Abstract
High-elevation insectivorous birds are currently confronted with the reality of a changing climate, land use shifts, and the decline of many prey groups. The diet dynamics among many imperiled animals in this group are still unresolved. Examining the diets of tree-line Passerine birds to the species level of the prey allows for stronger population predictions. This study uses DNA metabarcoding to identify prey arthropods from adult Passerine bird feces at and slightly below tree-line in a Pyrenean forest. Our objective was to quantify the intra-and inter-species richness and overlap of Passerine bird diet over time and space. The results showed that adult Passerine diets have high inter- and intra-species dietary variability and low inter- and intra-species dietary overlap. The lack of association between dietary richness and open space, season, and elevation and lack of differences between dietary overlap and open space and elevation suggest high-elevation Passerine birds have very high dietary flexibility. The results also showed that aphids known to be pests to conifers, and other conifer pests, were prevalent in the birds’ diets. The Passerine diets and high rate of rare dietary items are mainly in line with other recent DNA metabarcoding studies. Implications for the long-term projections relative to tree-line Passerine populations are discussed.
Similar content being viewed by others
Introduction
Insect-eating birds across the globe consume 400–500 million tons of insects annually (Nyffeler et al., 2018), surpassing the 350 million tons of meat that humans consume each year (Hicks et al., 2018). However, insectivorous bird populations are particularly at risk due to climate change. In North America, some terrestrial insectivorous bird populations have decreased by 33% in the past five decades (Tallamy & Shriver, 2021). Similarly, many insectivorous and non-insectivorous bird populations have faced a substantial decline over the past 30 years in Europe (Inger et al., 2015).
Upper elevation birds are particularly at risk because mountains are expected to be more affected by climate change than lowland areas due to faster and enhanced warming (Mountain Research Initiative EDW Working Group, 2015). For example, high-elevation populations of Canada jays (Perisoreus canadensis (L., 1766)) declined 50% over a 30-year period, and the decline was attributed to warmer and more variable weather (Sutton et al., 2021). This change in weather pattern increased the number of freeze–thaw events, which caused an increase in the spoilage of cached food items. The survival of another high-elevation Passerine, the white winged snowfinch (Montifringilla nivalis (L., 1766)) is in doubt because its foraging behavior is closely tied to snow retreat conditions which are becoming increasingly less consistent (Resano-Mayor et al., 2019). Finally, Barras et al. (2021) found that elevated ambient temperatures at the tree-line in the Swiss alps negatively affected nestling prey provisioning rates of the Alpine ring ouzel (Turdus torquatus alpestris L., 1758).
The flora and fauna of the Pyrenees mountains are especially threatened due to climate change and land use shifts (OPCC-CTP, 2018). The snowpack is warmer than many other mountain ranges across the world and thus particularly sensitive to slight changes in ambient temperature (Lopez-Moreno et al., 2017), and the decline of agropastoral practices in the Pyrenees has led to transitions of open grassland into forest (Roura et al., 2005). The Pyrenean tree-line is also shifting upward (Ameztegui et al., 2016). Tree-line dynamics globally are affected by a variety of factors, including precipitation, tree community composition, and soil structure (Grace et al., 2002; Körner, 2012). The upward shift and densification of the tree-line in the Pyrenees is generally linked to local agricultural abandonment (Batllori & Gutiérrez, 2008), but there can be locally important factors such as slope morphometry and lithology (Feuillet et al., 2020).
It is within this context that we examined the diet of Passerines at elevations located below and at tree-line. The diet of many European Passerine birds, e.g., Paridae, has been examined closely, even though most studies were limited to estimating the diet of nestlings using either methods that are invasive (neck collars (Barba & Gil-Delgado, 1990; Pagani-Núñez et al., 2011) and stomach flushing (Senécal et al., 2021)), noninvasive but less detailed (e.g., cameras and stable isotope analysis (Currie et al., 1996; Şekercioğlu et al., 2023, respectively)), or lethal (gizzard extraction (Sehhatisabet et al., 2008)). Recent advances in DNA metabarcoding technology (hereafter metabarcoding) have increased our ability to analyze the diet of adult Passerines in a noninvasive manner at a high level of taxonomic classification (see Crisol-Martínez et al., 2016; Ribeiro et al., 2019; da Silva et al., 2020; Shutt et al., 2020).
Metabarcoding supports high-throughput (i.e., massively parallel) taxonomic classification within a sample (Bush et al., 2019). A short portion of a gene (barcode) from an environmental or biological sample is amplified by a primer designed to provide taxonomic resolution of a target organism or taxonomic group (Deagle et al., 2014; Hajibabaei et al., 2007). However, there are many fundamental limitations of fecal metabarcoding. For example, raw prey abundances cannot be determined from the number of reads in a similar DNA sequence, and relative prey abundance is difficult to recover because of technological and biological biases including primer mismatch and differences in PCR amplification due to primer sequence length (Deagle et al., 2013; Krehenwinkel et al., 2017; Piñol et al., 2015). Sample contamination and differing rates of DNA preservation in the gut can also present issues (Galan et al., 2018; Nielsen et al., 2018).
A handful of studies have examined the diet of multiple adult Passerines species using metabarcoding (see Crisol-Martínez et al., 2016, Sottas et al., 2020, and Garfinkel et al., 2022). Most Passerine dietary metabarcoding studies have examined the adult diet of one species (McClenaghan et al., 2019; Moran et al., 2019; Shutt et al., 2020, and Snider et al., 2021 among others) or that of nestlings (see Rytkönen et al., 2019, da Silva et al., 2020, and Jarrett et al., 2020). The goal of our study was to examine the diets of co-occurring high-elevation Pyrenean Passerines and to assess the effects of open space, elevation, and season on them. We expected higher niche differentiation (i.e., difference in diet composition) in morphologically and behaviorally similar species and higher dietary richness and higher dietary overlap among species as spring progressed to autumn. Passerines that have similar traits often have competition-driven niche separation (Alatalo et al., 1986; Cowie & Hinsley, 1988; Sottas et al., 2020), and higher abundance of prey is linked to less dietary partitioning (Davies et al., 2022). In some species, we expected diet richness to positively correlate with the percentage of open space because patchier habitats have been shown to benefit some species but not others (Suarez-Seoane et al., 2002). Finally, we expected higher overlap in below tree-line plots because we expected the conditions to be more favorable to Passerines. Higher overlap is common in more favorable habitats (Hou et al., 2021).
Methods
Study area and feces collection
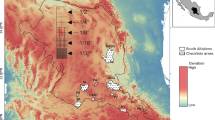
Ten plots were selected in grid format within a black pine forest (Pinus mugo species complex, hereafter P. mugo) in Vall d’Ordino, a valley located within three km of Vall de Sorteny Natural Park in the parish of Ordino, Andorra. Plots were situated between 1729 and 2352 m above sea level (hereafter MASL). Percent open space surrounding each plot (1000 m radius) was calculated using QGIS3.4 and the MCSA 2012 landcover map downloaded from the Institute of Andorran Studies (Centre de Biodiversitat de l'Institut d'Estudis Andorrans, 2012). Testing various scales of open space was out of the scope of this study; therefore, a landscape level scale of 1000 m radius was chosen as a 1000 m radius is a generally accepted radius for a landscape level measure of “forest amount” (Shoffner et al., 2018). Plots were characterized as “below tree-line” or “at tree-line” depending on positioning above or below the median elevation of all plots (i.e., 2077 MASL). In most of Andorra, tree-line occurs between 2200 and 2400 MASL and in a few areas between 2100 and 2500 MASL (Carreras et al., 1996).
Birds were captured using Ecotone mist-nets (9 m and 6 m long and 2.5 m high, with 5 shelves and a mesh size of 16 mm2) stretched between 4 m poles inserted perpendicularly in the ground. Three nets were used in eight of the sites, and two nets were used in the remaining two sites due to difficultly of access, and openness of site (see Supplementary material 1). Mist nets were deployed between May 15, 2018, and September 30, 2018, and number of net days for each site are also documented in Supplementary material 1. The start date for the field component of this study coincided with the date when snow historically has retreated from the Andorran tree-line. Mist nets were not set at a plot when there was precipitation or high winds to ensure good capture conditions and welfare of the bird. Once a bird was captured, it was placed in a single use individual paper bag. After defecation, feces were carefully removed from the paper bag using a single-use toothpick and stored in plastic vials. Vials were placed on ice in the field and transferred to long-term deep-freeze as soon as possible. All birds caught were identified at species level, ringed, aged, sexed, and measured following standard ringing procedure. Birds were handled by certified ringers and all the procedures approved by the Environment and Sustainability Department of Andorran Government.
A total of 132 fecal samples were collected. No bird was captured twice. Samples collected in May and June were considered to be from the spring, July and August samples were considered to be from the summer, and September samples were considered to be from the autumn.
DNA extraction and amplification
DNA in the collected fecal samples and in four DNA extraction blanks (i.e., vials with no fecal samples that served as contaminant control) was extracted using the QIAamp DNA Stool Mini Kit (QIAGEN) following the manufacture’s protocol with modifications suggested by Davies et al. (2022). Only samples of similar size (approximately 3 mg) were processed. Three samples were smaller than 3 mg and thus discarded from analysis. DNA concentration was quantified using a Qubit 3.0 Fluorometer (Thermo Fisher Scientific Inc.). Amplicon library preparation and sequencing were carried out by the Georgia Genomics and Bioinformatics Core (University of Georgia, Athens GA, USA) as follows: the concentration and size distribution of the DNA were determined using SYBR fluorometry and a fragment analyzer, respectively. The DNA samples were normalized to the same concentration before library preparation. The amplicon library was prepared beginning with a 2X KAPA HiFi HotStart ReadyMix (Roche Sequencing & Life Sciences). The first PCR was assembled with target-specific primers. Initial plans called for the use of a primer amplifying a longer region, but a preliminary experiment (data not shown) indicated higher efficacy of a shorter primer, the mini-barcode mitochondrial primer (ANML). ANML amplifies a smaller 180 bp segment on the cytochrome oxidase C subunit 1 (COI) (Jusino et al., 2017). In the second round of PCR, unique Illumina Indexed primers (i5 and i7) were added to each sample and at the end of each round of PCR, amplicon was purified with bead cleanup. Final libraries underwent quality checks by SYBR fluorometry, Fragment Analyzer and qPCR (KAPA Library Quantification—Roche Sequencing & Life Sciences). Libraries were normalized and pooled equally. The pooled amplicons were sequenced on the MiSeq platform (PE250) (Illumina).
Bioinformatic analysis
Within the QIIME 2 2020.6 environment, tagged feces sequence reads generated from the Illumina MiSeq sequencer were demultiplexed and primers trimmed to create fastq files (Bolyen et al., 2019). We then used the DADA2 pipeline for further downstream analysis, which created a table of amplicon sequence variants (ASVs) rather than traditional operational taxonomic units (OTUs), thereby improving reproducibility, comprehensiveness, and accuracy (Callahan et al., 2016). Potential contaminants in the ASV table using the “frequency” method were identified by the package Decontam using the default threshold of 0.1 (Davis et al., 2018). The “frequency” method identifies contaminants whose concentrations in blank samples are inversely correlated with sample DNA concentrations, based on a user-specified threshold. As the metabarcoding workflow introduces quantitative bias into results, ASV raw counts were transformed into a presence/absence matrix (Martoni et al., 2022).
ASVs were taxonomically classified by aligning sequences to those within the arthropod training database “tidybug” (O’Rourke et al., 2020) via the classy-sklearn naïve Bayes method implemented in QIIME 2’s q2-feature-classifier plugin. The full QIIME script can be found in Supplementary material 2. Each taxonomic assignment was further examined individually using the following protocol: (1) The geographic range of the assignment was assessed, and species that do not occur in Europe were removed. (2) Species considered to be rare in Europe but are not known to occur in the Pyrenees were flagged. (3) Flagged assignments were verified by submitting query sequences to the NCBI BLAST tool and assignments that did not score at or above 98% identity were removed. All single flagged species composed of multiple ASV sequences were aligned to check for sequencing error, and sequences above an 80% sequencing error were kept in the analysis (Brandt et al., 2021; Ritter et al., 2022). (4) If an ASV showed multiple hits with the same max score on the NCBI BLAST tool, the ASV was removed from the analysis.
Statistical analyses
A Wilcoxon–Mann–Whitney test was performed to test for correlations between open space and elevation as a discrete variable, using the percent of open space as a dependent variable. We calculated mean prey richness of all bird species and of the top two commonly collected bird species (Periparus ater (L., 1758) and Lophophanes cristatus (L., 1758)). Predictive roles of independent factors affecting of the combined prey richness of these two bird species (season, elevation as a continuous gradient, and open space) were found by fitting data to a negative binomial or Poisson model (GLMM) using the lme4 v.26 package (Bates et al., 2015) in R Version 1.3.1056 (R Core Team, 2021). Models were chosen through a combination of (1) residual plotting with the DHARMa package (Hartig, 2022); (2) model performance testing using Pearson, Kendall, and Spearman correlation coefficients; and (3) model accuracy evaluation by measuring the root mean square error and the mean-absolute deviation of each model. Plot was used as a random variable. We also tested “individual bird” as the only random factor and it yielded very similar results, quantitatively and qualitatively. Nevertheless, the “individual bird” choice also caused some models to fail convergence and these problems were more severe when both “plot” and “individual” were chosen together as a random factor. Therefore, our final choice for random factor was “plot.” Some models, however, did not accept any random factor and were thus run without one after numerical testing showed little differences between models with random factors and those without. After the prey richness model of each bird group was fit, post hoc Tukey tests were carried out to investigate error rates of the categorical factor of season. Comparison of beta diversity (i.e., dissimilarity of diet) among and within bird species was determined by a Jaccard dissimilarity matrix using the R vegan package, with a value of 1 indicating there were no shared species and a value of 0 indicating complete sharing of species (Oksanen et al., 2020). A Jaccard dissimilarity matrix was also used to examine the dissimilarity of diet of all individual birds combined and the individuals of the two most common bird species combined in regard to season and the discrete variable of elevation. Frequency of occurrence (number of times a prey item appeared in a fecal sample, divided by total number of samples) was calculated for the six most frequently occurring prey groups (Arachnida, Diptera, Coleoptera, Hemiptera, Hymenoptera, and Lepidoptera). Predictive roles affecting species richness within each prey group (season, elevation as a continuous gradient, and open space) were calculated using the same model selection process as bird groups. Sample coverage was examined using the iNEXT package to create sample size-based rarefaction curves and extrapolation curves (Chao & Jost, 2012; Chao et al., 2014).
Results
A total of 8.95 million sequence reads were produced in the feces samples, with ASV counts per feces sample ranging from four to 268,999 (Supplementary material 3). ASVs were taxonomically classified as the MOTUs of 713 arthropod classifications (representing six classes, 30 orders, 163 families, 466 genera) and then only MOTUs classified to genus or species were kept for a total of 590 arthropod MOTUs. We refer to these MOTUs as species.
Inter- and intra-species dietary richness of Passerines
Fecal samples were collected and analyzed from 14 bird species (Supplementary material 4). The bird species P. ater (coal tit), L. cristatus (crested tit), and Prunella modularis (L., 1758) (dunnock) accounted for 40%, 15%, and 14% (respectively) of the bird species from which samples were collected. No significant seasonal differences were found when all bird species were combined or when just P. ater and L. cristatus were combined (P values in Supplementary material 5). No captures of P. modularis were made in autumn. Seventy-three and 59 birds were caught in the high-elevation and low-elevation plots, respectively.
The mean prey richness per bird capture was 13.5 ± 6.9 species. Phoenicurus ochruros (Gmelin, 1774) (black redstart) displayed the highest species richness per bird capture, followed by Sylvia atricapilla (L., 1758) (blackcap) and P. modularis, although standard deviation bars do not show differences within these three species (Fig. 1).
According to Wilcoxon–Mann–Whitney tests, the percent of open space was significantly higher in the plots at tree-line (mean: 88 ± 13.7) than the plots below tree-line (mean: 45.2 ± 17.5) (Z = 4.25, P < 0.001). However, GLMM results showed that the percentage of open space was not correlated with prey richness in the diets of P. ater and L. cristatus when combined, or when all bird species were combined (Beta estimates and P values in Supplementary material 5). There were no elevational differences in prey richness when all bird species were combined, or when P. ater and L. cristatus were combined (Supplementary material 5).
Inter- and intra-species dietary overlap of Passerines
Jaccard dissimilarity index showed very little overlap in the diet between and within bird species, and there was a mean dissimilarity of 0.91 ± 0.05 in the diet of the 14 bird species. Beta diversity was very high among species. Compared to each other, P. modularis /L. cristatus and P. modularis /P. ater had a higher rate of dissimilarity (0.86 and 0.83, respectively) than the rate between P. ater/ L. cristatus (0.73). The dietary variability within P. ater, L. cristatus, and P. modularis was high as well: P. ater (0.82 ± 0.02), L. cristatus (0.93 ± 0.05), and P. modularis (0.95 ± 0.04). Accordingly, sample size-based rarefaction curves indicated it would be necessary to capture over 100 more P. ater individuals than P. modularis and L. cristatus to reach 99% sample coverage (Fig. 2a). At 99% coverage, P. modularis is expected to have a higher diversity of diet than P. ater and P. cristatus, while P. ater is expected to have the lowest (Fig. 2b). The mean overlap among all individual birds captured was not different between the three seasons (spring, 0.9 ± 0.05; summer, 0.93 ± 0.07; autumn, 0.93 ± 0.06) (Supplementary material 6a). Similarly, when the mean overlap between the two most captured birds (P. ater and L. cristatus) was calculated by season, no difference in overlap was recorded (spring, 0.92 ± 0.07; summer, 0.91 ± 0.08; autumn, 0.90 ± 0.07) (Supplementary material 6a). The mean overlap between all birds and between P. ater and P. cristatus in the plots at tree-line and below tree-line was similarly low (see Supplementary material 6b).
Sample size-based rarefaction curves (solid line) and extrapolation curves (dotted line) with 95% confidence intervals. a Number of birds caught per sample coverage. Numbers in parentheses indicate bird species and number of individual birds caught at 99% sample coverage. To reach 99% coverage, it would be necessary to capture over 100 more P. ater individuals than P. modularis and L. cristatus. b Prey diversity of bird species per number of individual birds sampled. Numbers in parentheses indicate number of captures of birds per species necessary to reach 99% coverage, and prey diversity at 99% coverage with 95% confidence intervals. Prunella modularis is expected to have a higher diversity of diet at 99% sample coverage, while P. ater is expected to have lowest diversity
Presence of prey species and prey species trends
Most prey species were rare; 58.64% of the prey species were collected only once (i.e., collected in one sample) (Table 1). However, eleven species were present in over 15% of samples (Supplementary material 7). Of these eleven species, five were conifer pests. Aphid conifer pests (Hemiptera) were the two species most likely to be present (Supplementary material 7). Diptera and Lepidoptera represented 21.78% and 20.11%, respectively, of all prey species (Fig. 3). GLMM results indicated that the richness of some groups of prey (Coleoptera, Hemiptera, and Hymenoptera) significantly varied between some seasons, and elevation and open space did not drive richness of prey groups (Beta estimates, P values, and seasonality Tukey tests in Supplementary material 8).
Discussion
The difficulty in accurately and directly identifying species-level dietary components of adult insectivorous Passerines in a non-lethal manner is a quandary that has long confounded ornithologists. A metabarcoding approach allows for the direct study of species composition of Passerine feces. With this technique, we were able to determine that: (1) there was extremely high inter- and intra-species variability and low inter- and intra-species overlap in the diet of captured Passerines, a result that contrasts with some traditional Passerine nestling studies and agrees with many metabarcoding studies; (2) dietary richness did not correlate with season nor with open space, and there was no difference in dietary overlap relative to open space or elevation, which suggests that high-elevation Passerine birds in our study have high dietary mobility; and (3) Passerine diets were dominated by conifer pests.
There was high biological richness in our analysis of the 132 fecal samples: over 594 taxonomically classified arthropod species were identified. Most prey species were rare; 58.64% of the prey species (or 346) were recorded in only one feces collection. A recent metabarcoding study examining diets of 25 North American grassland Passerines also found very diverse diets and rare prey items among bird species and linked this inter-species dietary diversity to opportunistic availability (Garfinkel et al., 2022). Similarly, authors of other metabarcoding studies examining (respectively) Eurasian blue tits (Cyanistes caeruleus (Shutt et al., 2020)), Rufous hummingbirds (Selasphorus rufus (Gmelin 1788)) (Moran et al., 2019), and barn swallows (Hirundo rustica (L. 1758)) (McClenaghan et al., 2019) postulated that the high intraspecific dietary variation found in the birds’ diets was likely due to prey availability and dietary flexibility.
Despite a mean richness per bird capture of only 13.5 ± 6.9 species, the dietary overlap among and within bird species was very low. We expected lower overlap (higher niche differentiation) between closely related bird species, as it is well established that segregated foraging behavior occurs between closely related European insectivorous Passerines. When a potential niche is left unoccupied by a Passerine bird species, the species that most resembles the absent species in body size is the species most likely to fill it (for a review of geographic niche changes in insectivorous Passerines, see Alatalo et al. (1986)). Segregated foraging behavior makes sense in light of a study showing negative impacts upon a less dominant but closely related species sharing geographic space; Parus major (L. 1758) (great tit) nestlings raised sympatrically with C. caeruleus weighed less than those raised allopatrically, suggesting that a large overlap of resource utilization exists between the two closely related species (Torok & Tóth, 1999). Most dietary studies comparing insectivorous hole-nesting Passerines (mainly some combination of P. ater, P. major, L. cristatus, and C. caeruleus) have historically examined nestling diets and reported a high overlap when dietary components are classified to a combination of class and family (Grzędzicka, 2018; Michalski et al., 2011; Nour et al., 1998). One study that classified Passerine prey of Lepidoptera and Arachnida to species also found high overlap (Atiénzar et al., 2013). However, at least two studies have shown that P. major and C. caeruleus feed differing sizes of caterpillar prey to nestlings (Ceia et al., 2016; Torok & Tóth, 1999), a result that would not be observable in a DNA-based study such as ours. In Ceia et al. (2016) the authors showed that the composition of prey (classified to family) between the two bird species were similar, and they postulated that difference in prey size resulted from either interspecific competition between P. major and C. caeruleus or the segregation of bird foraging guilds; Cyanistes caeruleus is primarily a foliage-gleaner and more likely to come in contact with smaller instar caterpillars, while P. major are bark–foliage gleaners and therefore more likely to come in contact with later instar caterpillars. Numerous dietary metabarcoding studies, however, have found lower inter- and intra-species overlap, including a dietary metabarcoding study of insectivorous Malaysian Babbler species (genera within the families Pellorneidae and Timaliidae) which found lower than 35% overlap in three of the ten species (Mansor et al., 2022). Lower inter- and intra-species overlap was also found in two other metabarcoding studies that examined dietary overlap within European insectivorous Passerines species (see Rytkönen et al., 2019; Shutt et al., 2020).
While inter- and intra-species overlap was very low in our study, the Jaccard dissimilarity indexes displayed slightly higher index values between P. modularis/L. cristatus and P. modularis/P. ater than between the more closely related P. ater and L. cristatus. These results are likely due to diverging foraging habits, a finding supported by a metabarcoding study that found differing dietary compositions of ground- vs arboreal-foraging bird species in macadamia orchards (Crisol-Martínez et al., 2016). Periparus ater and L. cristatus both forage in trees (Alatalo, 1981; Hartley, 1987; Lens, 1996), while P. modularis are mainly ground feeders (Bishton, 1986).
Historical data report divergent timing of clutch-laying of closely related insectivorous hole-nesting Passerines (Sanz et al., 2010), and a more recent study reported that many Passerine nestlings are provisioned with differing prey types depending on the nestling’s development stage (Orłowski et al., 2017). Historical data also suggest that resident insectivorous Passerines in many temperate forests are less segregated in both foraging sites and dietary components in summer, when insect prey is more abundant. Insect prey in autumn and winter is less abundant, leading to resource partitioning and inter-species competition (Betts, 1955; Gibb, 1954; Lister, 1980). There has been disagreement over seasonal segregation and diet. For example, Ulfstrand (1977) found spatial segregation in summer compared to autumn, whereas Wagner (1981) and Almeida and Granadeiro (2000) found no significant seasonal spatial differences. Obeso (1987) found no spatial difference but did find significantly different dietary components. Finally, a recent study found high dietary overlap between communities of insectivorous Passerines during times of limited insect availability, in contrast to many studies that show high dietary overlap during times of high resource availability. The researchers postulated this result indicated that during times of very low food availability, bird species were unable to avoid competition (Kent et al., 2022). Our study found no seasonal differences in dietary overlap between the two most commonly captured birds (P. ater and L. cristatus).
Our results also did not show seasonal dietary richness differences of these two bird species combined or when dietary richness of all individual birds captured were combined (Supplementary material 5). Our seasonal richness results are different from the findings of an adult metabarcoding study done in a deciduous forest (Shutt et al., 2020). In that study, Shutt et al. (2020) tested 793 fecal samples collected from March through early May of C. caeruleus and linked rising seasonal dietary richness to rising herbivorous insect abundance and availability. It is possible that the lack of differences in seasonal dietary richness found in our study is a result of our sampling time frame (mid-May through late September) and/or the relatively lower number of samples taken in our study (132). Regardless, our results (lack of seasonal prey overlap among bird species, no difference in dietary richness from spring to autumn, and no difference in richness levels among all bird species) likely indicate that the birds captured in our study have a high level of dietary flexibility.
We expected dietary richness levels of ground and shrub foraging species, such as P. modularis, to be correlated with open space, as structure and composition of vegetation can be very influential in nestling success of some European Passerine species (Orłowski et al., 2017). We also expected higher dietary overlap in plots below tree-line; while response of insects to elevation is species specific (Hodkinson, 2005), many montane fauna either decrease with elevation or have a humped shaped distribution along an elevational gradient (McCain, 2009; Rahbek, 2005). Temperature is known to be a major driver in insect community structure (Bale et al., 2002), and temperature swings are wider at higher elevations in the Pyrenees (Navarro-Serrano et al., 2020). Therefore, insects are likely more abundant in plots below tree-line, and thus, insectivorous Passerines might be less segregated at these lower elevations. However, in our study percent of open space and/or elevation had no effect upon the richness of the bird diets, or when the most common birds were examined separately. While more birds were caught in plots below tree-line than in plots at tree-line, the diet composition within both these groups showed low overlap, i.e., the diet among birds in plots at tree-line had as much overlap as the diet of birds in plots below tree-line. More data would be needed to document and compare the diet of each 14 bird species we studied, but this lack of link between open space and elevation may indicate high mobility of the more common generalists that occupy high-elevation Pyrenean landscapes.
Our adult Passerine barcoding study was performed in P. mugo forests, so it is unsurprising that five of the eleven species most likely to be present in the bird feces from this habitat were conifer pests. The two species most likely to be present in the bird feces were conifer aphids; 53.03% of the samples contained Cinara pini (L., 1758) and 37.88% contained Eulachnus rileyi (Williams, 1911) (Supplementary material 6). Cinara pini is a common and native conifer pest in Europe. Eulachnus riley, however, is considered rare to uncommon in its native range in Europe and is considered a pest outside its native range (Blackman & Eastop, 1994), so it is interesting to find this species to be common in our study. Even though, as a group, the European Paridae are some of most intensely studied birds in the world (Betts, 1955; Cowie & Hinsley, 1988; Gibb, 1954; Lack, 1964; Ulfstrand, 1976), data are limited relative to their adult diets. However, the abundance of aphids in adult diets in our study is similar to two other studies: Shutt et al. (2020) reported aphids comprised three of the top ten prey taxa and had the highest presence incidences, and Betts (1955) found aphids comprised over 50% of the adult diet of three species of hole-nesting European Paridae in June. Birds play important roles in top-down control of forest arthropod populations (Fayt et al., 2005; Gunnarsson, 1995, 1996; Philpott et al., 2004; Schwenk et al., 2010). It is possible the Passerines in our study are shaping arthropod communities and causing a trophic cascade by affecting tree growth. Research examining trophic cascade affects by bird predation, however, have revealed complex interactions or mixed results (Gruner, 2004; Schwenk et al., 2010).
Regardless, we now know many high-elevation species are under pressure (Öztürk et al., 2015), and alpine-breeding Passerines such as Anthus spinoletta (L., 1758) (water pipit) are precipitously declining in some areas (Flousek et al., 2015). We caught only one specimen of A. spinoletta; the remainder of our species are elevational generalists and not confined to high elevation. While our data does not shed light on alpine specialists, the lack of differences in dietary overlap and diversity relative to open space and elevation, not to mention the extremely low levels of intra-species dietary overlap, suggest that adult diet may not be a constraining factor in populational growth of some generalist insectivorous Passerines in and around the Pyrenean tree-line. At least one European generalist insectivorous Passerine bird seems to display extreme plasticity in timing of egg-laying (Wesołowski et al., 2016), and other ecological requirements such as suitable nesting sites and the provisioning needs for nestling could be more plausible population constraints. The upward migration of the tree-line in the Pyrenees, a phenomenon likely caused by both land-use shifts and climate change (Batllori & Gutiérrez, 2008; Batllori et al., 2010), may therefore not be a significant factor affecting the diet of some adult generalist insectivorous Passerines.
Much remains to be discovered regarding the diets of adult European insectivorous Passerines (Cholewa & Wesołowski, 2011). In the future, metabarcoding will undoubtably continue to elucidate the relationship between birds, insects, and landscape and has the potential to reveal vast quantities of dietary data. Our results showed very high prey diversity and very little overlap within and among insectivorous Passerines. Spatial trends (open space and elevation) had little effect on prey diversity and overlap. While these data indicate that the dietary plasticity of the more common birds is high, more studies are needed to reveal dietary components of mountain species, such as A. spinoletta.
Data availability
The raw data present at this link: https://doi.org/10.5281/zenodo.6968770.
References
Alatalo, R. V. (1981). Interspecific competition in tits: Parus spp. and the goldcrest Regulus regulus: Foraging shifts in multispecific flocks. Oikos, 37(3), 335–344.
Alatalo, R. V., Gustafsson, L., & Lundberg, A. (1986). Interspecific competition and niche changes in tits (Parus spp.): Evaluation of nonexperimental data. The American Naturalist, 127(6), 819–834.
Almeida, J., & Granadeiro, J. (2000). Seasonal variation of foraging niches in a guild of passerine birds in a cork-oak woodland. Ardea, 88, 243–252.
Ameztegui, A., Coll, L., Brotons, L., & Ninot, J. M. (2016). Land-use legacies rather than climate change are driving the recent upward shift of the mountain tree line in the Pyrenees. Global Ecology and Biogeography, 25(3), 263–273.
Atiénzar, F., Belda Eduardo, J., & Barba, E. (2013). Coexistence of mediterranean tits: A multidimensional approach. Écoscience, 20(1), 40–47.
Bale, J. S., Masters, G. J., Hodkinson, I. D., Awmack, C., Bezemer, T. M., Brown, V. K., Butterfield, J., Buse, A., Coulson, J. C., Farrar, J., & Good, J. E. (2002). Herbivory in global climate change research: Direct effects of rising temperature on insect herbivores. Global Change Biology, 8(1), 1–16.
Barba, E., & Gil-Delgado, J. (1990). Seasonal variation in nestling diet of the great tit Parus major in orange groves in eastern Spain. Ornis Scandinavica, 21, 296–298.
Barras, A. G., Niffenegger, C. A., Candolfi, I., Hunziker, Y. A., & Arlettaz, R. (2021). Nestling diet and parental food provisioning in a declining mountain passerine reveal high sensitivity to climate change. Journal of Avian Biology, 52(2). https://doi.org/10.1111/jav.02649
Bates, D., Mächler, M., Bolker, B., & Walker, S. (2015). Fitting linear mixed-effects models using lme4. Journal of Statistical Software, 67(1), 1–48.
Batllori, E., Camarero, J. J., & Gutiérrez, E. (2010). Current regeneration patterns at the tree line in the Pyrenees indicate similar recruitment processes irrespective of the past disturbance regime. Journal of Biogeography, 37(10), 1938–1950.
Batllori, E., & Gutiérrez, E. (2008). Regional tree line dynamics in response to global change in the Pyrenees. Journal of Ecology, 96(6), 1275–1288. https://doi.org/10.1111/j.1365-2745.2008.01429.x
Betts, M. M. (1955). The food of titmice in oak woodland. Journal of Animal Ecology, 24(2), 282–323.
Bishton, G. (1986). The diet and foraging behaviour of the dunnock Prunella modularis in a hedgerow habitat. Ibis, 128(4), 526–539.
Blackman, R. L., & Eastop, V. F. (1994). Aphids on the world’s trees: An identification and information guide. CAB International in association with The Natural History Museum.
Bolyen, E., Rideout, J. R., Dillon, M. R., Bokulich, N. A., Abnet, C. C., Al-Ghalith, G. A., Alexander, H., Alm, E. J., Arumugam, M., Asnicar, F., & Bai, Y. (2019). Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology, 37(8), 852–857.
Brandt, M. I., Trouche, B., Quintric, L., Günther, B., Wincker, P., Poulain, J., & Arnaud-Haond, S. (2021). Bioinformatic pipelines combining denoising and clustering tools allow for more comprehensive prokaryotic and eukaryotic metabarcoding. Molecular Ecology Resources, 21(6), 1904–1921.
Bush, A., Compson, Z. G., Monk, W. A., Porter, T. M., Steeves, R., Emilson, E., Gagne, N., Hajibabaei, M., Roy, M., & Baird, D. J. (2019). Studying ecosystems with DNA metabarcoding: Lessons from biomonitoring of aquatic macroinvertebrates. Frontiers in Ecology and Evolution, 7, 434.
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., & Holmes, S. P. (2016). DADA2: High-resolution sample inference from Illumina amplicon data. Nature Methods, 13(7), 581–583.
Carreras, J., Carrillo, E., Massalles, R., & Ninot, J. M. (1996). Delimitation of the supra-forest zone in the Catalan Pyrenees. Bulletin De La Societé Linnéenne De Provence, 47, 27–36.
Ceia, R., Machado, R., & Ramos, J. (2016). Nestling food of three hole-nesting passerine species and experimental increase in their densities in Mediterranean oak woodlands. European Journal of Forest Research, 135, 839–847.
Centre de Biodiversitat de l'Institut d'Estudis Andorrans. (2012). Sistema d’Informació Geogràfica i Mediambiental d’Andorra. Retrieved from 12 May 2021 https://www.iea.ad/mapa-de-cobertes-del-sol-d-andorra-2012.
Chao, A., Gotelli, N. J., Hsieh, T. C., Sander, E. L., Ma, K. H., Colwell, R. K., & Ellison, A. M. (2014). Rarefaction and extrapolation with Hill numbers: A framework for sampling and estimation in species diversity studies. Ecological Monographs, 84(1), 45–67.
Chao, A., & Jost, L. (2012). Coverage-based rarefaction and extrapolation: Standardizing samples by completeness rather than size. Ecology, 93(12), 2533–2547.
Cholewa, M., & Wesołowski, T. (2011). Nestling food of European hole-nesting Passerines: Do we know enough to test the adaptive hypotheses on breeding seasons? Acta Ornithologica, 46(2), 105–116.
Cowie, R. J., & Hinsley, S. A. (1988). Feeding ecology of great tits (Parus major) and blue tits (Parus caeruleus), breeding in suburban gardens. Journal of Animal Ecology, 57(2), 611–626.
Crisol-Martínez, E., Moreno-Moyano, L. T., Wormington, K. R., Brown, P. H., & Stanley, D. (2016). Using next-generation sequencing to contrast the diet and explore pest-reduction services of sympatric bird species in macadamia orchards in Australia. PLoS ONE, 11(3), e0150159.
Currie, D., Nour, N., & Adriaensen, F. (1996). A new technique for filming prey delivered to nestlings, making minimal alterations to the nest box. Bird Study, 43, 380–382.
da Silva, L. P., Mata, V. A., Lopes, P. B., Lopes, R. J., & Beja, P. (2020). High-resolution multi-marker DNA metabarcoding reveals sexual dietary differentiation in a bird with minor dimorphism. Ecology and Evolution, 10(19), 10364–10373.
Davies, S., Vaughan, I., Thomas, R., Marchbank, A., Drake, L., & Symondson, W. (2022). Seasonal and ontological variation in diet and age-related differences in prey choice, by an insectivorous songbird. Ecology and Evolution, 12(8), e9180.
Davis, N. M., Proctor, D. M., Holmes, S. P., Relman, D. A., & Callahan, B. J. (2018). Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome, 6(1), 226.
Deagle, B. E., Jarman, S. N., Coissac, E., Pompanon, F., & Taberlet, P. (2014). DNA metabarcoding and the cytochrome c oxidase subunit I marker: not a perfect match. Biology Letters, 10(9), 20140562.
Deagle, B. E., Thomas, A. C., Shaffer, A. K., Trites, A. W., & Jarman, S. N. (2013). Quantifying sequence proportions in a DNA-based diet study using Ion Torrent amplicon sequencing: Which counts count? Molecular Ecology Resources, 13(4), 620–633.
Fayt, P., Machmer, M. M., & Steeger, C. (2005). Regulation of spruce bark beetles by woodpeckers—a literature review. Forest Ecology and Management, 206(1), 1–14.
Feuillet, T., Birre, D., Milian, J., Godard, V., Clauzel, C., & Serrano-Notivoli, R. (2020). Spatial dynamics of alpine tree lines under global warming: What explains the mismatch between tree densification and elevational upward shifts at the tree line ecotone? Journal of Biogeography, 47(5), 1056–1068.
Flousek, J., Telenský, T., Hanzelka, J., & Reif, J. (2015). Population trends of central European montane birds provide evidence for adverse impacts of climate change on high-altitude species. PLoS ONE, 10(10), e0139465.
Galan, M., Pons, J. B., Tournayre, O., Pierre, É., Leuchtmann, M., Pontier, D., & Charbonnel, N. (2018). Metabarcoding for the parallel identification of several hundred predators and their prey: Application to bat species diet analysis. Molecular Ecology Resources, 18(3), 474–489.
Garfinkel, M., Minor, E., & Whelan, C. J. (2022). Using faecal metabarcoding to examine consumption of crop pests and beneficial arthropods in communities of generalist avian insectivores. Ibis, 164(1), 27–43.
Gibb, J. (1954). Feeding ecology of tits, with notes on treecreeper and goldcrest. Ibis, 96(4), 513–543.
Grace, J., Berninger, F., & Nagy, L. (2002). Impacts of climate change on the tree line. Annals of Botany, 90(4), 537–544.
Gruner, D. S. (2004). Attenuation of top-down and bottom-up forces in a complex terrestrial community. Ecology, 85(11), 3010–3022.
Grzędzicka, E. (2018). Habitat and diet variability of two coexisting tit species in central European forests. Bird Study, 65(1), 52–61.
Gunnarsson, B. (1995). Arthropods and passerine birds in coniferous forest: The impact of acidification and needle-loss. Ecological Bulletins, 44, 248–258.
Gunnarsson, B. (1996). Bird predation and vegetation structure affecting spruce-living arthropods in a temperate forest. Journal of Animal Ecology, 65(3), 389–397.
Hajibabaei, M., Singer, G. A. C., Hebert, P. D. N., & Hickey, D. A. (2007). DNA barcoding: How it complements taxonomy, molecular phylogenetics and population genetics. Trends in Genetics, 23(4), 167–172.
Hartig, F. (2022). DHARMa: Residual diagnostics for hierarchical (multi-level/mixed) regression models. R package version 0.4.6. R Development Core Team.
Hartley, P. H. T. (1987). Ecological aspects of the foraging behaviour of crested tits Parus cristatus. Bird Study, 34(2), 107–111.
Hicks, T. M., Knowles, S. O., & Farouk, M. M. (2018). Global provisioning of red meat for flexitarian diets. Frontiers in Nutrition, 5, 50–50.
Hodkinson, I. D. (2005). Terrestrial insects along elevation gradients: Species and community responses to altitude. Biological Reviews, 80(3), 489–513.
Hou, J., Li, L., Wang, Y., Wang, W., Zhan, H., Dai, N., & Lu, P. (2021). Influences of submerged plant collapse on diet composition, breadth, and overlap among four crane species at Poyang Lake, China. Frontiers in Zoology, 18(1), 24.
Inger, R., Gregory, R., Duffy, J. P., Stott, I., Vorisek, P., & Gaston, K. J. (2015). Common European birds are declining rapidly while less abundant species’ numbers are rising. Ecology Letters, 18(1), 28–36.
Jarrett, C., Powell, L. L., McDevitt, H., Helm, B., & Welch, A. J. (2020). Bitter fruits of hard labour: Diet metabarcoding and telemetry reveal that urban songbirds travel further for lower-quality food. Oecologia, 193, 377–388.
Jusino, M. A., Banik, M. T., Palmer, J. M., Wray, A. K., Xiao, L., Pelton, E., Barber, J. R., Kawahara, A. Y., Gratton, C., Peery, M. Z., & Lindner, D. L. (2017). An improved method for utilizing high-throughput amplicon sequencing to determine the diets of insectivorous animals. Molecular Ecology Resources, 19, 176–190.
Kent, C. M., Huh, K. M., Hunter, S. C., Judson, K., Powell, L. L., & Sherry, T. W. (2022). High resource overlap and small dietary differences are widespread in food-limited warbler (Parulidae) communities. Ibis, 164(1), 44–59.
Körner, C. (2012). Alpine treelines: Functional ecology of the global high elevation tree limits. Springer.
Krehenwinkel, H., Wolf, M., Lim, J. Y., Rominger, A. J., Simison, W. B., & Gillespie, R. G. (2017). Estimating and mitigating amplification bias in qualitative and quantitative arthropod metabarcoding. Science and Reports, 7, 17668–17612.
Lack, D. (1964). A long-term study of the great tit (Parus major). Journal of Animal Ecology, 33, 159–173.
Lens, L. (1996). Wind stress affects foraging site competition between crested tits and willow tits. Journal of Avian Biology, 27(1), 41–46.
Lister, B. C. (1980). Resource variation and the structure of British bird communities. Proceedings of the National Academy of Sciences—PNAS, 77(7), 4185–4187.
López-Moreno, J. I., Gascoin, S., Herrero, J., Sproles, E. A., Pons, M., Alonso-González, E., Hanich, L., Boudhar, A., Musselman, K. N., Molotch, N. P., & Sickman, J. (2017). Different sensitivities of snowpacks to warming in Mediterranean climate mountain areas. Environmental Research Letters, 12(7), 074006.
Mansor, M. S., Rozali, F. Z., Davies, S., Nor, S. M., & Ramli, R. (2022). High-throughput sequencing reveals dietary segregation in Malaysian babblers. Current Zoology, 68, 381–389.
Martoni, F., Piper, A. M., Rodoni, B. C., & Blacket, M. J. (2022). Disentangling bias for non-destructive insect metabarcoding. PeerJ, 10, e12981–e12981.
McCain, C. M. (2009). Global analysis of bird elevational diversity. Global Ecology and Biogeography, 18(3), 346–360.
McClenaghan, B., Nol, E., & Kerr, K. C. R. (2019). DNA metabarcoding reveals the broad and flexible diet of a declining aerial insectivore. The Auk, 136(1), 1–11.
Michalski, M., Nadolski, J., Marciniak, B., Loga, B., & Bańbura, J. (2011). Faecal analysis as a method of nestling diet determination in insectivorous birds: A case study in blue tits Cyanistes caeruleus and great tits Parus major. Acta Ornithologica, 46(2), 164–172.
Moran, A. J., Prosser, S. W. J., & Moran, J. A. (2019). DNA metabarcoding allows non-invasive identification of arthropod prey provisioned to nestling Rufous hummingbirds (Selasphorus rufus). PeerJ, 7, e6596.
Mountain Research Initiative EDW Working Group. (2015). Elevation-dependent warming in mountain regions of the world. Nat Clima Chang., 5(5), 424–430.
Navarro-Serrano, F., López-Moreno, J., Azorin-Molina, C., Alonso-González, E., Aznarez-Balta, M., Buisan, S., & Revuelto, J. (2020). Elevation effects on air temperature in a topographically complex mountain valley in the Spanish Pyrenees. Atmosphere, 11, 656.
Nielsen, J. M., Clare, E. L., Hayden, B., Brett, M. T., & Kratina, P. (2018). Diet tracing in ecology: Method comparison and selection. Methods in Ecology and Evolution, 9(2), 278–291.
Nour, N., Currie, D., Matthysen, E., Van Damme, R., & Dhondt, A. A. (1998). Effects of habitat fragmentation on provisioning rates, diet and breeding success in two species of tit (great tit and blue tit). Oecologia, 114(4), 522–530.
Nyffeler, M., Şekercioğlu, Ç. H., & Whelan, C. J. (2018). Insectivorous birds consume an estimated 400–500 million tons of prey annually. The Science of Nature, 105(7), 47.
O’Rourke, D., Bokulich, N., Jusino, M., MacManes, M., & Foster, J. (2020). A total crapshoot? Evaluating bioinformatic decisions in animal diet metabarcoding analyses. Ecology and Evolution, 10(18), 9721–9739.
Obeso, J. R. (1987). Uso del espacio y alimentation de los Parus spp en bosques mixtos de la Sierra de Cazorla. Ardeola, 34(1), 61–77.
Oksanen, J., Blanchet, F.G., Kindt, R., Legendre, P., Minchin, P.R., O'Hara, R.B. and Wagner, H. (2020). vegan: Community Ecology Package Version R package version 2.5-7
OPCC-CTP. (2018). Climate change in the Pyrenees: Impacts, vulnerabilities and adaptation. Observatory PCC.
Orłowski, G., Frankiewicz, J., & Karg, J. (2017). Nestling diet optimization and condition in relation to prey attributes and breeding patch size in a patch-resident insectivorous passerine: An optimal continuum and habitat constraints. Journal of Ornithology, 158(1), 169–184.
Öztürk, M. A., Hakeem, K. R., Hanum, I. F., & Efe, R. (2015). Climate change impacts on high-altitude ecosystems. Springer.
Pagani-Núñez, E., Ruiz, I., Quesada, J., Negro, J., & Senar, J. C. (2011). The diet of great tit Parus major nestlings in a Mediterranean Iberian forest: The important role of spiders. Animal Biodiversity and Conservation, 34, 355–361.
Philpott, S. M., Greenberg, R., Bichier, P., & Perfecto, I. (2004). Impacts of major predators on tropical agroforest arthropods: comparisons within and across taxa. Oecologia, 140(1), 140–149.
Piñol, J., Mir, G., Gomez-Polo, P., & Agusti, N. (2015). Universal and blocking primer mismatches limit the use of high-throughput DNA sequencing for the quantitative metabarcoding of arthropods. Molecular Ecology Resources, 15(4), 819–830.
R Core Team. (2021). R: A language and environment for statistical computing. R foundation for statistical computing version 1.3.1056. R Core Team.
Rahbek, C. (2005). The role of spatial scale and the perception of large-scale species-richness patterns. Ecology Letters, 8(2), 224–239.
Resano-Mayor, J., Korner-Nievergelt, F., Vignali, S., Horrenberger, N., Barras, A. G., Braunisch, V., Pernollet, C. A., & Arlettaz, R. (2019). Snow cover phenology is the main driver of foraging habitat selection for a high-alpine passerine during breeding: Implications for species persistence in the face of climate change. Biodiversity and Conservation, 28(10), 2669–2685.
Ribeiro, Â., Smit, B., & Gilbert, M. (2019). 31° South: Dietary niche of an arid-zone endemic passerine. Environmental DNA, 1, 109–118.
Ritter, C. D., Dal Pont, G., Stica, P. V., Horodesky, A., Cozer, N., Netto, O. S. M., Henn, C., Ostrensky, A., & Pie, M. R. (2022). Wanted not, wasted not: Searching for non-target taxa in environmental DNA metabarcoding by-catch. Environmental Advances, 7, 100169.
Roura, N., Pons, P., Etienne, M., & Lambert, B. (2005). Transformation of a rural landscape in the eastern Pyrenees between 1953 and 2000. Mountain Research and Development, 25, 252–261.
Rytkönen, S., Vesterinen, E. J., Westerduin, C., Leviäkangas, T., Vatka, E., Mutanen, M., Välimäki, P., Hukkanen, M., Suokas, M., & Orell, M. (2019). From feces to data: A metabarcoding method for analyzing consumed and available prey in a bird-insect food web. Ecology and Evolution, 9(1), 631–639.
Sanz, J. J., Garcia-Navas, V., & Ruiz-Peinado, J. V. (2010). Effect of habitat type and nest-site characteristics on the breeding performance of great and blue tits (Parus major and P. caeruleus) in a Mediterranean landscape. Ornis Fennica, 87(2), 41.
Schwenk, W. S., Strong, A. M., & Sillett, T. S. (2010). Effects of bird predation on arthropod abundance and tree growth across an elevational gradient. Journal of Avian Biology, 41(4), 367–377.
Sehhatisabet, M. E., Kiabi, B., Pazuki, A., Alipanah, H., Khaleghizadeh, A., Barari, H., Basiri, R., & Aghabeigi, F. (2008). Food diversity and niche-overlap of sympatric tits (great tit, Parus major, blue tit, Cyanistes caeruleus and coal tit Periparus ater) in the Hyrcanian Plain forests. Zoology in the Middle East, 44(1), 18–30.
Şekercioğlu, Ç. H., Fullwood, M. J., Cerling, T. E., Brenes, F. O., Daily, G. C., Ehrlich, P. R., Chamberlain, P., & Newsome, S. D. (2023). Using stable isotopes to measure the dietary responses of Costa Rican forest birds to agricultural countryside. Frontiers in Ecology and Evolution, 11, 202.
Senécal, S., Riva, J.-C., O’Connor, R. S., Hallot, F., Nozais, C., & Vézina, F. (2021). Poor prey quality is compensated by higher provisioning effort in passerine birds. Science and Reports, 11, 11182.
Shoffner, A., Wilson, A. M., Tang, W., & Gagné, S. A. (2018). The relative effects of forest amount, forest configuration, and urban matrix quality on forest breeding birds. Science and Reports, 8, 17140.
Shutt, J. D., Nicholls, J. A., Trivedi, U. H., Burgess, M. D., Stone, G. N., Hadfield, J. D., & Phillimore, A. B. (2020). Gradients in richness and turnover of a forest passerine’s diet prior to breeding: A mixed model approach applied to faecal metabarcoding data. Molecular Ecology, 29(6), 1199–1213.
Snider, A. M., Bonisoli-Alquati, A., Pérez-Umphrey, A. A., Stouffer, P. C., & Taylor, S. S. (2021). Metabarcoding of stomach contents and fecal samples provide similar insights about Seaside Sparrow diet. Ornithological Applications, 124(1), 1–12.
Sottas, C., Reif, J., Kreisinger, J., Schmiedová, L., Sam, K., Osiejuk, T. S., & Reifová, R. (2020). Tracing the early steps of competition-driven eco-morphological divergence in two sister species of passerines. Evolutionary Ecology, 34(4), 501–524.
Suarez-Seoane, S., Osborne, P. E., & Baudry, J. (2002). Responses of birds of different biogeographic origins and habitat requirements to agricultural land abandonment in northern Spain. Biological Conservation, 105(3), 333–344.
Sutton, A. O., Strickland, D., Freeman, N. E., & Norris, D. R. (2021). Climate-driven carry-over effects negatively influence population growth rate in a food-caching boreal passerine. Global Change Biology, 27(5), 983–992.
Tallamy, D. W., & Shriver, W. G. (2021). Are declines in insects and insectivorous birds related? Ornithological Applications, 123(1), 8.
Torok, J., & Tóth, L. (1999). Asymmetric competition between two tit species: A reciprocal removal experiment. Journal of Animal Ecology, 68(2), 338–345.
Ulfstrand, S. (1976). Feeding niches of some passerine birds in a south Swedish coniferous plantation in winter and summer. Ornis Scandinavica, 7(1), 21–27.
Ulfstrand, S. (1977). Foraging niche dynamics and overlap in a guild of Passerine birds in a south Swedish coniferous woodland. Oecologia, 27(1), 23–45.
Wagner, J. L. (1981). Seasonal change in guild structure: Oak woodland insectivorous birds. Ecology, 62(4), 973–981.
Wesołowski, T., Cholewa, M., Hebda, G., Maziarz, M., & Rowiński, P. (2016). Immense plasticity of timing of breeding in a sedentary forest passerine, Poecile palustris. Journal of Avian Biology, 47(1), 129–213.
Acknowledgements
Many people were integral to the success of this project. Earthwatch Institute volunteers donated countless hours dedicated to capturing birds and collecting samples. Cristina Ametller Quero was extremely helpful with organizing and filtering DNA sequences and initiated much of the statistical work. Cesc Murria and Josep Piñol were especially helpful in regard to the bioinformatic processes. Roberto Molowny provided critical expertise to statistics sections. Gerald Dinkins contributed thoughtful proofreading assistance. Amie Carlone’s GIS advice was very helpful. Funding was provided by Earthwatch Institute, Daniel B. Warnell School of Forestry and Natural Resources within the University of Georgia, and the Collections Section of the Museu de Ciències Naturals de Barcelona
Funding
Open Access Funding provided by Universitat Autonoma de Barcelona.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bookwalter, J., Niyas, A.M.M., Caballero-López, B. et al. DNA metabarcoding Passerine bird feces at tree-line uncovers little intra- and inter-species dietary overlap. COMMUNITY ECOLOGY 24, 147–157 (2023). https://doi.org/10.1007/s42974-023-00148-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42974-023-00148-4