Abstract
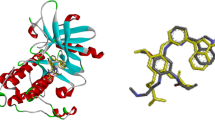
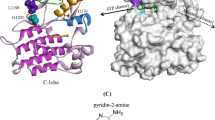
Epidermal growth factor receptor (EGFR) inhibitors have been proven as a high-potential therapeutic target for treating advanced non-small-cell lung cancer (NSCLC). However, many patients still suffer from drug-resistant mutations and drug side effects. The current study aims to investigate Osimertinib-based compounds inhibitory activity against double mutant EGFRL858R/T790M by computational methods of 3D-QSAR, molecular docking, and molecular dynamic (MD) simulations. CoMFA and CoMSIA approaches were used to create the 3D-QSAR models. Molecular docking and molecular dynamics simulation was performed to generate the binding mode and stability of the investigated inhibitors. The CoMFA model achieved good predictability with Q2 = 0.663, R2 = 0.978, SEE = 0.115, and an acceptable value for the coefficient of determination Rtest2 = 0.756. In addition, based on the information retained by the CoMFA contour maps, we proposed four new molecules (T1–T4) with significantly higher inhibitory activity. Furthermore, molecular docking and MD simulation analysis were utilized to confirm the 3D-QSAR results, supporting the stability of the proposed molecules in the 3W2O receptor. Finally, the selected molecules showed favorable pharmacokinetic properties and were non-toxic. The results provide significant information about the proposed Osimertinib-based compounds as potential H1975 Cell Line drugs that encourage further innovative experimental and clinical research.
Graphical Abstract











Similar content being viewed by others
Referencess
Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S, Isobe H, Gemma A, Harada M, Yoshizawa H, Kinoshita I, Fujita Y, Okinaga S, Hirano H, Yoshimori K, Harada T, Ogura T, Ando M, Miyazawa H, Tanaka T, Saijo Y, Hagiwara K, Morita S, Nukiwa T (2010) Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N Engl J Med 362(25):2380–2388. https://doi.org/10.1056/NEJMoa0909530
Wang Y, Schmid-Bindert G, Zhou C (2012) Erlotinib in the treatment of advanced non-small cell lung cancer: an update for clinicians. Ther Adv Med Oncol 4(1):19–29. https://doi.org/10.1177/1758834011427927
Zhou W, Ercan D, Chen L, Yun CH, Li D, Capelletti M, Cortot AB, Chirieac L, Iacob RE, Padera R, Engen JR, Wong KK, Eck MJ, Gray NS, Jänne PA (2009) Novel mutant-selective EGFR kinase inhibitors against EGFR T790M. Nature 462(7276):1070–1074. https://doi.org/10.1038/nature08622
Walter AO, Sjin RT, Haringsma HJ, Ohashi K, Sun J, Lee K, Dubrovskiy A, Labenski M, Zhu Z, Wang Z, Sheets M, Martin TS, Karp R, Kalken D, Chaturvedi P, Niu D, Nacht M, Petter RC, Westlin W, Lin K, Allen A (2013) Discovery of a mutant-selective covalent inhibitor of EGFR that overcomes T790M mediated resistance in NSCLC. Cancer Discov 3:1404–1415. https://doi.org/10.1158/2159-8290.CD-13-0314
Finlay MR, Anderton M, Ashton S, Ballard P, Bethel PA, Box MR, Brad-bury RH, Brown SJ, Butterworth S, Campbell A, Chorley C, Colclough N, Cross DA, Currie GS, Grist M, Hassall L, Hill GB, James D, James M, Kemmitt PGL (2014) Wrigley Discovery of a potent and selective EGFR inhibitor (AZD9291) of both sensitizing and T790M resistance mutations that spares the wild type form of the receptor. J Med Chem 57:8249–8267
Ahmad I, Shaikh M, Surana S, Ghosh A, Patel H (2022) p38α MAP kinase inhibitors to overcome EGFR tertiary C797S point mutation associated with osimertinib in non-small cell lung cancer (NSCLC): emergence of fourth-generation EGFR inhibitor. J Biomol Struct Dyn 40(7):3046–3059
Jänne PA, Yang JCH, Kim DW, Planchard D, Ohe Y, Ramalingam SS, Ahn MJ, Kim SW, Su WC, Horn L, Haggstrom D, Felip E, Kim JH, Frewer P, Cantarini M, Brown KH, Dickinson PA, Ghiorghiu S, Ranson M (2015) AZD9291 in EGFR inhibitor–resistant non–small-cell lung cancer. N Engl J Med 372(18):1689–1699. https://doi.org/10.1056/NEJMoa1411817
Patel HM, Shaikh M, Ahmad I, Lokwani D, Surana SJ (2021) BREED based de novo hybridization approach: generating novel T790M/C797S-EGFR tyrosine kinase inhibitors to overcome the problem of mutation and resistance in non-smallcell lung cancer (NSCLC). J Biomol Struct Dyn 39(8):2838–2856. https://doi.org/10.1080/07391102.2020.1754918
Miyashita Y, Ko R, Shimada N, Mitsuishi Y, Miura K, Matsumoto N, Asao T, Shukuya T, Shibayama R, Koyama R, Takahashi K (2021) Impact of the generation of EGFR-TKIs administered as prior therapy on the efficacy of Osimertinib in patients with non-small cell lung cancer harboring EGFR T790M mutation. Thorac Cancer 12(3):329–338. https://doi.org/10.1111/1759-7714.13742
Oxnard GR, Hu Y, Mileham KF, Husain H, Costa DB, Tracy P, Feeney N, Sholl LM, Dahlberg SE, Redig AJ, Kwiatkowski DJ, Rabin MS, Paweletz CP, Thress KS, Jänne PA (2018) Assessment of resistance mechanisms and clinical implications in patients with EGFR T790M-positive lung cancer and acquired resistance to osimertinib. JAMA Oncol 4(11):1527–1534
Harun P, Iqrar A, Harsha J, Rahul P, Deepak L, Sanjay S (2022) Investigating the impact of different acrylamide (electrophilic warhead) on Osimertinib’s pharmacological spectrum by molecular mechanic and quantum mechanic approach. Comb Chem High Throughput Screen 25:149-166(18). https://doi.org/10.2174/1386207323666201204125524
Al-Mulla A (2017) A review: biological importance of heterocyclic compounds. Der Pharma Chem 9(13):141–147
Reutskaya E, Osipyan A, Sapegin A, Novikov AS, Krasavin M (2018) Rethinking hydrolytic imidazoline ring expansion: a common approach to the preparation of medium-sized rings via side-chain insertion into [1.4] oxa-and [1.4] thiazepinone scaffolds. J Org Chem 84(4):1693–1705
Othman IM, Alamshany ZM, Tashkandi NY, Gad-Elkareem MA, Anwar MM, Nossier ES (2021) New pyrimidine and pyrazole-based compounds as potential EGFR inhibitors: synthesis, anticancer, antimicrobial evaluation and computational studies. Bioorg Chem 114:105078
Sherbiny FF, Bayoumi AH, El-Morsy AM, Sobhy M, Hagras M (2021) Design, synthesis, biological evaluation, and molecular docking studies of novel Pyrazolo [3, 4-d] Pyrimidine derivative scaffolds as potent EGFR inhibitors and cell apoptosis inducers. Bioorg Chem 116:105325
Mikherdov AS, Novikov AS, Kinzhalov MA, Zolotarev AA, Boyarskiy VP (2018) Intra-/intermolecular bifurcated chalcogen bonding in crystal structure of thiazole/thiadiazole derived binuclear (diaminocarbene) PdII complexes. Cryst 8(3):112
Kardile RA, Sarkate AP, Lokwani DK, Tiwari SV, Azad R, Thopate SR (2023) Design, synthesis, and biological evaluation of novel quinoline derivatives as small molecule mutant EGFR inhibitors targeting resistance in NSCLC: In vitro screening and ADME predictions. Eur J Med Chem 245:114889
Osipyan A, Sapegin A, Novikov AS, Krasavin M (2018) Rare medium-sized rings prepared via hydrolytic imidazoline ring expansion (HIRE). J Org Chem 83(17):9707–9717
Joshi G, Nayyar H, Kalra S, Sharma P, Munshi A, Singh S, Kumar R (2017) Pyrimidine containing epidermal growth factor receptor kinase inhibitors: synthesis and biological evaluation. Chem Biol Drug Des 90(5):995–1006
Nasser AA, Eissa IH, Oun MR, El-Zahabi MA, Taghour MS, Belal A, Saleh AM, Mehany ABM, Luesch H, Mostafa AE, Afifi WM, Rocca JR, Mahdy HA (2020) Discovery of new pyrimidine-5-carbonitrile derivatives as anticancer agents targeting EGFR WT and EGFR T790M. Org Biomol Chem 18(38):7608–7634
Gaber AA, Bayoumi AH, El-Morsy AM, Sherbiny FF, Mehany AB, Eissa IH (2018) Design, synthesis and anticancer evaluation of 1H-pyrazolo [3, 4-d] pyrimidine derivatives as potent EGFRWT and EGFRT790M inhibitors and apoptosis inducers. Bioorg Chem 80:375–395
Hao Y, Lyu J, Qu R, Tong Y, Sun D, Feng F, Tong L, Yang L, Zhao Z, Zhu L, Ding J, Xu Y, Xie H, Li H (2018) Design, Synthesis, and biological evaluation of pyrimido[4, 5-d] pyrimidine-2, 4 (1H, 3H)-diones as potent and selective epidermal growth factor receptor (EGFR) inhibitors against L858R/T790M resistance mutation. J Med Chem 61(13):5609–5622
Li J, An B, Song X, Zhang Q, Chen C, Wei S, Fan R, Li X, Zou Y (2021) Design, synthesis and biological evaluation of novel 2, 4-diaryl pyrimidine derivatives as selective EGFRL858R/T790M inhibitors. Eur J Med Chem 212:113019
Ding S, Dong X, Gao Z, Zheng X, Ji J, Zhang M, Liu F, Wu S, Li M, Song W, Shen J, Duan W, Liu J, Chen Y (2022) Design synthesis and biological evaluation of novel N-(3-amino-4-methoxyphenyl) acrylamide derivatives as selective EGFRL858R/T790M kinase inhibitors. Bioorg Chem 118:105471. https://doi.org/10.1016/j.bioorg.2021.105471
Abdullahi SH, Uzairu A, Shallangwa GA, Uba S, Umar AB (2022) Structure based design of some novel 3-methylquinoxaline derivatives through molecular docking and pharmacokinetics studies as novel VEGFR-2 inhibitors. Chem Afr 5:1967–1978
Zhu J, Wu Y, Xu L, Jin J (2020) Theoretical studies on the selectivity mechanisms of glycogen synthase kinase 3β (GSK3β) with pyrazine ATP-competitive inhibitors by 3D-QSAR, molecular docking, molecular dynamics simulation and free energy calculations. Curr Comput-Aided Drug Des 16(1):17–30
Abdullahi SH, Uzairu A, Shallangwa GA, Uba S, Umar AB (2023) 2D and 3D-QSAR modeling of 1H-pyrazole derivatives as EGFR inhibitors: molecular docking, and pharmacokinetic profiling. Chem Afr 6:1381–1398. https://doi.org/10.1007/s42250-023-00592-9
Dhingra N, Kar A, Sharma R (2020) Towards further understanding the structural requirements of combretastatin-like chalcones as inhibitors of microtubule polymerization. Curr Comput-Aided Drug Des 16(2):155–166
Arakawa M, Hasegawa K, Funatsu K (2007) The recent trend in QSAR modeling-variable selection and 3D-QSAR methods. Curr Comput-Aided Drug Des 3(4):254–262
Clark M, Cramer RD, Opdenbosch NV (1989) Validation of the general purpose tripos 5.2 force field. J Comput Chem 10:982–1012. https://doi.org/10.1002/jcc.540100804
Powell MJD (1977) Restart procedures for the conjugate gradient method. Math Program 12:241–254
Tsai KC, Chen YC, Hsiao NW, Wang CL, Lin CL, Lee YC, Li M, Wang BA (2010) Comparison of different electrostatic potentials on prediction accuracy in CoMFA and CoMSIA studies. Eur J Med Chem 45(4):1544–1551
Sogabe S, Kawakita Y, Igaki S, Iwata H, Miki H, Cary DR, Ishikawa T (2013) Structure-based approach for the discovery of pyrrolo [3, 2-d] pyrimidine-based EGFR T790M/L858R mutant inhibitors. ACS Med Chem Lett 4(2):201–205. https://doi.org/10.1021/ml300327z
BIOVIA DS (2016) Discovery studio modeling environment, release 2017, San Diego: DassaultSystèmes, 2016
DeLano W (2017) The PyMOL molecular graphics system. Delano Scientific, Palo Alto
Tabti K, Elmchichi L, Sbai A, Maghat H, Bouachrine M, Lakhlifi T (2022) HQSAR, CoMFA, CoMSIA docking studies and simulation MD on quinazolines/quinolines derivatives for DENV virus inhibitory activity. Chem Afr 5(6):1937–1958
Abraham MJ, Murtola T, Schulz R, Pall S, Smith JC, Hess B, Lindahl E (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1:19–25
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26(16):1701–1718
Kumari R, Kumar R (2014) Open Source Drug Discovery Consortium, Lynn, A. g_mmpbsa A GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54(7):1951–1962
Safavi A, Ghodousi ES, Ghavamizadeh M, Sabaghan M, Azadbakht O, Babaei H, Zarezade V (2021) Computational investigation of novel farnesyltransferase inhibitors using 3D-QSAR pharmacophore modeling, virtual screening, molecular docking and molecular dynamics simulation studies: a new insight into cancer treatment. J Mol Struct 1241:130667. https://doi.org/10.1016/j.molstruc.2021.130667
Ghaleb A, Aouidate A, Ayouchia HBE, Aarjane M, Anane H, Stiriba SE (2022) In silico molecular investigations of pyridine N-Oxide compounds as potential inhibitors of SARS-CoV-2: 3D QSAR, molecular docking modeling, and ADMET screening. J Biomol Struct Dyn 40(1):143–153. https://doi.org/10.1080/07391102.2020.1808530
Pires DE, Blundell TL, Ascher DB (2015) pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem 58(9):4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7(1):42717. https://doi.org/10.1038/srep42717
Author information
Authors and Affiliations
Contributions
All contributors aided the study's inception and design. El Bahi S, Boutalaka M, El Alaouy M, Koubi Y, and El Khatabi K: presented idea, draft preparation, data handling, data analysis, and writing; Alaqarbeh M: performed the calculations of molecular dynamics, writing, review, and editing; Choukrad M, Sbai A, and Bouachrine M: data analysis, study justification, reviewing, and supervision; Lakhlifi T: reviewing supervision, and project administration.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
El Bahi, S., Boutalaka, M., Alaqarbeh, M. et al. In-Silico Investigation of Osimertinib Based Compounds as Potential Double Mutant EGFR Kinase Inhibitors Against H1975 Cell Line: Integrating QSAR Modeling, Molecular Docking, MD Simulations, and ADME/Tox Studies. Chemistry Africa 7, 111–129 (2024). https://doi.org/10.1007/s42250-023-00744-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42250-023-00744-x