecular dynamic simulations have made conformational exploration easier, thereby making molecular docking a more powerful tool for virtual screening lead compounds.
2.1.2 Homology modeling
Owing to the high cost and time-intensive feature of the experimental measurements, homology modeling has become one of the most precise computational methods to obtain 3D coordinates of proteins from their amino acid sequences [45, 46]. The fundamental tenet of homology modeling is that the target sequence of interest is likely to share a similar structure and function to a homologous template [47, 48]. The quality of the predicted structure is determined by the level of similarity between the target and the template sequence [49]. Sequence-based similarity search algorithms, such as FASTA and BLAST, are commonly employed to identify homologous templates from the Protein Data Bank (PDB) dataset [50].
2.1.3 Molecular dynamics
Proteins were once believed to be static structures, but this view has been challenged and overturned [51]. Molecular dynamics (MD) is a computer simulation technique employed to predict protein movements and conformational changes over time [52]. Initially introduced by Alder and Wainwright [53], MD simulations have evolved from methods based on Newtonian mechanics to more advanced quantum mechanical approaches [54, 55]. MD simulations are widely used to study drug-target interactions, capture multiple conformational changes in proteins, calculate ligand-target binding energies, and identify cryptic or allosteric sites [56, 57]. With rapid development and iteration of graphics processing units, the precision of MD simulations has significantly improved, thereby enabling the exploration of more sophisticated biochemical processes [58].
2.1.4 High-throughput screening and virtual screening
Both high-throughput screening (HTS) and virtual screening (VS) share the goal of identifying potential target molecules from a large, diverse library of compounds [59]. VS primarily relies on computational simulations to predict the interactions between small molecules and a target [60]. On the other hand, HTS involves the experimental testing of a large number of compounds in the wet lab. Though these two approaches possess certain distinctions, they are often used complementarily and researchers even use these terms interchangeably. Depending on the availability of structural data, VS/HTS can be conducted using either receptor-based or ligand-based methods [61]. However, in the practical application of anesthetic drug development, the utilization of structure-based methods is more prevalent.
2.2 Ligand–based drug design
In cases where structural information about drug targets is not available, ligand-based drug design (LBDD) can be a preferred approach [62]. The principle behind LBDD is that molecules with similar structures tend to share similar biological properties. Common LBDD methods include quantitative structure-affinity relationship (QSAR), similarity search, and pharmacophore modeling [63]. The main objective of LBDD techniques is to identify key structural or physicochemical features that account for the observed biological activity of a set of compounds [64].
2.2.1 Similarity search
Similarity searching is the most effective and straightforward approach to selecting structures similar to the input compound based on chemical or physiochemical characteristics [65, 66]. Choosing the right molecular descriptor is an important part of similarity measuring. Common molecular descriptors include 2D and 3D descriptors as well as various physicochemical properties [67]. The most widely used 2D descriptor is the molecular fingerprint, which compares similarity through bit strings [68]. Similarity searching is used in a wide range of scenarios, such as molecular novelty measurement, clustering analysis, docking searches, and reaction similarity evaluation [69]. This method can be a more advantageous alternative when lacking enough ligands with known bioactivity [70].
2.2.2 Pharmacophore modeling
Pharmacophores represent abstract characteristics essential for biological activity, rather than actual chemical or functional groups [71]. Pharmacophore features commonly include cations, anions, aromatics, and hydrogen bond acceptors or donors [72, 73]. Ligand-based pharmacophore modeling is commonly utilized in virtual screening to reveal features crucial for receptor binding. It generally includes the following steps: analyzing the conformational space of ligands in the training set, aligning molecular conformations, and identifying the best overlay of pharmacophore features [74].
2.2.3 QSAR
QSAR modeling is a process of predicting the physical and biological properties of untested molecules based on their structure [75, 76]. Pharmacophore modeling identifies essential 3D features for biological activity, while QSAR quantifies the relationship between structural features and activity, offering complementary information for drug development. QSAR modeling has undergone constant evolution and reinforcement, becoming one of the most frequently utilized approaches for the statistical analysis of biological data [77]. However, the phenomenon of “activity cliffs”, where slight changes in chemical structures can result in significant changes in target activity, can impede QSAR model performance [78]. It’s an inherent challenge but may be improved with other advanced computational techniques.
The selection of chemical descriptors plays a pivotal role in improving the performance of the QSAR models. 2D QSAR studies are based on various 2D properties such as constitutional descriptors, topological descriptors, and electronic descriptors. But their limited ability to account for the 3D structure of molecules undermines their predictive capabilities [79]. To overcome this limitation, 3D-QSAR leverages the ligands’ three-dimensional characteristics like molecular shape, electrostatic potentials, hydrophobic regions, and steric interactions, while 4D-QSAR adds conformational and alignment freedom, improving the model’s accuracy and comprehensibility [80].
2.2.4 Clinical demand-oriented reverse drug design
Anesthesia, especially general anesthesia is an intricate physiological activity that encompasses multiple targets, and a comprehensive understanding of each target’s mechanism remains elusive. These hurdles make it more complicated to develop novel anesthetics through the target-based drug design paradigm. In addition, clinicians consider the clinical features of a molecule as the most crucial information and the primary indicator of its potential as a new drug. Although descriptors commonly used in QSAR pertain to the molecule’s druggability, they do not provide a direct reflection of the molecule’s biological activity when used in patients.
Therefore, our team led by Jin Liu (the professor and chairman of the Anesthesiology Department and director of the Translational Neuroscience Center at West China Hospital, Sichuan University, Chengdu, China) presents a novel strategy termed “clinical demand-oriented reverse drug design”. Our strategy prioritizes the direct optimization of compound clinical properties regardless of the information of targets, distinguishing itself from the conventional target-based drug discovery paradigm. Moreover, our model has the ability to understand clinical requirements described in natural language. It can process a descriptive paragraph outlining clinical demands as input and subsequently generate potential candidates, setting it apart from phenotype-based drug discovery, which primarily revolves around screening as its central methodology.
Our team has undertaken efforts to develop novel general anesthetics through clinical demand-oriented reverse drug design. We have selected several clinical characteristics as optimization goals, encompassing anesthesia efficacy, anesthetic onset and duration time, blood pressure, abnormal nervous system excitation, and respiratory depression. Taking inspiration from Jin’s work [81], our initial step involved training a molecular variational autoencoder (VAE) capable of encoding and decoding molecules. The VAE model enabled us to encode input molecule graphs into latent vectors and decode modified vectors into compounds with optimized properties. In addition, the latent space provided by the VAE model can be utilized to train predictors for diverse clinical properties.
To generate and optimize potential compounds, we randomly sampled vectors from the latent space (i.e., random initialization). Subsequently, we calculated the necessary gradients for each property and applied the gradient descent algorithm to optimize each property individually. The optimized compounds exhibiting high predictive scores for all the clinical goals underwent further experimentation in the wet lab. Our research paradigm has already yielded preliminary results and may shed light on the development of new anesthetic drugs.
3 Principles of machine learning
The ultimate goal of artificial intelligence (AI) is to create computer systems or machines that have human-like intelligence and independence [82]. Learning and problem-resolving are typical tasks that represent human intelligence, thereby making "machine learning" a core area of artificial intelligence [83]. The major focus of machine learning is to develop computer systems that can acquire knowledge, make predictions or decisions, and enhance their performance using provided data instead of explicit programming [84, 85].
ML techniques are typically classified into three main categories (Fig. 2): supervised learning, unsupervised learning, and reinforcement learning. In addition, deep learning, a specialized approach within ML, and transfer learning, an extension or application of ML, have also demonstrated their utility in the field of drug discovery.
3.1 Supervised learning
Supervised learning encompasses the process of training a model using annotated data, where input samples are paired with corresponding target labels [86]. Through this approach, the model acquires the ability to make predictions or classify unseen data by leveraging the patterns and relationships learned from the labeled examples.
3.1.1 Classification
Classification refers to assigning input data instances to specific predefined categories or classes. The evaluation of a classification model is typically based on metrics such as precision, recall, accuracy, F1 score, and the area under the receiver operator characteristic curve (AUC-ROC) [87].
Naive Bayes (NB)
Naive Bayes is a classification method based on Bayes theorem that assumes the independence of features given class labels. This method has advantages including simplicity, computational efficiency, and ability to handle high-dimensional or unbalanced data [88]. Nonetheless, in real-world scenarios, it is uncommon for features to be truly independent, which restricts the effectiveness of this method when substantial interactions exist between features.
k-Nearest Neighbor (k-NN)
The k-NN algorithm classifies or predicts new data points based on their proximity to labeled training examples within a feature space. k-NN exhibits versatility by accommodating both classification and regression tasks. To generate predictions, the algorithm identifies the k nearest neighbors in proximity to the new data point and assigns the most prevalent class label among them. The algorithm’s performance hinges upon the careful selection of the distance metric and the optimal value for k [89]. Noteworthy advantages of k-NN encompass its simplicity, ease of implementation, capacity to handle multi-class classification, and adaptability to novel data. Nevertheless, k-NN’s main disadvantage is computational complexity, especially with large datasets, owing to the necessity of comparing the query point against all training examples [90].
Support Vector Machine (SVM)
SVM aims to identify an optimal hyperplane within a high-dimensional space to separate data points into different classes, maximizing the margin between the classes. SVM is widely regarded as one of the most robust predictive models and finds extensive utilization in the domain of drug development [91]. SVMs are capable of handling high-dimensional data but are not directly applicable to multi-class tasks.
Decision Tree (DT)
The decision tree algorithm constructs a hierarchical structure of nodes to facilitate decision-making based on simple decision rules derived from data features [92]. Conceptually, a tree can be perceived as a piecewise constant approximation. This approach possesses a white-box property, offering interpretability through visually comprehensible tree structures, and it can effectively handle both numerical and categorical data. These advantages have established decision trees as one of the most prevalent and potent machine learning methods [93]. The limitations of decision trees include their proneness to overfitting when deep trees are utilized, as well as their sensitivity to minor data variations, which can lead to high variance [94]. To address these limitations, ensemble techniques like random forests or gradient boosting can be utilized.
Gradient Boosting Machine (GBM)
Boosting is a technique that transforms weak learners into strong learners, and one notable implementation of this principle is gradient boosting. GBM sequentially combines weak predictive models to create a robust ensemble model and can be used in regression and classification tasks. When decision trees serve as the weak learners, the resulting algorithm is referred to as a gradient-boosted tree. This iterative process allows GBM to effectively capture complex, non-linear relationships in the data, providing higher accuracy compared to single machine learning models [95].
Random Forest (RF)
Random forest is a versatile machine-learning algorithm that harnesses the collective strength of an ensemble of decision trees. Each tree in the forest is constructed independently by randomly selecting subsets of features and data samples. The final prediction is then determined by aggregating the outputs through majority voting or averaging. RF exhibits several advantages, including its proficiency in handling high-dimensional data, capturing complex relationships, and effectively managing missing values and outliers. Although Random Forest frequently achieves superior accuracy compared to a single decision tree [96], it sacrifices some of the inherent interpretability in decision trees.
Synthetic Minority Oversampling Technique (SMOTE)
Label imbalance is a common challenge encountered in datasets utilized for drug discovery [97], and SMOTE is employed to tackle this issue [98]. It addresses the scarcity of data in the minority class by generating synthetic samples that closely resemble the existing minority class instances. This approach effectively balances the class distribution and enhances the classifier’s ability to accurately predict the minority class. However, SMOTE may introduce synthetic samples that are noisy or less representative of the actual data.
3.1.2 Regression
Regression aims to predict a continuous numerical value as the output. Regression models can be linear or non-linear and are evaluated using metrics such as mean squared error, mean absolute error, or R-squared.
Linear regression
The linear regression model is renowned for its simplicity and effectiveness. It establishes a linear relationship between a dependent variable and one or more independent variables, seeking to identify the best-fit line that minimizes the overall difference between the predicted and actual values [99]. Notably, linear regression offers interpretability through coefficients that convey the direction and magnitude of the variable relationships. But this method is not suitable for nonlinear scenarios.
Ordinary least square
One of the commonly used methods in Linear Regression is Ordinary Least Squares, which estimate the parameters of a linear model by minimizing the sum of the squared errors.
Logistic regression
Logistic regression gets its name from the logistic function, also known as the sigmoid function, which lies at the core of this method. The sigmoid function maps real-valued numbers to a range between 0 and 1. Despite the resemblance to linear regression in terms of representation, logistic regression differs by focusing on binary values rather than continuous ones [100]. The simplicity and interpretability of logistic regression make it advantageous, as it offers insights into the importance and direction of input variables. Furthermore, logistic regression demonstrates computational efficiency and versatility in handling both numerical and categorical features, making it a valuable tool for various types of data analysis.
3.2 Unsupervised learning
Unsupervised learning deals with unlabeled data, allowing the model to discover hidden patterns or data groupings without explicit target labels or human intervention. Clustering and dimensionality reduction are two of the most common tasks in unsupervised learning.
3.2.1 Clustering
Clustering is used to automatically group similar objects together based on their inherent characteristics or patterns, without any prior knowledge of class labels or output values. In the process of drug discovery, clustering can be used to cluster compounds based on structural similarities, or group molecular targets based on functional characteristics.
K-Means
K-Means is an iterative algorithm that partitions data into k clusters by assigning data points to the cluster with the closest centroid and updating the centroids based on the mean of the data points within each cluster. The value of k, representing the number of clusters, is a user-defined parameter. K-Means boasts simplicity and efficiency, making it well-suited for large datasets. It operates without the need for labeled data and can handle both numerical and categorical features. However, it is crucial to consider that K-Means algorithm is sensitive to the initial placement of centroids, which can lead to convergence on different solutions [101].
Hierarchical clustering
Hierarchical Clustering is a method that groups similar data points into nested clusters based on pairwise distances [102]. It differs from K-Means in that it constructs a hierarchical structure of clusters, while K-Means directly partitions the data. Divisive clustering begins with all data points in a single cluster and recursively splits them into smaller clusters until the desired number of clusters is reached. This approach allows for capturing the inherent hierarchical structure of the data without requiring the prior specification of the number of clusters. But the choice of distance metric and linkage method can impact the resulting clusters.
3.2.2 Dimensionality reduction
Dimensionality reduction techniques are used to transform high-dimensional data into a lower-dimensional subspace while preserving the most important features. By reducing the dimensionality, researchers can simplify data analysis, remove noise, and identify key patterns.
Principle Component Analysis (PCA)
PCA addresses the complexity of high-dimensional data by extracting smaller features from a large set of variables [103]. It’s widely acknowledged as one of the most prominent methods within this category. PCA identifies the directions, known as principal components, along which the data exhibits the most significant variability. PCA finds applications in various domains, including molecular descriptors analysis, feature space reduction, and high-dimensional data visualization. However, reducing the number of variables naturally comes at the expense of accuracy.
Linear Discriminant Analysis (LDA)
LDA focuses on projecting data onto a lower-dimensional space while preserving class-specific information. It assumes that the data follow a Gaussian distribution and that the covariance matrices of different classes are equal [104]. LDA calculates class-specific means and covariance matrices to determine a discriminant function that optimally separates the classes. This method offers benefits such as simplicity, interpretability, and the ability to handle multi-class problems. But it can’t capture complex non-linear relationships in the data.
Time-structure Independent Components Analysis (tICA)
tICA is a method used to analyze data that changes over time, like molecular dynamics simulations or time series data. It simplifies the data and finds important patterns in the information. tICA can be a useful tool for understanding slow movements and significant changes in complex systems, thereby accelerating the process of drug development.
3.3 Deep learning
Deep learning (DL) algorithms are based on artificial neural networks, which aim to emulate the functionality of the human brain and learn from examples [105]. In the human brain, millions of interconnected neurons collaborate to acquire and process information [106]. Likewise, deep learning neural networks comprise multiple layers of artificial neurons, operating collectively within a computer system.
3.3.1 Multilayer Perceptron (MLP)
MLP is an artificial neural network that comprises multiple interconnected layers of nodes called neurons. It consists of three types of layers: the input layer, the output layer, and the hidden layer. Each neuron in the network receives input from the previous layer, applies a non-linear activation function to generate an output, and passes it to the next layer. The hidden layers, positioned between the input and output layers, enable MLPs to approximate any arbitrary function and learn complex features [107]. MLPs are trained using the backpropagation technique, where the network adjusts the weights and biases of its connections to minimize the discrepancy between predicted and actual values [108].
3.3.2 Convolutional Neural Network (CNN)
CNN is a neural network architecture that draws inspiration from the structure and functioning of the human visual system [109]. It consists of multiple layers, including convolutional layers, pooling layers, and fully connected layers. The convolutional layers play a central role in CNNs as they extract important features from the input [110]. Pooling layers then downsample the output, reducing its dimensionality. Fully connected layers connect all the neurons from the previous layer to the next, enabling classification or regression tasks. The hierarchical structure of CNNs allows them to automatically learn pertinent features from raw data, eliminating the necessity for manual feature engineering.
Though originally designed for visual data analysis, CNNs have expanded their applications to various domains, including drug discovery. CNNs are now utilized in analyzing intricate molecular and biological data, facilitating de novo drug design, aiding decision-making in pharmaceutical research, and processing imaging data like high-content screening images or histopathology slides [111].
3.3.3 Graph Neural Network (GNN)
A graph is a data structure consisting of two fundamental components: nodes (or vertices) and edges [112]. GNN is a specific type of deep learning model specifically designed to operate on graph-structured data. The primary goal of GNNs is to acquire node or entire graph representations through the propagation of information within the graph structure. An exemplary technique in this realm is the Graph Convolutional Network, which extends conventional convolutional operations to encompass graphs.
Graphs provide an effective representation of the structure and properties of chemical molecules, and they have garnered significant attention in the field of drug discovery [113, 114]. GNNs offer the capability to model molecular structures and exploit both local and global molecular interactions, making them valuable tools in the exploration of novel drug candidates.
3.3.4 Recurrent Neural Network (RNN)
RNN is a general framework for processing sequential data. Unlike feedforward neural networks, RNNs have feedback connections, allowing them to maintain an internal memory or state. This memory allows the network to process information not only based on the current input but also on previously seen inputs in the sequence.
RNNs suffer from vanishing or exploding gradients, which can make long-term dependencies challenging to learn. To mitigate this issue, specialized variations of RNNs, such as Long Short-Term Memory (LSTM) and Gated Recurrent Unit (GRU), have been introduced. LSTMs achieve this through separate memory cells and gating mechanisms [115], while GRUs simplify the architecture by using combined and reset gates [116].
3.4 Transfer learning
Transfer learning is a machine learning method that applies knowledge obtained from training one task to a distinct but related task [117]. Instead of commencing from scratch, a pre-existing model, typically trained on a substantial dataset, is used as a foundation for a new task [118]. The pre-trained model has already learned general features and patterns from the initial task, which can be leveraged to improve learning and performance on the new task even with limited labeled datasets.
The integration of transfer learning and deep learning architectures frequently gives rise to a powerful framework known as deep transfer learning, offering a convenient means of fine-tuning parameters. Transfer learning has demonstrated promising applications, including biological sequence analysis, molecule bioactivity prediction, molecular generation, virtual screening, and protein-protein interaction prediction [119]. These advancements highlight the potential of transfer learning to enhance various aspects of drug discovery research.
This promising method can save computational resources and minimize the requirement for labeled data. However, effective transfer learning depends on the similarity between the pre-trained task and the target task. Failing to select an appropriate pre-trained model may result in the "negative transfer", wherein the learner’s performance deteriorates compared to not employing transfer learning at all [120].
3.5 Reinforcement learning
Reinforcement learning (RL) stands as a distinct paradigm within the field of machine learning, wherein an agent interacts with an environment, receives feedback or rewards, and adjusts its behavior through trial and error [121]. Although both supervised learning and reinforcement learning aim to optimize their performance over time, they differ in three main aspects: Firstly, supervised learning relies on labeled training data, while reinforcement learning doesn’t require pre-labeled datasets. Secondly, supervised learning aims to predict output accurately for new inputs, whereas reinforcement learning seeks an optimal strategy to maximize cumulative rewards over time. Thirdly, feedback in supervised learning comes from labeled training data, while reinforcement learning receives feedback as rewards or penalties based on agent actions in the environment, which can be delayed and sparse. Similar to how children explore their surroundings and learn to achieve a goal, RL models operate autonomously and self-teach. RL algorithms hold considerable power and promise as they can learn optimal actions for achieving success in an unfamiliar environment without the need for intervention or guidance from a supervisor.
Reinforcement learning has found practical applications in the realm of drug discovery, encompassing a range of areas such as enhancing precision medicine, improving clinical trials, optimizing biochemical properties, and advancing pharmacological research [122, 123]. Traditional generative models may generate novel molecules with limited synthetic accessibility. This challenge can be effectively tackled by integrating reinforcement learning algorithms to explore the synthesizable chemical space [124].
RL boasts numerous advantages and diverse applications, but it is not exempt from limitations. For example, RL algorithms grapple with the delicate task of striking a balance between exploring new actions and exploiting existing knowledge. Additionally, designing appropriate reward functions poses a significant challenge [125].
4 Applications in anesthetics discovery
Drug discovery typically involves identifying and validating targets, discovering potential hits and leads, optimizing lead compounds, conducting preclinical testing, progressing through clinical trials, and obtaining regulatory approval (Fig. 3). CADD and ML methods have been widely applied at various steps of the process (Table 2).
4.1 Target elucidation
Target elucidation is the fundamental basis for the design and development of effective drugs that modulate target activity and deliver disease treatment. It encompasses two primary parts: target identification and target validation. Target identification involves the identification of biological targets (proteins, enzymes, receptors, etc.) and the exploration of their structural characteristics [169]. Target validation, on the other hand, aims to understand target-ligand interactions, uncover the underlying molecular mechanisms, and demonstrate the druggable potential of the target [170]. Obtaining comprehensive information about targets is vital for guiding the subsequent drug discovery process.
4.1.1 Prediction of protein structures
Proteins in living cells continuously undergo conformational changes, altering their atomistic-level structures and binding affinities with chemicals [154]. Researchers have found that certain protein conformations exhibit significantly stronger chemical binding abilities than others [171]. Furthermore, a protein’s shape determines its function. Erroneous protein structures may lead to toxicity and adverse reactions. Therefore, correctly predicting the structure of proteins and understanding protein conformational dynamics are vital to uncovering the underlying mechanisms of diseases and then advancing the drug discovery process.
Traditional structural biology experiments are time- and money-consuming, thus computational methods have recently flourished to produce less expensive and quicker computer simulations, helping researchers to predict the structure of targets related to anesthesia [172]. For example, Gamma-aminobutyric acid (GABA) has been widely recognized as the primary inhibitory neurotransmitter in the central nervous system [173]. The characterization of \(\textrm{GABA}_{\textrm{A}}\) and \(\textrm{GABA}_{\textrm{B}}\) receptors has been crucial for understanding some physiological activities in the nervous system [174]. Sripriya Akondi et al. applied SVM to classify \(\textrm{GABA}_{\textrm{A}}\textrm{Rs}\) based on the features extracted from Chou’s pseudo-amino acid composition [158, 175]. Liao et al. trained four ML classifiers to distinguish \(\textrm{GABA}_{\textrm{A}}\textrm{Rs}\) from non-\(\textrm{GABA}_{\textrm{A}}\textrm{Rs}\), using only the protein sequence information. Among the four classifiers, the gradient boosting decision tree and a library for support vector machine (libSVM) marginally overperformed RF and k-NN [157].
Opioids are widely used for pain relief and there are four primary subtypes of opioid receptors. Opioid receptor Kappa 1 (OPRK1) agonists have been demonstrated to activate pain-inhibitory pathways within the central nervous system [176]. Recently, Sripriya Akondi et al. implemented a decision fusion strategy that integrated multiple machine-learning algorithms to identify potential drug-binding molecular conformations of the OPRK1 protein. To address class imbalance issues, a synthetic minority oversampling technique (SMOTE) was incorporated. The decision fusion strategy utilizing LR (Logistic regression) + SMOTE + GB (Gaussian Naive Bayes) and LR + SMOTE + k-NN techniques outperformed those using a single technique [154]. Feinberg et al. employed a combination of molecular dynamics simulations and unsupervised machine learning techniques, including the tICA algorithm and K-Means clustering, to identify new conformational states of \(\mu\)OR. The authors then utilized docking scores to develop random forest classifier models that were capable of virtual screening agonists and binders for \(\mu\)OR [134].
4.1.2 Prediction of targets and binding sites
Effective target discovery is an indispensable prerequisite in the modern target-based drug discovery paradigm, as the selection of appropriate targets directly correlates with the success of drug development [177,178,179]. Therefore, prioritizing the identification and validation of targets is of paramount importance in the drug discovery process.
For example, Kang et al. created a novel deep-learning model and discovered a distinct druggable binding site for the P2X3 receptor (P2X3R). Their finding of 16 unique hit compounds may expedite the search for new P2X3R antagonists and lead to more effective neuropathic pain treatments [166].
Bertaccini et al. conducted a series of studies utilizing homology modeling to construct atomic-level models of the GABA receptor and the glycine alpha one receptor (GlyRa1). These models uncovered potential anesthetic binding sites and suggested a link between ligand docking scores and drug potency. Their findings have made a significant contribution to the current understanding of anesthetic action and have facilitated the development of novel anesthetics [128,129,130].
4.1.3 Prediction of protein-ligand interactions
Protein-ligand interactions take an irreplaceable part in various biological processes, including signal transduction, gene regulation, and immunologic reaction [180]. Therefore, exploring the mechanisms of protein-ligand recognition and binding is crucial for providing a theoretical foundation for drug development [181, 182].
In a recent study, Cheng and Ding developed SVR-based QSAR models to estimate the compounds’ binding affinities after docking with the \(\textrm{GABA}_{\textrm{A}}\) receptor and identified six essential characteristics that influence ligand-receptor binding [147]. Wijeyesakere et al. successfully built in silico tools to access the GABAergic potential of uncharacterized compounds — two binary classifiers based on random forest [164]. Liu et al. showed the effectiveness of molecular docking methods in predicting both the binding sites and affinities of anesthetics for both water-soluble and membrane proteins [137]. By employing molecular docking and absorption, distribution, metabolism, excretion, and toxicity (ADMET) prediction techniques, Meraj et al. assessed a group of Disopyramide analogs and discovered five potential lead compounds that were capable of binding to human voltage-gated sodium channel (NaV) proteins [135], which is beneficial for the development of anesthetics, anticonvulsants, and antiarrhythmic drugs [183]. Similarly, Lv et al. discovered the potential of a traditional Chinese medicine named gastrodin as a new anesthetic drug[136].
A major goal of computational biology has been to explore the mechanisms of protein-ligand binding. Based on homology modeling, Elgarf et al. discovered that different benzodiazepine ligands employ discrepant modes for binding to \(\textrm{GABA}_{\textrm{A}}\) receptors [126]. Pan et al. integrated several computational methods, including homology modeling, molecular docking, and molecular dynamics simulations, to investigate the interaction between benzhydrylpiperazine agonists and human \(\delta\) opioid receptors [127]. Yuan et al. combined molecular dynamics simulations and molecular docking to explore the potential binding mechanism of propofol to \(\textrm{GABA}_{\textrm{A}}\textrm{Rs}\) [131]. Lima Neto et al. were the pioneers in introducing quantum biochemical computational approaches to investigate the binding mechanism of existing agonists and antagonists to the \(\textrm{GABA}_{\textrm{B}}\) receptor [139]. A recent study by Manzur-Villalobos et al. evaluated the association between the Nav1.7 channel and local anesthetics through virtual screening, molecular docking and dynamics [133].
4.2 Hit and lead discovery
In drug discovery, the term "hit and lead discovery" encompasses the initial stages involved in identifying and optimizing potential drug candidates. This umbrella term includes two phases: hit discovery and hit-to-lead. The hit discovery phase focuses on the identification of compounds or molecules that exhibit promising activity against a particular target or disease [184]. Once hits are identified, the hit-to-lead phase begins, where the selected hits undergo further optimization to enhance their potency, selectivity, and other desirable properties [185]. These two stages typically involve high-throughput screening or virtual screening of large compound libraries, de novo drug design, and structure-activity relationship studies.
4.2.1 High-throughput screening and virtual screening
Since the mid-1990s, HTS has been widely considered the primary catalyst for hit discovery [186, 187]. This method enables the identification of potential "hits" from large-scale chemical libraries [188]. In recent years, there has been a notable advancement in VS techniques, which can complement high-throughput discovery methods. Both academia and industry have replaced traditional drug screening methods with HTS and VS due to their simplicity, cost-effectiveness, and efficiency [189].
Through HTS and VS, Feinberg et al. identified a promising molecule named FMP4, which is an agonist for \(\mu\)OR and possesses a unique structure [134]. Cayla et al. identified a group of lead compounds based on a distinctive chemical core. These compounds exhibited comparable anesthetic efficacy while minimizing detrimental cardiovascular side effects, making them superior to traditional intravenous anesthetics [140]. Chandra et al. introduced a tiered approach for the identification of Nav1.7 inhibitors. A novel compound named DA-0218 exhibited analgesic and anti-pruritic properties [132]. Ebalunode et al. created a structure-based pharmacophore modeling approach to identify novel anesthetic compounds [146]. Peng et al. discovered prospective anesthetics by virtual screening 50,000 compounds based on drug-likeness and ADMET properties, with five lead compounds demonstrating the potential for binding to \(\textrm{GABA}_{\textrm{A}}\) protein [143]. Yang et al. discovered two innovative compounds with reversible sedative-hypnotic activity [141]. Peng et al. identified two FDA-approved drugs that had not been recognized as sigma 1 receptor (S1R) antagonists previously [142].
Specifically, fluorescence-based techniques are becoming the most commonly used detection method due to their great sensitivity and automation-friendliness[189, 190]. For example, Lea et al. utilized a surrogate binding target for anesthetics and an environment-sensitive fluorescent probe to identify novel anesthetics [144]. McKinstry-Wu et al. developed a high-throughput assay by choosing a fluorescent anesthetic agent and apoferritin as a surrogate for the anesthetic target. In their study, two potent agents with low toxicity were selected from a large-scale library [145].
4.2.2 De novo drug design
Over the past 15 years, de novo drug design—the process of generating innovative compounds with desirable properties but without a beginning template has emerged as a potential area of study [191]. De novo drug design can produce entirely novel and distinctive molecules to enhance existing chemical libraries, whereas traditional virtual screening often screens candidate compounds from existing synthetically accessible, druglike chemical space [192, 193]. Nevertheless, lead compounds generated through this approach typically exhibit low synthetic accessibility [194].
With the emergence and flourishing of deep learning and reinforcement learning techniques, de novo drug design has offered promising opportunities for developing new and effective drugs. For instance, CNNs [195], RNNs [192], generative adversarial networks [196] and VAEs [197] have already been applied in this field.
Given the limited brain retention capacity of the commonly used opioid antagonist Naloxone, the development of improved opioid antagonists is crucial in tackling the opioid epidemic. Deng et al. created a deep reinforcement learning (DRL) framework to generate molecular simplified molecular-inputine-entry system (SMILES), predict chemical properties, and optimize SMILES into desired properties. In this study, they sampled 10k SMILES and filtered out 6 potential lead compounds, demonstrating their DRL framework’s ability to discover better opioid antagonists [165].
4.2.3 QSAR
Using computational methods like QSAR to calculate the ligand-binding affinity is of special interest because it can accelerate the drug discovery process at an early stage [198, 199]. In a study by Lu and Zhou, a novel method was proposed by bridging 3D-QSAR and receptor modeling to predict the binding affinity of imidazobenzodiazepines to \(\textrm{GABA}_{\textrm{A}}\) receptor subtypes [148]. By constructing 3D-QSAR models, Peng et al. accurately predicted the binding affinity between ligands and the S1R [142], Pan et al. evaluated the binding affinity of a set of benzhydrylpiperazine \(\delta\) opioid receptor agonists [127]. Floresta et al. built three distinct QSAR models based on a class of fentanyl-like structures to predict the binding affinity of \(\mu\)OR ligands [153]. Jia et al. presented a fresh workflow to discover new analgesic opioids with higher binding affinities. For each bioassay endpoint taken from the PubChem dataset, they created 12 individual QSAR models using a mix of four different chemical descriptors and three machine learning techniques (k-NN, RF, and SVM) [149]. Mehdipour et al. estimated the anesthetic potency of polyhalogenated ethers by developing a QSAR model that incorporates four parameters: logP, molecular polarizability, most positive charge, and electrostatic potential parameters [150].
4.3 Lead optimization
After identifying lead compounds with the desired bioactivity and selectivity, the subsequential step is lead optimization, which aims to improve their drug-like properties and enhance the chance of developing drug candidates. This involves iterative cycles of compound design, synthesis, and testing to optimize various aspects, including potency, selectivity, pharmacokinetics (PD), pharmacodynamics (PK), metabolic stability, and safety [200].
4.3.1 Evaluation of ADMET properties
PD and PK parameters, including ADMET, have long been critical considerations for researchers in the drug discovery process [201]. Lack of efficacy and safety has been a key factor contributing to the late-stage attrition of drug candidates [202]. The emergence of in silico prediction offers a new dimension for assessing multiple ADMET properties at every stage of drug discovery, enabling the optimization of virtual screening and experimental testing process by filtering out compounds with adverse ADMET profiles [203, 204].
Manzur-Villalobos et al. predicted the ADMET properties of the amide-type local anesthetic analogs and identified a potential brand-new local anesthetic that showed high binding capacity [133]. Jagannathan endeavored to discover novel psychoactive metabolites from Cannabis sativa, the smoke of C. sativa, and other phytocannabinoid matrices, based on the physiochemical descriptors and ADMET properties predicted by four ML techniques (NB, SVM, MLP, and hierarchical clustering) [156]. Coli Louvisse de Abreu et al. evaluated multiple types of toxicity of the epinephrine and norepinephrine degradation products using an ADMET predictor [151].
Different from de novo drug design, there are two typical approaches to designing new drugs based on existing drug molecules. One is to screen new compounds from derivatives of FDA-approved drugs[205]. In a recent study, Azamatov et al. introduced computational AMDET prediction methods to develop novel local anesthetics with lower toxicity. Through the study of a group of 1-aryl tetrahydroisoquinoline alkaloid derivatives, they also proposed explainable correlations between chemical structures and acute toxicity [152]. Another approach is to refine currently available drugs by optimizing PK/PD parameters. Jiang et al. utilized molecular docking and PK prediction techniques to design etomidate analogs that exhibit comparable anesthetic potency while avoiding adrenocortical suppression [138].
A variety of anesthetic and psychotropic drugs exert their effects on the central nervous system (CNS). Predicting the penetration of the blood-brain barrier (BBB) is a significant part of assessing ADMET properties and designing CNS drugs [206, 207]. A study by Yu et al. proposed an innovative hybrid method for identifying potential CNS drug candidates by combining three traditional ML algorithms (DT, RF, SVM) with the DL algorithm (Graph Convolutional Network). The resulting model generated six explainable sub-structural features that enable the rapid assessment of a molecule’s potential to penetrate BBB and become a CNS drug [162].
PK/PD modeling is a widely utilized method for assessing dose-concentration-response correlations, providing valuable insights for the understanding of drug delivery systems and the process of drug discovery [208, 209]. Sevoflurane is one of the most commonly used inhalation anesthetics but may lead to neurotoxicity and cognitive dysfunction [210]. Dhandore et al. combined PK/PD modeling and six machine-learning regression techniques (Linear Regression, Support Vector Regression, Bayesian Ridge, Decision Tree Regressor, Gaussian Process Regressor, and XGBoost) to investigate the effects of sevoflurane concentration changes on ten body parameters. The XGBoost model exhibited the least errors and identified \(\textrm{SpO2}\) as the most crucial body parameter to study drug effects during surgical anesthesia [160].
4.3.2 Prediction of side effects
Unexpected side effects are a major contributor to late-stage drug development failures and can even lead to the withdrawal of FDA-approved drugs [211,212,213]. Traditional approaches for evaluating adverse effects like animal models and clinical research, are time-wasting, costly, and not easily scalable [214,215,216]. Consequently, there is growing interest in using computational methods to detect and predict drug side effects.
Accurately predicting postinduction hypotension (PIH) can greatly assist clinicians in recognizing the potential risk of medication use and selecting the appropriate induction agent. In a study by Kendale et al., eight supervised ML techniques (logistic regression, SVM, NB, k-NN, linear discriminant analysis, random forest, neural nets, and GBM) were employed to assess the risk of hypotension [155]. Lundberg et al. created an ensemble machine learning model that significantly augments anesthesiologists’ predictive capabilities concerning the risk of intraoperative hypoxemia, both in preoperative assessment and real-time monitoring. Their ML system leverages time-series data extracted from the patient’s monitors and anesthesia machine, alongside static information, including age, gender, smoking status, height, and weight, which serve as inputs for the model. Meanwhile, the system enhances interpretability by furnishing insights into the contribution and trends of various risk factors [163].
Abnormal movements like convulsions, seizures, or severe involuntary muscular contractions occasionally happened during anesthesia induction. Nagata et al. developed a convulsion prediction model based on heart rate variability to predict drug-induced convulsions. Their model is built based on an anomaly detection algorithm named multivariate statistical process control [168]. Zhang et al. presented an enhanced SVM model to predict the seizure liability of chemicals, which was characterized by its cost-effectiveness, expeditiousness, and superior accuracy [161]. Gao et al. developed an ML model that combined PCA, SVM and CNNs to effectively differentiate seizure-inducing drugs from safe drugs [159].
To improve clinical decision-making, monitoring and predicting muscle relaxation in patients is important, as muscle relaxants are a critical component of balanced anesthesia. Wang et al. utilized DL techniques (RNN, GRU, LSTM) to predict the real-time Train-of-four ratio of cisatracurium, a commonly used muscle relaxant. Additionally, they implemented transfer learning by calculating patient similarity based on body mass index and age and constructing pre-trained models [167].
5 Challenges
Despite the widespread use of CADD and ML methods in the field of anesthetic drug discovery in recent years, the inherent disadvantages and limitations of these methods, along with the complexity of the anesthetic process, are still inescapable hurdles to the development of new anesthetic drugs.
5.1 Limitations of CADD methods
SBDD heavily relies on the availability of high-resolution structural data, such as protein crystal structures. However, acquiring experimental structures for all target proteins of interest poses significant challenges and overheads. Furthermore, SBDD often overlooks the impact of solvent molecules like water in protein-ligand interactions [217]. Another limitation of SBDD is its tendency to focus on a single target, disregarding the fact that drugs can interact with multiple targets within the human body. Complex diseases and physiological activities can involve multiple targets, thereby restricting the applicability of SBDD in such scenarios.
LBDD requires an ample supply of high-quality ligand data for the target protein. The efficacy of LBDD is greatly influenced by the accessibility and quality of the training set. In addition, LBDD predominantly concentrates on designing drugs based on known ligands that have been previously identified and characterized, but it often neglects the significance of considering protein structure and flexibility [218]. Also, Computational methods used in LBDD such as search algorithms, and scoring functions still have limited accuracy, necessitating further validation in the wet lab.
The limitations specific to each of the commonly used CADD methods are described below. To start with, the accuracy of homology modeling greatly relies on the quality and availability of target-template sequence alignment [219]. Secondly, the sampling methods utilized in molecular dynamics simulations still require further improvement to enhance their accuracy and efficiency [56]. Thirdly, targets and receptors exhibit considerable flexibility, making molecular docking a particularly challenging task [39]. In the case of virtual screening, a major limitation lies in the absence of robust scoring metrics [71]. Moreover, the selection of training compounds significantly influences the quality of the generated pharmacophore model [74]. Eventually, it bears to mention that many CADD methods require substantial computational power and storage resources, which can pose constraints on their widespread application.
5.2 Disadvantages of ML/DL methods
The quality of the dataset is one of the most significant factors affecting the performance of machine learning models [220]. However, data currently available are often not only low in quantity but also far from high-quality, well-annotated, balanced labeling and comprehensive [221, 222]. In particular, there are few specialist datasets available for anesthetic drug development. Secondly, drug discovery datasets incorporate various forms of biological, structural, and chemical information, such as adverse medication effects in patients, multi-omics data in pharmacology, and bioactivity assays in the wet lab [223]. Researchers must carefully select appropriate data handling methods since problems often can’t be treated as simple classification or regression tasks. Lastly, the creation of dedicated databases often requires significant time and effort from experienced professionals for manual data annotation. Due to competition among pharmaceutical companies and high dataset construction costs, high-quality datasets are frequently not publicly available [224].
Lack of transparency and explicability is one of the intrinsic flaws of machine learning that has been widely criticized [223, 225]. "Black box" machine learning models make predictions but without providing explanations of how and why the model reaches a particular conclusion [226, 227]. Nowadays, only a few studies have investigated the explanatory power of their models [222]. This problem greatly hinders the promotion of ML methods in the drug discovery process since researchers are more interested in the biological mechanism behind the prediction.
There are some other important factors closely related to the performance of machine learning models. Firstly, many algorithms are not out-of-the-box and require re-initialization and fine-tuning of parameters, resulting in poor repeatability of results [224]. For example, the starting values like weights and biases in neural networks are usually chosen randomly, which means different initializations can lead to different trained models with varying performance. Additionally, overfitting and underfitting often occur during the training of the model, which significantly impairs predictive accuracy. Finally, training ML/DL models often necessitates extensive storage and computing resources [228]. Large-scale models can’t be trained in a short period of time without a sufficient number of consumer-grade graphical processing units or tensor-processing units.
5.3 Unclear mechanism of anesthesia
Although the notion that general anesthetics act as "drugs without receptors" has been disproved, the precise mechanism of anesthesia remains elusive, making it more challenging to develop novel anesthetic drugs [229, 230].
Recently, there has been growing evidence to suggest that the primary molecular targets for general anesthetics are neurotransmitter- and voltage-gated receptors, such as \(\textrm{GABA}_{\textrm{A}}\) receptors, glycine receptors, nicotinic acetylcholine receptors, and glutamate receptors [231, 232]. However, a drug’s effectiveness is not solely determined by its activity on the intended target, but also by its potential effects on other targets. This is especially relevant in complex therapeutic procedures such as general anesthesia, which often require a combination of drugs to achieve the desired effect on numerous targets [233]. Therefore, the development of novel anesthetics necessitates the consideration of multiple targets, further amplifying the complexity of this task.
Furthermore, the development of effective and safe drugs requires a meticulous equilibrium between biological activity, appropriate pharmacodynamic/pharmacokinetic parameters, and manageable toxicity. These diverse properties are interrelated yet distinct. To optimize multiple properties without compromising one another, researchers must either construct an ensemble model capable of predicting all intended properties or train multiple individual models to optimize each property sequentially [220, 222].
6 Conclusion
Undoubtedly, CADD and ML methods have been adopted by more and more pharmaceutical companies and research teams for designing novel anesthetic drugs. It is worth noting that Large Language Models (LLMs), such as ChatGPT, have made remarkable strides and garnered unprecedented attention recently. LLMs have also exhibited significant potential in bioinformatic analysis and drug discovery tasks, aiding researchers in generating functional protein sequences, identifying new potential targets, predicting toxic effects of drugs, interpreting drug-drug interactions, and extracting accurate pharmacological data from the scientific literature.
Despite the successful identification of several promising anesthetic compounds through computational methods, the development of new commercially available anesthetics remains a formidable challenge. Multiple obstacles continue to hinder progress, such as the scarcity of high-quality datasets, inherent limitations of computational methods, and the incomplete understanding of the underlying mechanisms of anesthesia. In a nutshell, further explorations of the physiological basis of anesthesia and continuous advancements in computational techniques are imperative. Only through these efforts can researchers develop novel anesthetics with reduced expense and shorter development timelines.
Availability of data and materials
Not applicable.
Abbreviations
- ADMET:
-
Absorption, distribution, metabolism, excretion, and toxicity
- AI:
-
Artificial Intelligence
- AUC-ROC:
-
Area under the receiver operator characteristic curve
- BBB:
-
Blood-brain barrier
- CADD:
-
Computer-aided drug design
- CNN:
-
Convolutional neural network
- CNS:
-
Central nervous system
- DL:
-
Deep learning
- DRL:
-
Deep reinforcement learning
- DT:
-
Decision tree
- DTL:
-
Deep transfer learning
- FDA:
-
Food and Drug Administration
- GABA:
-
Gamma-aminobutyric acid
- GB:
-
Gaussian naive bayes
- GBM:
-
Gradient boosting machine
- GNN:
-
Graph neural network
- GRU:
-
Gated recurrent unit
- HTS:
-
High-throughput screening
- K-NN:
-
K-nearest neighbor
- LBDD:
-
Ligand-based drug design
- LDA:
-
Linear discriminant analysis
- LLM:
-
Large language model
- LR:
-
Logistic regression
- LSTM:
-
Long short-term memory
- MAE:
-
Mean absolute error
- MD:
-
Molecular dynamics
- ML:
-
Machine learning
- MLP:
-
Multilayer perceptron
- MSE:
-
Mean squared error
- NaV:
-
Voltage-gated sodium channel
- NB:
-
Naive bayes
- OPRK1:
-
Opioid receptor Kappa 1
- P2X3R:
-
P2X3 receptor
- PCA:
-
Principle component analysis
- PD:
-
Pharmacokinetics
- PDB:
-
Protein Data Bank
- PIH:
-
Post-induction hypotension
- PK:
-
Pharmacodynamics
- PPI:
-
Protein-protein interaction
- QSAR:
-
Quantitative structure-affinity relationship
- RF:
-
Random forest
- RL:
-
Reinforcement learning
- RNN:
-
Recurrent neural network
- S1R:
-
Sigma 1 receptor
- SAR:
-
Structure-activity relationship
- SBDD:
-
Structure-based drug design
- SMOTE:
-
Synthetic minority oversampling technique
- SMILES:
-
Simplifiedmolecular-inputline-entry system
- SVM:
-
Support vector machine
- tICA:
-
Time-structure independent components analysis
- TOFR:
-
Train-of-four ratio
- VAE:
-
Variational autoencoder
- VS:
-
Virtual screening
- \(\mu\)OR:
-
mu-opioid receptor
References
Robinson DH, Toledo AH. Historical development of modern anesthesia. J Invest Surg. 2012;25(3):141–9. https://doi.org/10.3109/08941939.2012.690328.
Alkire MT, Hudetz AG, Tononi G. Consciousness and anesthesia. Science. 2008;322(5903):876–80. https://doi.org/10.1126/science.1149213.
Solt K, Forman SA. Correlating the clinical actions and molecular mechanisms of general anesthetics. Curr Opin Anaesthesiol. 2007;20(4):300–6. https://doi.org/10.1097/ACO.0b013e32816678a5.
Eger EI. Characteristics of anesthetic agents used for induction and maintenance of general anesthesia. AJHP. 2004;61 Suppl 4:S3–10. https://doi.org/10.1093/ajhp/61.suppl_4.S3.
Urban B, Bleckwenn M. Concepts and correlations relevant to general anaesthesia. Br J Anaesth. 2002;89(1):3–16. https://doi.org/10.1093/bja/aef164.
Rosenberg M, Weaver J. General anesthesia. Anesth Prog. 1991;38(4–5):172–86.
Yin J, Zhao Y, He Q, Hai A, Peng Y, Zuo Z, et al. Design, synthesis and biological evaluation of novel procaine derivatives for intravenous anesthesia. Bioorganic Med Chem Lett. 2022;60:128587. https://doi.org/10.1016/j.bmcl.2022.128587.
McCaughey W. Adverse effects of local anaesthetics. Drug Saf. 1992;7(3):178–89. https://doi.org/10.2165/00002018-199207030-00003.
Berthoud MC, Reilly CS. Adverse effects of general anaesthetics. Drug Saf. 1992;7(6):434–59. https://doi.org/10.2165/00002018-199207060-00005.
Cai Y, Nong L, Li H, Luo Q, Zhu Y, Shu H. Effect of bilateral superficial cervical plexus block on postoperative pain, nausea, and vomiting in thyroid surgery: a systematic review and meta-analysis. APS. 2023;1(2):13. https://doi.org/10.1007/s44254-023-00012-8.
Schneemilch CE, Schilling T, Bank U. Effects of general anaesthesia on inflammation. Best Pract Res Clin Anaesthesiol. 2004;18(3):493–507. https://doi.org/10.1016/j.bpa.2004.01.002.
Deng C, Gong D, Yang J, Ke B, Kang Y, Liu J, et al. New insights for screening etomidate analogues in the human H295R cell model. Toxicol in Vitro. 2020;68:104934. https://doi.org/10.1016/j.tiv.2020.104934.
Leelananda SP, Lindert S. Computational methods in drug discovery. Beilstein J Org Chem. 2016;12:2694–718. https://doi.org/10.1124/pr.112.007336.
Schaefer S, Kolkhof P. Failure is an option: learning from unsuccessful proof-of-concept trials. Drug Discov Today. 2008;13(21–22):913–6. https://doi.org/10.1016/j.drudis.2008.03.026.
Munos B. Lessons from 60 years of pharmaceutical innovation. Nat Rev Drug Discov. 2009;8(12):959–68. https://doi.org/10.1038/nrd2961.
Dickson M, Gagnon JP. The cost of new drug discovery and development. Discov Med. 2009;4(22):172–9.
Borhani DW, Shaw DE. The future of molecular dynamics simulations in drug discovery. J Comput Aided Mol Des. 2012;26(1):15–26. https://doi.org/10.1007/s10822-011-9517-y.
Gertrudes JC, Maltarollo VG, Silva R, Oliveira PR, Honorio KM, Da Silva A. Machine learning techniques and drug design. Curr Med Chem. 2012;19(25):4289–97. https://doi.org/10.2174/092986712802884259.
Sneyd JR. Thiopental to desflurane - an anaesthetic journey. Where are we going next? Br J Anaesth. 2017;119 Suppl 1:i44–52. https://doi.org/10.1093/bja/aex328.
Shelton CL, Sutton R, White SM. Desflurane in modern anaesthetic practice: walking on thin ice (caps)? Br J Anaesth. 2020;125(6):852–6. https://doi.org/10.1016/j.bja.2020.09.013.
Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR, et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 2018;46(D1):D1074–82. https://doi.org/10.1093/nar/gkx1037.
Yasiry Z, Shorvon SD. How phenobarbital revolutionized epilepsy therapy: the story of phenobarbital therapy in epilepsy in the last 100 years. Epilepsia. 2012;53 Suppl 8:26–39. https://doi.org/10.1111/epi.12026.
Chidambaran V, Costandi A, D’Mello A. Propofol: a review of its role in pediatric anesthesia and sedation. CNS Drugs. 2015;29(7):543–63. https://doi.org/10.1007/s40263-015-0259-6.
Lim GFS, Huether MJ, Brodland DG. 58 - Local Anesthetics. In: Wolverton SE, editor. Comprehensive Dermatologic Drug Therapy (Fourth Edition). Amsterdam: Elsevier; 2021. p. 631–49.e6. https://doi.org/10.1016/B978-0-323-61211-1.00058-9.
Skarda RT, Muir WW, Hubbell JAE. Chapter 11-Local anesthetic drugs and techniques. In: Muir WW, Hubbell JAE, editors. Equine anesthesia. 2nd ed. Saint Louis: W.B. Saunders 2009. p. 210–42. https://doi.org/10.1016/B978-1-4160-2326-5.00011-0.
Tetracaine. In: Aronson JK, editor. Meyler’s side effects of drugs. 16th ed. Oxford: Elsevier; 2016. p. 768–9. https://doi.org/10.1016/B978-0-444-53717-1.01522-5.
Gordh T, Gordh TE, Lindqvist K, Warner DS. Lidocaine: the origin of a modern local anesthetic. Anesthesiology. 2010;113(6):1433–7. https://doi.org/10.1097/ALN.0b013e3181fcef48.
Chapter 25 - Pharmacology. In: Malamed SF, editor. Sedation. 5th ed. Saint Louis: Mosby; 2010. p. 316–54. https://doi.org/10.1016/B978-0-323-05680-9.00029-1.
McClish A. Diazepam as an intravenous induction agent for general anaesthesia. Can Anaes Soc J. 1966;13(6):562–75. https://doi.org/10.1007/BF03002226.
Hammond R, Christie M, Nicholson A. Chapter 14 - Opioid analgesics. In: MADDISON JE, PAGE SW, CHURCH DB, editors. Small Animal Clinical Pharmacology. 2nd ed. Edinburgh: W.B. Saunders; 2008. p. 309–29. https://doi.org/10.1016/B978-070202858-8.50016-6.
Kim S, Chen J, Cheng T, Gindulyte A, He J, He S, et al. PubChem 2023 update. Nucleic Acids Res. 2022 10;51(D1):D1373–D80. https://doi.org/10.1093/nar/gkac956.
Atracurium dibesilate. In: Aronson JK, editor. Meyler’s side effects of drugs: the international encyclopedia of adverse drug reactions and interactions. 15th ed. Amsterdam: Elsevier; 2006. p. 370–4. https://doi.org/10.1016/B0-44-451005-2/00228-X.
Van Drie JH. Computer-aided drug design: the next 20 years. J Comput Aided Mol Des. 2007;21(10–11):591–601. https://doi.org/10.1007/s10822-007-9142-y.
Lionta E, Spyrou G, K Vassilatis D, Cournia Z. Structure-based virtual screening for drug discovery: principles, applications and recent advances. SCurr Top Med Chem. 2014;14(16):1923–38. https://doi.org/10.1208/s12248-012-9322-0.
Zhang S. Computer-aided drug discovery and development. Methods Mol Biol. 2011;716:23–38. https://doi.org/10.1007/978-1-61779-012-6_2.
Grinter SZ, Zou X. Challenges, applications, and recent advances of protein-ligand docking in structure-based drug design. Molecules. 2014;19(7):10150–76. https://doi.org/10.3390/molecules190710150.
Jorgensen WL. Pulled from a protein’s embrace. Nature. 2010;466(7302):42–3. https://doi.org/10.1038/466042a.
Taylor RD, Jewsbury PJ, Essex JW. A review of protein-small molecule docking methods. J Comput Aided Mol Des. 2002;16(3):151–66. https://doi.org/10.1023/A:1020155510718.
Meng XY, Zhang HX, Mezei M, Cui M. Molecular docking: a powerful approach for structure-based drug discovery. Curr Comput Aided Drug Des. 2011;7(2):146–57. https://doi.org/10.2174/157340911795677602.
Saikia S, Bordoloi M. Molecular docking: challenges, advances and its use in drug discovery perspective. Curr Drug Targets. 2019;20(5):501–21. https://doi.org/10.2174/1389450119666181022153016.
Morris GM, Lim-Wilby M. Molecular docking. In: Kukol A, editor. Molecular Modeling of Proteins. Humana Totowa: Humana Press; 2008. p. 365–382. https://doi.org/10.1007/978-1-59745-177-2_19.
Ferreira LG, Dos Santos RN, Oliva G, Andricopulo AD. Molecular docking and structure-based drug design strategies. Molecules. 2015;20(7):13384–421. https://doi.org/10.3390/molecules200713384.
Chaudhary KK, Mishra N. A review on molecular docking: novel tool for drug discovery. JSM Chem. 2016;4(3):1029.
Pinzi L, Rastelli G. Molecular docking: shifting paradigms in drug discovery. Int J Mol Sci. 2019;20(18):4331. https://doi.org/10.3390/ijms20184331.
Vyas V, Ukawala R, Ghate M, Chintha C. Homology modeling a fast tool for drug discovery: current perspectives. Indian J Pharm Sci. 2012;74(1):1–17. https://doi.org/10.4103/0250-474X.102537.
Floudas C, Fung H, McAllister S, Mönnigmann M, Rajgaria R. Advances in protein structure prediction and de novo protein design: A review. Chem Eng Sci. 2006;61(3):966–88. https://doi.org/10.1016/j.ces.2005.04.009.
Imam SS, Gilani SJ. Computer aided drug design: A novel loom to drug discovery. Org Med Chem Int J. 2017;1(4):002 555567. https://doi.org/10.19080/OMCIJ.2017.01.555567.
Sanjeevi M, Hebbar PN, Aiswarya N, Rashmi S, Rahul CN, Mohan A, et al. Methods and applications of machine learning in structure-based drug discovery. In: Tripathi T, Dubey VK, editors. Advances in Protein Molecular and Structural Biology Methods. London: Academic Press; 2022. p. 405–437. https://doi.org/10.1016/B978-0-323-90264-9.00025-8.
Sun H. Homology modeling and ligand-based molecule design. In: A practical guide to rational drug design. Sawston: Woodhead Publishing; 2016. p. 109–60. https://doi.org/10.1016/B978-0-08-100098-4.00004-1.
Abeln S, Feenstra KA, Heringa J. Protein three-dimensional structure prediction. In: Ranganathan S, Gribskov M, Nakai K, Schönbach C, editors. Encyclopedia of bioinformatics and computational biology: ABC of bioinformatics. Netherlands: Elsevier; 2019. p. 497–511.
Karplus M, McCammon JA. Molecular dynamics simulations of biomolecules. Nat StructBiol. 2002;9(9):646–52. https://doi.org/10.1038/nsb0902-646.
Shukla R, Tripathi T. Molecular dynamics simulation in drug discovery: opportunities and challenges. In: Singh SK, editor. Innovations and implementations of computer aided drug discovery strategies in rational drug design. Singapore: Springer; 2021. p. 295–316. https://doi.org/10.1007/978-981-15-8936-2_12.
Battimelli G, Ciccotti G. Berni Alder and the pioneering times of molecular simulation. Eur Phys J Plus. 2018;43:303–35. https://doi.org/10.1140/epjh/e2018-90027-5.
Ganesan A, Coote ML, Barakat K. Molecular dynamics-driven drug discovery: leaping forward with confidence. Drug Discov Today. 2017;22(2):249–69. https://doi.org/10.1016/j.drudis.2016.11.001.
Salo-Ahen OMH, Alanko I, Bhadane R, Bonvin AM, Honorato RV, Hossain S, et al. Molecular dynamics simulations in drug discovery and pharmaceutical development. Processes. 2020;9(1):71. https://doi.org/10.3390/pr9010071.
Durrant JD, McCammon JA. Molecular dynamics simulations and drug discovery. BMC Biol. 2011;9:71. https://doi.org/10.1186/1741-7007-9-71.
De Vivo M, Masetti M, Bottegoni G, Cavalli A. Role of molecular dynamics and related methods in drug discovery. J Med Chem. 2016;59(9):4035–61. https://doi.org/10.1021/acs.jmedchem.5b01684.
Liu X, Shi D, Zhou S, Liu H, Liu H, Yao X. Molecular dynamics simulations and novel drug discovery. Expert Opin Drug Discov. 2018;13(1):23–37. https://doi.org/10.1080/17460441.2018.1403419.
Bajorath J. Integration of virtual and high-throughput screening. Nat Rev Drug Discov. 2002;1(11):882–94. https://doi.org/10.1038/nrd941.
Maia EHB, Assis LC, De Oliveira TA, Da Silva AM, Taranto AG. Structure-based virtual screening: from classical to artificial intelligence. Front Chem. 2020;8:343. https://doi.org/10.3389/fchem.2020.00343.
Subramaniam S, Mehrotra M, Gupta D. Virtual high throughput screening (vHTS)-A perspective. Bioinformation. 2008;3(1):14–7. https://doi.org/10.6026/97320630003014.
Sliwoski G, Kothiwale S, Meiler J, Lowe EW. Computational methods in drug discovery. Pharmacol Rev. 2014;66(1):334–95. https://doi.org/10.1124/pr.112.007336.
Hung CL, Chen CC. Computational approaches for drug discovery. Drug Dev Res. 2014;75(6):412–8. https://doi.org/10.1002/ddr.21222.
Macalino SJY, Gosu V, Hong S, Choi S. Role of computer-aided drug design in modern drug discovery. Arch Pharm Res. 2015;38(9):1686–701. https://doi.org/10.1007/s12272-015-0640-5.
Johnson MA, Maggiora GM. Concepts and applications of molecular similarity. New York: Wiley; 1990.
Yu W, MacKerell AD. Computer-aided drug design methods. Methods Mol Biol. 2017;1520:85–106. https://doi.org/10.1007/978-1-4939-6634-9_5.
Kaushik AC, Kumar A, Bharadwaj S, Chaudhary R, Sahi S. Ligand-based approach for in-silico drug designing. Bioinformatics techniques for drug discovery: applications for complex diseases. Berlin: Springer Cham; 2018. p. 11–9. https://doi.org/10.1007/978-3-319-75732-2_2.
Muegge I, Mukherjee P. An overview of molecular fingerprint similarity search in virtual screening. Expert Opin Drug Discov. 2016;11(2):137–48. https://doi.org/10.1517/17460441.2016.1117070.
Willett P, Barnard JM, Downs GM. Chemical similarity searching. J Chem Inf Comput Sci. 1998;38(6):983–96. https://doi.org/10.1021/ci9800211.
Shim J, MacKerell AD Jr. Computational ligand-based rational design: role of conformational sampling and force fields in model development. MedChemComm. 2011;2(5):356–70. https://doi.org/10.1039/C1MD00044F.
Qing X, Lee XY, De Raeymaecker J, Tame JR, Zhang KY, De Maeyer M, et al. Pharmacophore modeling: advances, limitations, and current utility in drug discovery. J Recept Ligand Channel Res. 2014;7:81–92. https://doi.org/10.2147/JRLCR.S46843.
Langer T, Wolber G. Pharmacophore definition and 3D searches. Drug Discov Today. 2004;1(3):203–7. https://doi.org/10.1016/j.ddtec.2004.11.015.
Koes DR. Pharmacophore modeling: methods and applications. In: Zhang W, editor. Computer-aided drug discovery. New York: Humana; 2016. p. 167–88. https://doi.org/10.1007/7653_2015_46.
Yang SY. Pharmacophore modeling and applications in drug discovery: challenges and recent advances. Drug Discov Today. 2010;15(11–12):444–50. https://doi.org/10.1016/j.drudis.2010.03.013.
Gramatica P. On the development and validation of QSAR models. Methods Mol Biol. 2013;930:499–526. https://doi.org/10.1007/978-1-62703-059-5_21.
Tropsha A. Best practices for QSAR model development, validation, and exploitation. Mol Inform. 2010;29(6–7):476–88.
Cherkasov A, Muratov EN, Fourches D, Varnek A, Baskin II, Cronin M, et al. QSAR modeling: where have you been? Where are you going to? J Med Chem. 2014;57(12):4977–5010. https://doi.org/10.1021/jm4004285.
Maggiora GM. On outliers and activity cliffs why QSAR often disappoints. 2006;46(4):1535. https://doi.org/10.1021/ci060117s.
Verma J, Khedkar VM, Coutinho EC. 3D-QSAR in drug design-a review. Curr Top Med Chem. 2010;10(1):95–115. https://doi.org/10.2174/156802610790232260.
Hopfinger A, Wang S, Tokarski JS, Jin B, Albuquerque M, Madhav PJ, et al. Construction of 3D-QSAR models using the 4D-QSAR analysis formalism. J Am Chem Soc. 1997;119(43):10509–24. https://doi.org/10.1021/ja9718937.
Jin W, Barzilay R, Jaakkola T. Junction tree variational autoencoder for molecular graph generation. In: Dy J, Krause A, editors. Proceedings of the 35th International Conference on Machine Learning, 2018 July 10–15; Stockholm, Sweden. Stockholm: ICME; 2018. p. 2323–32. https://doi.org/10.48550/arXiv.1802.04364.
Shinde PP, Shah S. A review of machine learning and deep learning applications. In: 2018 Fourth international conference on computing communication control and automation (ICCUBEA); 2018 August 16–18; Pune, India. New York: IEEE; 2018. https://doi.org/10.1109/ICCUBEA.2018.8697857.
Jordan MI, Mitchell TM. Machine learning: Trends, perspectives, and prospects. Science. 2015;349(6245):255–60. https://doi.org/10.1126/science.aaa8415.
Wang H, Ma C, Zhou L. A brief review of machine learning and its application. In: International Conference on Information Engineering and Computer Science; 2009 December 19–20; Wuhan, China. New York: IEEE; 2009. p. 1–4. https://doi.org/10.1109/ICIECS.2009.5362936.
Dhall D, Kaur R, Juneja M. Machine learning: a review of the algorithms and its applications. In: Singh PK, Kar AK, Singh Y, Kolekar MH, Tanwar S, editors. Proceedings of ICRIC 2019: Recent Innovations in Computing. 1st ed. Berlin: Springer; 2020. p. 47–63. https://doi.org/10.1007/978-3-030-29407-6_5.
Nasteski V. An overview of the supervised machine learning methods. Horizons b. 2017;4:51–62.
Kwofie SK, Adams J, Broni E, Enninful KS, Agoni C, Soliman ME, et al. Artificial intelligence, machine learning, and big data for ebola virus drug discovery. Pharmaceuticals. 2023;16(3):332. https://doi.org/10.3390/ph16030332.
Zhang H, Ding L, Zou Y, Hu SQ, Huang HG, Kong WB, et al. Predicting drug-induced liver injury in human with Naïve Bayes classifier approach. J Comput AID Mol Des. 2016;30(10):889–98. https://doi.org/10.1007/s10822-016-9972-6.
Sun S, Huang R. An adaptive k-nearest neighbor algorithm. In: 2010 seventh international conference on fuzzy systems and knowledge discovery; 2010 August 10–12; Yantai, China. New York: IEEE; 2010. p. 91–4. https://doi.org/10.1109/FSKD.2010.5569740.
Taunk K, De S, Verma S, Swetapadma A. A brief review of nearest neighbor algorithm for learning and classification. In: 2019 International Conference on Intelligent Computing and Control Systems (ICCS); 2019 May 15–17; Madurai: India. New York: IEEE; 2019. p. 1255–60. https://doi.org/10.1109/ICCS45141.2019.9065747.
Rodríguez-Pérez R, Bajorath J. Evolution of support vector machine and regression modeling in chemoinformatics and drug discovery. J Comput AID Mol Des. 2022;36(5):355–62. https://doi.org/10.1007/s10822-022-00442-9.
Safavian SR, Landgrebe D. A survey of decision tree classifier methodology. IEEE Trans Syst Man Cybern. 1991;21(3):660–74. https://doi.org/10.1109/21.97458.
Wu X, Kumar V, Ross Quinlan J, Ghosh J, Yang Q, Motoda H, et al. Top 10 algorithms in data mining. Knowl Inf Syst. 2008;14:1–37. https://doi.org/10.1007/s10115-007-0114-2.
Hammann F, Drewe J. Decision tree models for data mining in hit discovery. Expert Opin Drug Dis. 2012;7(4):341–52. https://doi.org/10.1517/17460441.2012.668182.
Natekin A, Knoll A. Gradient boosting machines, a tutorial. Front Neurorobotics. 2013;7:21. https://doi.org/10.3389/fnbot.2013.00021.
Biau G, Scornet E. A random forest guided tour. Test. 2016;25:197–227. https://doi.org/10.1007/s11749-016-0481-7.
Korkmaz S. Deep learning-based imbalanced data classification for drug discovery. J Chem Inf Model. 2020;60(9):4180–90. https://doi.org/10.1021/acs.jcim.9b01162.
Wang X, Yu B, Ma A, Chen C, Liu B, Ma Q. Protein-protein interaction sites prediction by ensemble random forests with synthetic minority oversampling technique. Bioinformatics. 2019;35(14):2395–402. https://doi.org/10.1093/bioinformatics/bty995.
Weisberg S. Applied linear regression. 4th ed. Hoboken: Wiley; 2005. https://doi.org/10.1002/0471704091.
Sperandei S. Understanding logistic regression analysis. Biochem Medica. 2014;24(1):12–18. https://doi.org/10.11613/BM.2014.003.
Pham DT, Dimov SS, Nguyen CD. Selection of K in K-means clustering. P I Mech Eng C-J Mec. 2005;219(1):103–19. https://doi.org/10.1243/095440605X8298.
Johnson SC. Hierarchical clustering schemes. Psychometrika. 1967;32(3):241–54. https://doi.org/10.1007/BF02289588.
Lever J, Krzywinski M, Altman N. Points of significance: Principal component analysis. Nat Methods. 2017;14(7):641–2. https://doi.org/10.1038/nmeth.4346.
Ioffe S. Probabilistic linear discriminant analysis. In: Leonardis A, Bischof H, Pinz A, editors. Computer Vision–ECCV 2006: 9th European Conference on Computer Vision, Proceedings, Part IV 9; 2006 May 7-13; Graz, Austria. Heidelberg: Springer Berlin; 2006. p. 531–42. https://doi.org/10.1007/11744085_41.
Krogh A. What are artificial neural networks? Nat Biotechnol. 2008;26:195–7. https://doi.org/10.1038/nbt1386.
Jain AK, Mao J, Mohiuddin KM. Artificial neural networks: A tutorial. Computer. 1996;29(3):31–44. https://doi.org/10.1109/2.485891.
Abirami SP, Chitra P. Chapter Fourteen - Energy-efficient edge based real-time healthcare support system. Adv Comput. 2020;117(1):339–68. https://doi.org/10.1016/bs.adcom.2019.09.007.
Lillicrap TP, Santoro A, Marris L, Akerman CJ, Hinton G. Backpropagation and the brain. Nat Rev Neurosci. 2020;21(6):335–46. https://doi.org/10.1038/s41583-020-0277-3.
Ajit A, Acharya K, Samanta A. A Review of Convolutional Neural Networks. In: 2020 International Conference on Emerging Trends in Information Technology and Engineering (ic-ETITE); 2020 February 24–25; Vellore, India. New York: IEEE; 2020. p. 1–5. https://doi.org/10.1109/ic-ETITE47903.2020.049.
Li Z, Liu F, Yang W, Peng S, Zhou J. A survey of convolutional neural networks: analysis, applications, and prospects. IEEE T Neur Net Lear. 2021;33(12):6999–7019. https://doi.org/10.1109/TNNLS.2021.3084827.
Gu J, Wang Z, Kuen J, Ma L, Shahroudy A, Shuai B, et al. Recent advances in convolutional neural networks. Pattern Recogn. 2018;77:354–77. https://doi.org/10.1016/j.patcog.2017.10.013.
Zhou J, Cui G, Hu S, Zhang Z, Yang C, Liu Z, et al. Graph neural networks: A review of methods and applications. AI Open. 2020;1:57–81. https://doi.org/10.1016/j.aiopen.2021.01.001.
Bongini P, Bianchini M, Scarselli F. Molecular generative graph neural networks for drug discovery. Neurocomputing. 2021;450:242–52. https://doi.org/10.1016/j.neucom.2021.04.039.
Liu X, Luo Y, Li P, Song S, Peng J. Deep geometric representations for modeling effects of mutations on protein-protein binding affinity. PLoS Comput Biol. 2021;17(8):e1009284. https://doi.org/10.1371/journal.pcbi.1009284.
Yu Y, Si X, Hu C, Zhang J. A review of recurrent neural networks: LSTM cells and network architectures. Neural Comput. 2019;31(7):1235–70. https://doi.org/10.1162/neco_a_01199.
Fu R, Zhang Z, Li L. Using LSTM and GRU neural network methods for traffic flow prediction. In: 2016 31st Youth Academic Annual Conference of Chinese Association of Automation (YAC); 2016 November 11–13; Wuhan: China. New York: IEEE; 2016. p. 324–328. https://doi.org/10.1109/YAC.2016.7804912.
Li L, Zhang S, Wang B. Apple leaf disease identification with a small and imbalanced dataset based on lightweight convolutional networks. Sensors. 2022;22(1):173. https://doi.org/10.3390/s22010173.
Zhuang D, Ibrahim AK. Deep learning for drug discovery: a study of identifying high efficacy drug compounds using a cascade transfer learning approach. Appl Sci. 2021;11(17):7772. https://doi.org/10.3390/app11177772.
Cai C, Wang S, Xu Y, Zhang W, Tang K, Ouyang Q, et al. Transfer Learning for Drug Discovery. J Med Chem. 2020;63(16):8683–94. https://doi.org/10.1021/acs.jmedchem.9b02147.
Zhuang F, Qi Z, Duan K, Xi D, Zhu Y, Zhu H, et al. A comprehensive survey on transfer learning. Proceedings of the IEEE. 2020;109(1):43–76. https://doi.org/10.48550/arXiv.1911.02685.
Kaelbling LP, Littman ML, Moore AW. Reinforcement learning: A survey. J Artif Intell Res. 1996;4:237–285. https://doi.org/10.48550/arXiv.cs/9605103.
Zhou Z, Kearnes S, Li L, Zare RN, Riley P. Optimization of molecules via deep reinforcement learning. Sci Rep. 2019;9(1):10752. https://doi.org/10.1038/s41598-019-47148-x.
Tan RK, Liu Y, Xie L. Reinforcement learning for systems pharmacology-oriented and personalized drug design. Expert Opin Drug Dis. 2022;17(8):849–63. https://doi.org/10.1080/17460441.2022.2072288.
Gottipati SK, Sattarov B, Niu S, Pathak Y, Wei H, Liu S, et al. Learning to navigate the synthetically accessible chemical space using reinforcement learning. In: III HD, Singh A, editors. Proceedings of the 37th International Conference on Machine Learning; 2020 July 13–18. Online: PMLR; 2020. p. 3668–79. https://doi.org/10.48550/arXiv.2004.12485.
Dulac-Arnold G, Levine N, Mankowitz DJ, Li J, Paduraru C, Gowal A, et al. Challenges of real-world reinforcement learning. Mach Learn. 2021;110:2419–68. https://doi.org/10.1007/s10994-021-05961-4.
Elgarf AA, Siebert DC, Steudle F, Draxler A, Li G, Huang S, et al. Different benzodiazepines bind with distinct binding modes to \({\rm GABA_{A}}\) receptors. ACS Chem Biol. 2018;13(8):2033–9. https://doi.org/10.1021/acschembio.8b00144.
Pan C, Meng H, Zhang S, Zuo Z, Shen Y, Wang L, et al. Homology modeling and 3D-QSAR study of benzhydrylpiperazine \(\delta\) opioid receptor agonists. Comput Biol Chem. 2019;83:107109. https://doi.org/10.1016/j.compbiolchem.2019.107109.
Bertaccini EJ, Shapiro J, Brutlag DL, Trudell JR. Homology modeling of a human glycine alpha 1 receptor reveals a plausible anesthetic binding site. J Chem Inf Model. 2005;45(1):128–35. https://doi.org/10.1021/ci0497399.
Bertaccini EJ, Wallner B, Trudell JR, Lindahl E. Modeling anesthetic binding sites within the glycine alpha one receptor based on prokaryotic ion channel templates: the problem with TM4. J Chem Inf Model. 2010;50(12):2248–55. https://doi.org/10.1021/ci100266c.
Bertaccini EJ, Yoluk O, Lindahl ER, Trudell JR. Assessment of homology templates and an anesthetic binding site within the \(\gamma\)-aminobutyric acid receptor. Anesthesiology. 2013;119(5):1087–95. https://doi.org/10.1097/ALN.0b013e31829e47e3.
Yuan X, Zhang D, Mao S, Wang Q. Filling the Gap in Understanding the Mechanism of \({\rm GABA_{A}R}\) and Propofol Using Computational Approaches. J Chem Inf Model. 2021;61(4):1889–901. https://doi.org/10.1021/acs.jcim.0c01290.
Chandra S, Wang Z, Tao X, Chen O, Luo X, Ji RR, et al. Computer-aided discovery of a new Nav1.7 inhibitor for treatment of pain and itch. Anesthesiology. 2020;133(3):611–627. https://doi.org/10.1097/ALN.0000000000003427.
Manzur-Villalobos I, Contreras-Puentes N, Díaz-Caballero A, Durán-Lengua M, Alviz-Amador A. In silico study of local anesthetics analogues on sodium channel Nav1.7 a pharmacological target on inflamed dental pulp. Inform Med Unlocked. 2022;34:101117. https://doi.org/10.1016/j.imu.2022.101117.
Feinberg EN, Farimani AB, Uprety R, Hunkele A, Pasternak GW, Majumdar S, et al. Machine Learning Harnesses Molecular Dynamics to Discover New \(\mu\) Opioid Chemotypes. arXiv:1803.04479v1[Preprint]. 2018. p. 28. [Cited 2023 June 17]. Available from: https://doi.org/10.48550/arXiv.1803.04479.
Meraj K, Mahto MK, Christina NB, Desai N, Shahbazi S, Bhaskar M. Molecular modeling, docking and ADMET studies towards development of novel Disopyramide analogs for potential inhibition of human voltage gated sodium channel proteins. Bioinformation. 2012;8(23):1139–46. https://doi.org/10.6026/97320630081139.
Lv M, Wang XT, Li K, Wang YL. \({\rm GABA_{A}}\)receptor binding molecules from Traditional Chinese Medicine: An in silico approach. Bangladesh J Pharmacol. 2014;9:447–51. https://doi.org/10.3329/bjp.v9i4.19731.
Liu R, Perez-Aguilar JM, Liang D, Saven JG. Binding site and affinity prediction of general anesthetics to protein targets using docking. Anesth Analg. 2012;114(5):947. https://doi.org/10.1213/ANE.0b013e31824c4def.
Jiang Jh, Xu Xq, Jiang Wg, Wang T, Liu X, Zeng Lg, et al. Discovery of the EL-0052 as a potential anesthetic drug. Comput Struct Biotechnol J. 2021;19:710–8. https://doi.org/10.1016/j.csbj.2021.01.002.
Lima Neto JX, Bezerra KS, Barbosa ED, Oliveira JIN, Manzoni V, Soares-Rachetti VP, et al. Exploring the binding mechanism of \({\rm GABA_{B}}\) receptor agonists and antagonists through in silico simulations. J Chem Inf Model. 2020;60(2):1005–18. https://doi.org/10.1021/acs.jcim.9b01025.
Cayla NS, Dagne BA, Wu Y, Lu Y, Rodriguez L, Davies DL, et al. A newly developed anesthetic based on a unique chemical core. Proc Natl Acad Sci U S A. 2019;116(31):15706–15. https://doi.org/10.1073/pnas.1822076116.
Yang X, Jounaidi Y, Dai JB, Marte-Oquendo F, Halpin ES, Brown LE, et al. High-throughput screening in larval zebrafish identifies novel potent sedative-hypnotics. Anesthesiology. 2018;129(3):459–76. https://doi.org/10.1097/ALN.0000000000002281.
Peng Y, Dong H, Welsh WJ. Comprehensive 3D-QSAR model predicts binding affinity of structurally diverse sigma 1 receptor ligands. J Chem Inf Model. 2018;59(1):486–97. https://doi.org/10.1021/acs.jcim.8b00521.
Peng QX, Guan XH, Yi ZG, Su YP. Insilico approaches in anesthetic drug development: Computer aided drug designing. Drug Res (Stuttg). 2015;65(11):587–91. https://doi.org/10.1055/s-0034-1395564.
Lea WA, Xi J, Jadhav A, Lu L, Austin CP, Simeonov A, et al. A high-throughput approach for identification of novel general anesthetics. PLoS ONE. 2009;4(9):e7150. https://doi.org/10.1371/journal.pone.0007150.
McKinstry-Wu AR, Bu W, Rai G, Lea WA, Weiser BP, Liang DF, et al. Discovery of a novel general anesthetic chemotype using high-throughput screening. Anesthesiology. 2015;122(2):325–33. https://doi.org/10.1097/ALN.0000000000000505.
Ebalunode JO, Dong X, Ouyang Z, Liang J, Eckenhoff RG, Zheng W. Structure-based shape pharmacophore modeling for the discovery of novel anesthetic compounds. Bioorg Med Chem. 2009;17(14):5133–8. https://doi.org/10.1016/j.bmc.2009.05.060.
Cheng S, Ding Y. Construction of QSAR model between the ligand and \(\gamma\)-Aminobutyric acid type A receptor using support vector regression algorithm. In: 2020 19th International Symposium on Distributed Computing and Applications for Business Engineering and Science (DCABES). 2020 October 16–19; Xuzhou, China. New York: IEEE; 2020. p. 202–5. https://doi.org/10.1109/DCABES50732.2020.00060.
Lu A, Zhou J. Pseudoreceptor models and 3D-QSAR for imidazobenzodiazepines at \({\rm GABA_{A}}\)/BzR subtypes \(\alpha _{x}\)\(\beta _{3}\)\(\gamma _{2}\) [x= 1–3, 5, and 6] via flexible atom receptor model. J Chem Inf Comput Sci. 2004;44(3):1130–6. https://doi.org/10.1021/ci034281g.
Jia X, Ciallella HL, Russo DP, Zhao L, James MH, Zhu H. Construction of a virtual opioid bioprofile: a data-driven QSAR modeling study to identify new analgesic opioids. ACS Sustain Chem Eng. 2021;9(10):3909–19. https://doi.org/10.1021/acssuschemeng.0c09139.
Mehdipour AR, Hemmateenejad B, Miri R. QSAR studies on the anesthetic action of some polyhalogenated ethers. Chem Biol Drug Des. 2007;69(5):362–8. https://doi.org/10.1111/j.1747-0285.2007.00506.x.
Coli Louvisse de Abreu L, Abrahim-Vieira BA, Mendonça Teles de Souza A, Pinto EC, Gonçalves MDS, Simon A, et al. Forced degradation studies of norepinephrine and epinephrine from dental anesthetics: Development of stability-indicating HPLC method and in silico toxicity evaluation. Biomed Chromatogr. 2020;34(7):e4832. https://doi.org/10.1002/bmc.4832.
Azamatov AA, Zhurakulov SN, Vinogradova VI, Tursunkhodzhaeva F, Khinkar RM, Malatani RT, et al. Evaluation of the Local Anesthetic Activity, Acute Toxicity, and Structure-Toxicity Relationship in Series of Synthesized 1-Aryltetrahydroisoquinoline Alkaloid Derivatives In Vivo and In Silico. Molecules. 2023;28(2):477. https://doi.org/10.3390/molecules28020477.
Floresta G, Rescifina A, Abbate V. Structure-Based Approach for the Prediction of Mu-opioid Binding Affinity of Unclassified Designer Fentanyl-Like Molecules. Int J Mol Sci. 2019;20(9):2311. https://doi.org/10.3390/ijms20092311.
Sripriya Akondi V, Menon V, Baudry J, Whittle J. Novel big data-driven machine learning models for drug discovery application. Molecules. 2022;27(3):594. https://doi.org/10.3390/molecules27030594.
Kendale S, Kulkarni P, Rosenberg AD, Wang J. Supervised machine-learning predictive analytics for prediction of postinduction hypotension. Anesthesiology. 2018;129(4):675–88. https://doi.org/10.1097/ALN.0000000000002374.
Jagannathan R. Identification of psychoactive metabolites from Cannabis sativa, its smoke, and other phytocannabinoids using machine learning and multivariate methods. ACS OMEGA. 2020;5(1):281–95. https://doi.org/10.1021/acsomega.9b02663.
Liao Z, Huang Y, Yue X, Lu H, Xuan P, Ju Y. In silico prediction of gamma-aminobutyric acid type-A receptors using novel machine-learning-based SVM and GBDT approaches. BioMed Res Int. 2016;2016:2375268. https://doi.org/10.1155/2016/2375268.
Mohabatkar H, Beigi MM, Esmaeili A. Prediction of \({\rm {GABA_{A}}}\) receptor proteins using the concept of Chou’s pseudo-amino acid composition and support vector machine. J Theor Biol. 2011;281(1):18–23. https://doi.org/10.1016/j.jtbi.2011.04.017.
Gao M, Igata H, Takeuchi A, Sato K, Ikegaya Y. Machine learning-based prediction of adverse drug effects: an example of seizure-inducing compounds. J Pharmacol Sci. 2017;133(2):70–8. https://doi.org/10.1016/j.jphs.2017.01.003.
Dhandore A, Mhatre P, Bhole K. Prediction of Drug Events using Machine Learning. In: 2022 4th International Conference on Advances in Computing, Communication Control and Networking (ICAC3N); 2022 December 16–17; Greater Noida, India. New York: IEEE; 2022. p. 577–82. https://doi.org/10.1109/ICAC3N56670.2022.10074327.
Zhang H, Li W, Xie Y, Wang WJ, Li LL, Yang SY. Rapid and accurate assessment of seizure liability of drugs by using an optimal support vector machine method. Toxicol in Vitro. 2011;25(8):1848–54. https://doi.org/10.1016/j.tiv.2011.05.015.
Yu TH, Su BH, Battalora LC, Liu S, Tseng YJ. Ensemble modeling with machine learning and deep learning to provide interpretable generalized rules for classifying CNS drugs with high prediction power. Brief Bioinform. 2022;23(1):bbab377. https://doi.org/10.1093/bib/bbab377.
Lundberg SM, Nair B, Vavilala MS, Horibe M, Eisses MJ, Adams T, et al. Explainable machine-learning predictions for the prevention of hypoxaemia during surgery. Nat Biomed Eng. 2018;2(10):749–60. https://doi.org/10.1038/s41551-018-0304-0.
Wijeyesakere SJ, Wilson D, Auernhammer T, Parks A, Marty S. Machine-Learning Model Predicts Interaction with \(\gamma\)-Amino Butyric Acid (GABA) ergic Neurotransmission. Appl In Vitro Toxicol. 2022;8(4):129–38. https://doi.org/10.1089/aivt.2021.0028.
Deng J, Yang Z, Li Y, Samaras D, Wang F. Towards better opioid antagonists using deep reinforcement learning. arXiv:2004.04768v1[Preprint]. 2004 [Cited 2023 June 17]: [10 p.]. Available from: https://doi.org/10.48550/arXiv.2004.04768.
Kang KM, Lee I, Nam H, Kim YC. AI-based prediction of new binding site and virtual screening for the discovery of novel P2X3 receptor antagonists. Eur J Med Chem. 2022;240:114556. https://doi.org/10.1016/j.ejmech.2022.114556.
Wang K, Gao B, Liu H, Chen H, Liu H. The Real-Time and Patient-Specific Prediction for Duration and Recovery Profile of Cisatracurium Based on Deep Learning Models. Front Pharmacol. 2022;12:831149. https://doi.org/10.3389/fphar.2021.831149.
Nagata S, Fujiwara K, Kuga K, Ozaki H. Prediction of GABA receptor antagonist-induced convulsion in cynomolgus monkeys by combining machine learning and heart rate variability analysis. J Pharmacol Toxicol Methods. 2021;112:107127. https://doi.org/10.1016/j.vascn.2021.107127.
Schenone M, Dančík V, Wagner BK, Clemons PA. Target identification and mechanism of action in chemical biology and drug discovery. Nat Chem Biol. 2013;9(4):232–40. https://doi.org/10.1038/nchembio.1199.
Chen XP, Du GH. Target validation: A door to drug discovery. Drug Discov Ther. 2007;1(1):23–9.
Evangelista W, Weir RL, Ellingson SR, Harris JB, Kapoor K, Smith JC, et al. Ensemble-based docking: From hit discovery to metabolism and toxicity predictions. Bioorg Med Chem. 2016;24(20):4928–35. https://doi.org/10.1016/j.bmc.2016.07.064.
Dill KA, Ozkan SB, Shell MS, Weikl TR. The protein folding problem. Annu Rev Biophys. 2008;37:289–316. https://doi.org/10.1146/annurev.biophys.37.092707.153558.
Jazvinscak Jembrek M, Vlainic J. GABA receptors: pharmacological potential and pitfalls. Curr Pharm Des. 2015;21(34):4943–59.
Enna SJ. The GABA receptors. In: Enna SJ, Möhler H, editors. The GABA Receptors. Totowa: Springer; 2007. p. 1–12. https://doi.org/10.1007/978-1-59745-465-0_1.
Chou KC. Prediction of protein cellular attributes using pseudo-amino acid composition. Proteins. 2001;43(3):246–55. https://doi.org/10.1002/prot.1035.
Vanderah TW. Delta and kappa opioid receptors as suitable drug targets for pain. Clin J Pain. 2010;26 Suppl 10:S10–5. https://doi.org/10.1097/AJP.0b013e3181c49e3a.
Lindsay MA. Target discovery. Nat Rev Drug Discov. 2003;2(10):831–8. https://doi.org/10.1038/nrd1202.
Yang Y, Adelstein SJ, Kassis AI. Target discovery from data mining approaches. Drug Discov Today. 2012;17:S16–23. https://doi.org/10.1016/j.drudis.2011.12.006.
Sams-Dodd F. Drug discovery: selecting the optimal approach. Drug Discov Today. 2006;11(9–10):465–72. https://doi.org/10.1016/j.drudis.2006.03.015.
Fu Y, Zhao J, Chen Z. Insights into the molecular mechanisms of protein-ligand interactions by molecular docking and molecular dynamics simulation: A case of oligopeptide binding protein. Comput Math Methods Med. 2018;2018:3502514. https://doi.org/10.1155/2018/3502514.
Li L, Koh CC, Reker D, Brown JB, Wang H, Lee NK, et al. Predicting protein-ligand interactions based on bow-pharmacological space and Bayesian additive regression trees. Sci Rep. 2019;9(1):7703. https://doi.org/10.1038/s41598-019-43125-6.
Du X, Li Y, Xia YL, Ai SM, Liang J, Sang P, et al. Insights into protein-ligand interactions: mechanisms, models, and methods. Int J Mol Sci. 2016;17(2):144. https://doi.org/10.3390/ijms17020144.
Yu FH, Catterall WA. Overview of the voltage-gated sodium channel family. Genome Biol. 2003;4(3):207. https://doi.org/10.1186/gb-2003-4-3-207.
Kalyaanamoorthy S, Chen YPP. Structure-based drug design to augment hit discovery. Drug Discov Today. 2011;16(17–18):831–9. https://doi.org/10.1016/j.drudis.2011.07.006.
Hevener KE, Pesavento R, Ren J, Lee H, Ratia K, Johnson ME. Chapter Twelve - Hit-to-Lead: Hit Validation and Assessment. In: Lesburg CA, editor. Modern Approaches in Drug Discovery, vol. 610. Boston: Elsevier; 2018. p. 265–309. https://doi.org/10.1016/bs.mie.2018.09.022.
Mayr LM, Bojanic D. Novel trends in high-throughput screening. Curr Opin Pharmacol. 2009;9(5):580–8. https://doi.org/10.1016/j.coph.2009.08.004.
Hertzberg RP, Pope AJ. High-throughput screening: new technology for the 21st century. Curr Opin Chem Biol. 2000;4(4):445–51. https://doi.org/10.1016/S1367-5931(00)00110-1.
Entzeroth M, Flotow H, Condron P. Overview of high-throughput screening. Curr Protoc Pharmacol. 2009;44(1):9–4. https://doi.org/10.1002/0471141755.ph0904s44.
Liu B, Li S, Hu J. Technological advances in high-throughput screening. Am J Pharmacogenomics. 2004;4(4):263–76. https://doi.org/10.1016/S1367-5931(98)80089-6.
Zuo Z, Kang T, Hu S, Su W, Gan Y, Miao Z, et al. A bioluminescent probe for detecting norepinephrine in vivo. Anal Chem. 2022;94(17):6441–5. https://doi.org/10.1021/acs.analchem.2c00460.
Hartenfeller M, Schneider G. De novo drug design. Methods Mol Biol. 2011;672:299–323. https://doi.org/10.1007/978-1-60761-839-3_12.
Popova M, Isayev O, Tropsha A. Deep reinforcement learning for de novo drug design. Sci Adv. 2018;4(7):eaap7885. https://doi.org/10.1126/sciadv.aap7885.
Gupta A, Müller AT, Huisman BJ, Fuchs JA, Schneider P, Schneider G. Generative recurrent networks for de novo drug design. Mol Inform. 2018;37(1–2):1700111. https://doi.org/10.1002/minf.201700111.
Mouchlis VD, Afantitis A, Serra A, Fratello M, Papadiamantis AG, Aidinis V, et al. Advances in de novo drug design: from conventional to machine learning methods. Int J Mol Sci. 2021;22(4):1676. https://doi.org/10.3390/ijms22041676.
Li Y, Zhang L, Liu Z. Multi-objective de novo drug design with conditional graph generative model. J Cheminformatics. 2018;10:1–24. https://doi.org/10.1186/s13321-018-0287-6.
Putin E, Asadulaev A, Ivanenkov Y, Aladinskiy V, Sanchez-Lengeling B, Aspuru-Guzik A, et al. Reinforced adversarial neural computer for de novo molecular design. J Chem Inf Model. 2018;58(6):1194–204. https://doi.org/10.1021/acs.jcim.7b00690.
Blaschke T, Olivecrona M, Engkvist O, Bajorath J, Chen H. Application of generative autoencoder in de novo molecular design. Mol Inform. 2018;37(1–2):1700123. https://doi.org/10.1002/minf.201700123.
Heck GS, Pintro VO, Pereira RR, de Ávila MB, Levin NMB, de Azevedo WF. Supervised machine learning methods applied to predict ligand-binding affinity. Curr Med Chem. 2017;24(23):2459–70. https://doi.org/10.2174/0929867324666170623092503.
Parenti MD, Rastelli G. Advances and applications of binding affinity prediction methods in drug discovery. Biotechnol Adv. 2012;30(1):244–50. https://doi.org/10.1016/j.biotechadv.2011.08.003.
Showell GA, Mills JS. Chemistry challenges in lead optimization: silicon isosteres in drug discovery. Drug Discov Today. 2003;8(12):551–6. https://doi.org/10.1016/S1359-6446(03)02726-0.
Ferreira LL, Andricopulo AD. ADMET modeling approaches in drug discovery. Drug Discov Today. 2019;24(5):1157–65. https://doi.org/10.1016/j.drudis.2019.03.015.
Moroy G, Martiny VY, Vayer P, Villoutreix BO, Miteva MA. Toward in silico structure-based ADMET prediction in drug discovery. Drug Discov Today. 2012;17(1–2):44–55. https://doi.org/10.1016/j.drudis.2011.10.023.
Van De Waterbeemd H, Gifford E. ADMET in silico modelling: towards prediction paradise? Nat Rev Drug Discov. 2003;2(3):192–204. https://doi.org/10.1038/nrd1032.
Daoud NEH, Borah P, Deb PK, Venugopala KN, Hourani W, Alzweiri M, et al. ADMET profiling in drug discovery and development: perspectives of in silico, in vitro and integrated approaches. Curr Drug Metab. 2021;22(7):503–22. https://doi.org/10.2174/1389200222666210705122913.
Deng C, Liu J, Zhang W. Structural Modification in Anesthetic Drug Development for Prodrugs and Soft Drugs. Front Pharmacol. 2022;13:923353. https://doi.org/10.3389/fphar.2022.923353.
Danon JJ, Reekie TA, Kassiou M. Challenges and opportunities in central nervous system drug discovery. Trends Chem. 2019;1(6):612–24. https://doi.org/10.1016/j.trechm.2019.04.009.
Rankovic Z. CNS drug design: balancing physicochemical properties for optimal brain exposure. J Med Chem. 2015;58(6):2584–608. https://doi.org/10.1021/jm501535r.
Meibohm B, Derendorf H. Basic concepts of pharmacokinetic/pharmacodynamic (PK/PD) modelling. Int J Clin Pharmacol Ther. 1997;35(10):401–13.
Zou H, Banerjee P, Leung SSY, Yan X. Application of pharmacokinetic-pharmacodynamic modeling in drug delivery: development and challenges. Front Pharmacol. 2020;11:997. https://doi.org/10.3389/fphar.2020.00997.
Song S, Zhao W, Ji Y, Huang Q, Li Y, Chen S, et al. SHANK2 protein contributes to sevoflurane-induced developmental neurotoxicity and cognitive dysfunction in C57BL/6 male mice. APS. 2023;1(1):2. https://doi.org/10.1007/s44254-023-00005-7.
Shaked I, Oberhardt MA, Atias N, Sharan R, Ruppin E. Metabolic network prediction of drug side effects. Cell Syst. 2016;2(3):209–13. https://doi.org/10.1016/j.cels.2016.03.001.
Sachdev K, Gupta MK. A comprehensive review of computational techniques for the prediction of drug side effects. Drug Dev Res. 2020;81(6):650–70. https://doi.org/10.1002/ddr.21669.
Giacomini KM, Krauss RM, Roden DM, Eichelbaum M, Hayden MR, Nakamura Y. When good drugs go bad. Nature. 2007;446(7139):975–7. https://doi.org/10.1038/446975a.
Ji ZL, Wang Y, Yu L, Han LY, Zheng CJ, Chen YZ. In silico search of putative adverse drug reaction related proteins as a potential tool for facilitating drug adverse effect prediction. Toxicol Lett. 2006;164(2):104–12. https://doi.org/10.1016/j.toxlet.2005.11.017.
Zhao X, Chen L, Lu J. A similarity-based method for prediction of drug side effects with heterogeneous information. Math Biosci. 2018;306:136–44. https://doi.org/10.1016/j.mbs.2018.09.010.
Martin L, Hutchens M, Hawkins C, Radnov A. How much do clinical trials cost. Nat Rev Drug Discov. 2017;16(6):381–2.
Hassan Baig M, Ahmad K, Roy S, Mohammad Ashraf J, Adil M, Haris Siddiqui M, et al. Computer aided drug design: success and limitations. Curr Pharm Des. 2016;22(5):572–81.
Acharya C, Coop A, E Polli J, D MacKerell A. Recent advances in ligand-based drug design: relevance and utility of the conformationally sampled pharmacophore approach. Curr Comput Aided Drug Des. 2011;7(1):10–22. https://doi.org/10.2174/157340911793743547.
Ginalski K. Comparative modeling for protein structure prediction. Curr Opin Struct Biol. 2006;16(2):172–7. https://doi.org/10.1016/j.sbi.2006.02.003.
Martinelli D. Generative machine learning for de novo drug discovery: A systematic review. Comput Biol Med. 2022;145:105403. https://doi.org/10.1016/j.compbiomed.2022.105403.
Tripathi MK, Nath A, Singh TP, Ethayathulla A, Kaur P. Evolving scenario of big data and Artificial Intelligence (AI) in drug discovery. Mol Divers. 2021;25(3):1439–60. https://doi.org/10.1007/s11030-021-10256-w.
Gupta R, Srivastava D, Sahu M, Tiwari S, Ambasta RK, Kumar P. Artificial intelligence to deep learning: machine intelligence approach for drug discovery. Mol Divers. 2021;25(3):1315–60. https://doi.org/10.1007/s11030-021-10217-3.
Vatansever S, Schlessinger A, Wacker D, Kaniskan HÜ, Jin J, Zhou MM, et al. Artificial intelligence and machine learning-aided drug discovery in central nervous system diseases: State-of-the-arts and future directions. Med Res Rev. 2021;41(3):1427–73. https://doi.org/10.1002/med.21764.
Vamathevan J, Clark D, Czodrowski P, Dunham I, Ferran E, Lee G, et al. Applications of machine learning in drug discovery and development. Nat Rev Drug Discov. 2019;18(6):463–77. https://doi.org/10.1038/s41573-019-0024-5.
Dara S, Dhamercherla S, Jadav SS, Babu CM, Ahsan MJ. Machine learning in drug discovery: a review. Artif Intell Rev. 2022;55(3):1947–99. https://doi.org/10.1007/s10462-021-10058-4.
Deng J, Yang Z, Ojima I, Samaras D, Wang F. Artificial intelligence in drug discovery: applications and techniques. Brief Bioinform. 2022;23(1):bbab430. https://doi.org/10.1093/bib/bbab430.
Jiménez-Luna J, Grisoni F, Schneider G. Drug discovery with explainable artificial intelligence. Nat Mach Intell. 2020;2(10):573–84. https://doi.org/10.1038/s42256-020-00236-4.
Jiménez-Luna J, Grisoni F, Weskamp N, Schneider G. Artificial intelligence in drug discovery: Recent advances and future perspectives. Expert Opin Drug Discov. 2021;16(9):949–59. https://doi.org/10.1080/17460441.2021.1909567.
Hameroff SR. The entwined mysteries of anesthesia and consciousness: is there a common underlying mechanism? Anesthesiology. 2006;105(2):400–12. https://doi.org/10.1097/00000542-200608000-00024.
Hemmings HC, Akabas MH, Goldstein PA, Trudell JR, Orser BA, Harrison NL. Emerging molecular mechanisms of general anesthetic action. Trends Pharmacol Sci. 2005;26(10):503–10. https://doi.org/10.1016/j.tips.2005.08.006.
Son Y. Molecular mechanisms of general anesthesia. Korean J Anesthesiol. 2010;59(1):3–8. https://doi.org/10.1038/300487a0.
Mashour GA. Integrating the science of consciousness and anesthesia. Anesth Analg. 2006;103(4):975–82. https://doi.org/10.1213/01.ane.0000232442.69757.4a.
Askr H, Elgeldawi E, Aboul Ella H, Elshaier YA, Gomaa MM, Hassanien AE. Deep learning in drug discovery: an integrative review and future challenges. Artif Intell Rev. 2023;56(7):5975–6037. https://doi.org/10.1007/s10462-022-10306-1.
Acknowledgements
Not applicable.
Funding
Xianggen Liu was supported by the National Natural Science Foundation of China (No. 62206192) and the Natural Science Foundation of Sichuan Province (No. 2023NSFSC1408).
Author information
Authors and Affiliations
Contributions
Jiancheng Lv and Bowen Ke conceived and designed the study; Xianggen Liu and Zhe Xue wrote the paper; Mingmin Luo collected the data. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors gave their content for publication.
Competing interests
Dr. Bowen Ke is a member of the Editorial Board in Anesthesiology and Perioperative Science and recuses himself from every editorial procedure of this submission including peer-review and academic decisions. The author declares that he has no known competing financial interests or personal relationships that influenced the work reported here.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, X., Xue, Z., Luo, M. et al. Anesthetic drug discovery with computer-aided drug design and machine learning. APS 2, 7 (2024). https://doi.org/10.1007/s44254-023-00047-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44254-023-00047-x