Abstract
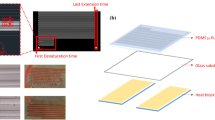
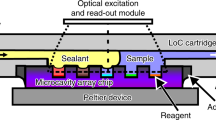
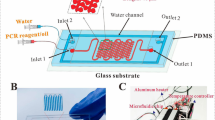
Digital PCR (dPCR) enables sensitive and precise quantification of template nucleic acid without calibration. However, dPCR is not yet in widespread use, probably due to the need for expensive specialized instruments. In this paper, we describe a dPCR system using a simple microfluidic chip and common laboratory tools. The microfluidic chip consists of two parts: a PDMS part with 24,840 × 0.25 nL microwells and a PDMS-coated flat glass plate. Human RNase P gene was adopted as the model template. Commercial products of human genomic DNA and real-time PCR reagents were mixed to make a PCR mixture. The PCR mixture was confined to the microwells by the PDMS degas-driven liquid control technique. The thermal cycling was performed on a common well-type thermal cycler with a minor modification. During the thermal cycling, evaporation of the PCR mixture was prevented with a handmade water holder. In the fluorescence image, bright (positive) microwells and dim (negative) ones were clearly discriminated. The number of the positive microwells was counted using software, and was used for estimation of the template concentration in the sample based on the theory of the Poisson distribution. The estimated concentrations well agreed with the input template concentrations in the range from 1.32 copies/µL to 13 200 copies/µL. The techniques presented in this paper will pave the way for facile dPCR in a broad range of laboratories without the need for expensive instruments.
Graphical Abstract







Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
S. Basu, SLAS Technol. (2017). https://doi.org/10.1177/2472630317705680
P.L. Quan, M. Sauzade, E. Brouzes, Sensors (2018). https://doi.org/10.3390/s18041271
C.M. Hindson, J.R. Chevillet, H.A. Briggs, E.N. Gallichotte, I.K. Ruf, B.J. Hindson, R.L. Vessella, M. Tewari, Nat. Methods (2013). https://doi.org/10.1038/nmeth.2633
B.J. Hindson, K.D. Ness, D.A. Masquelier, P. Belgrader, N.J. Heredia, A.J. Makarewicz, I.J. Bright, M.Y. Lucero, A.L. Hiddessen, T.C. Legler, T.K. Kitano, M.R. Hodel, J.F. Petersen, P.W. Wyatt, E.R. Steenblock, P.H. Shah, L.J. Bousse, C.B. Troup, J.C. Mellen, D.K. Wittmann, N.G. Erndt, T.H. Cauley, R.T. Koehler, A.P. So, S. Dube, K.A. Rose, L. Montesclaros, S.L. Wang, D.P. Stumbo, S.P. Hodges, S. Romine, F.P. Milanovich, H.E. White, J.F. Regan, G.A. Karlin-Neumann, C.M. Hindson, S. Saxonov, B.W. Colston, Anal. Chem. (2011). https://doi.org/10.1021/ac202028g
A.S. Whale, J.F. Huggett, S. Cowen, V. Speirs, J. Shaw, S. Ellison, C.A. Foy, D.J. Scott, Nucleic Acids Res. (2012). https://doi.org/10.1093/nar/gks203
E.A. Ottesen, J.W. Hong, S.R. Quake, J.R. Leadbetter, Science (2006). https://doi.org/10.1126/science.1131370
F. Shen, W.B. Du, J.E. Kreutz, A. Fok, R.F. Ismagilov, Lab Chip (2010). https://doi.org/10.1039/c004521g
N.R. Beer, B.J. Hindson, E.K. Wheeler, S.B. Hall, K.A. Rose, I.M. Kennedy, B.W. Colston, Anal. Chem. (2007). https://doi.org/10.1021/ac701809w
D. Pekin, Y. Skhiri, J. C. Baret, D. Le Corre, L. Mazutis, C. Ben Salem, F. Millot, A. El Harrak, J. B. Hutchison, J. W. Larson, D. R. Link, P. Laurent-Puig, A. D. Griffiths, V. Taly, Lab Chip (2011) https://doi.org/10.1039/c1lc20128j
K.A. Heyries, C. Tropini, M. VanInsberghe, C. Doolin, O.I. Petriv, A. Singhal, K. Leung, C.B. Hughesman, C.L. Hansen, Nat. Methods (2011). https://doi.org/10.1038/nmeth.1640
Y.F. Men, Y.S. Fu, Z.T. Chen, P.A. Sims, W.J. Greenleaf, Y.Y. Huang, Anal. Chem. (2012). https://doi.org/10.1021/ac300761n
A.M. Thompson, A. Gansen, A.L. Paguirigan, J.E. Kreutz, J.P. Radich, D.T. Chiu, Anal. Chem. (2014). https://doi.org/10.1021/ac5035924
Q.Y. Zhu, L. Qiu, B.W. Yu, Y.N. Xu, Y.B. Gao, T.T. Pan, Q.C. Tian, Q. Song, W. Jin, Q.H. Jin, Y. Mu, Lab Chip (2014). https://doi.org/10.1039/c3lc51327k
Q.C. Tian, Q. Song, Y.N. Xu, Q.Y. Zhu, B.W. Yu, W. Jin, Q.H. Jin, Y. Mu, Anal. Methods (2015). https://doi.org/10.1039/c4ay02604g
Y.Z. Wang, K.M. Southard, Y. Zeng, Analyst (2016). https://doi.org/10.1039/c6an00164e
Y.Y. Fu, H.B. Zhou, C.P. Jia, F.X. Jing, Q.H. Jin, J.L. Zhao, G. Li, Sens. Actuators B (2017). https://doi.org/10.1016/j.snb.2017.01.161
Q.Y. Zhu, Y.N. Xu, L. Qiu, C.C. Ma, B.W. Yu, Q. Song, W. Jin, Q.H. Jin, J.Y. Liu, Y. Mu, Lab Chip (2017). https://doi.org/10.1039/c7lc00267j
C.D. Ahrberg, J.M. Lee, B.G. Chung, Biochip J. (2019). https://doi.org/10.1007/s13206-019-3302-8
X. Zhou, G.C. Ravichandran, P. Zhang, Y. Yang, Y. Zeng, Lab Chip (2019). https://doi.org/10.1039/c9lc00932a
X. Cui, L. Wu, Y. Wu, J.H. Zhang, Q. Zhao, F.X. Jing, L. Yi, G. Li, Anal. Chim. Acta (2020). https://doi.org/10.1016/j.aca.2020.02.010
H.Q. Si, G.W. Xu, F.X. Jing, P. Sun, D. Zhao, D.P. Wu, Sens. Actuators B. (2020). https://doi.org/10.1016/j.snb.2020.128197
G.W. Xu, H.Q. Si, F.X. Jing, P. Sun, D. Zhao, D.P. Wu, Micromachines (2020). https://doi.org/10.3390/mi11121025
G.W. Xu, H.Q. Si, F.X. Jing, P. Sun, D.P. Wu, Biosens (2021). https://doi.org/10.3390/bios11050158
J.M. Hu, L.B. Chen, P.F. Zhang, K.W. Hsieh, H. Li, S. Yang, T.H. Wang, Lab Chip (2021). https://doi.org/10.1039/d1lc00636c
T.B. Xie, P. Wang, L. Wu, B.Y. Sun, Q. Zhao, G. Li, Lab Chip (2021). https://doi.org/10.1039/d1lc00448d
C.Y. Sung, C.C. Huang, Y.S. Chen, K.F. Hsu, G.B. Lee, Lab Chip (2021). https://doi.org/10.1039/d1lc00663k
T.C. Merkel, V.I. Bondar, K. Nagai, B.D. Freeman, I. Pinnau, J. Polym. Sci. Part B: Polym. Phys. (2000). https://doi.org/10.1002/(SICI)1099-0488(20000201)38:3%3c415::AID-POLB8%3e3.0.CO;2-Z
K. Hosokawa, K. Sato, N. Ichikawa, M. Maeda, Lab Chip (2004). https://doi.org/10.1039/B403930K
T. Ito, A. Inoue, K. Sato, K. Hosokawa, M. Maeda, Anal. Chem. (2005). https://doi.org/10.1021/ac050122f
K. Hosokawa, M. Omata, K. Sato, M. Maeda, Lab Chip (2006). https://doi.org/10.1039/B513424B
K. Hosokawa, M. Omata, M. Maeda, Anal. Chem. (2007). https://doi.org/10.1021/ac070659o
L.F. Xu, H. Lee, D. Jetta, K.W. Oh, Lab Chip (2015). https://doi.org/10.1039/c5lc00716j
K. Hosokawa, Anal. Sci. (2021). https://doi.org/10.2116/analsci.20SCR04
J.N. Lee, C. Park, G.M. Whitesides, Anal. Chem. (2003). https://doi.org/10.1021/ac0346712
M. Baer, T.W. Nilsen, C. Costigan, S. Altman, Nucleic Acids Res. (1990). https://doi.org/10.1093/nar/18.1.97
G.L. Long, J.D. Winefordner, Anal. Chem. (1983). https://doi.org/10.1021/ac00258a001
S.K. Wee, S.P. Sivalingam, E.P.H. Yap, Genes (2020). https://doi.org/10.3390/genes11060664
G. Choi, T.J. Moehling, R.J. Meagher, Expert Rev. Mol. Diagn. (2023). https://doi.org/10.1080/14737159.2023.2169071
Acknowledgements
This work was partly supported by Japan Society for the Promotion of Science (JSPS) KAKENHI Grant Number 18K04917.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hosokawa, K., Ohmori, H. Digital PCR using a simple PDMS microfluidic chip and standard laboratory equipment. ANAL. SCI. 39, 2067–2074 (2023). https://doi.org/10.1007/s44211-023-00425-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s44211-023-00425-2