Abstract
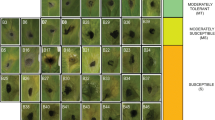
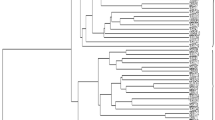
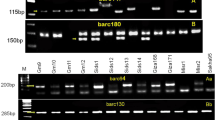
Wheat (Triticum spp.) is a global staple food crop, contributing significantly to the world's food security. Understanding and harnessing the genetic diversity within wheat cultivars is paramount for developing resilient and high-yielding varieties. The present study reports rust response of 31 registered rust resistant genetic stocks of wheat against recently identified and most virulent pathotypes of all three rust pathogens and their morphological and molecular diversity assessment. Analysis of variance (ANOVA) showed indicated significant differences among the genotypes for all the studied traits. Among 31 genetic stocks 30, 15, and 8 were found resistant against all the tested pathotypes of stem, leaf and stripe rust pathogens, respectively, whereas only two (FLW21 and FLW28) conferred resistance against all three rusts. Molecular profiling with 59 polymorphic SSRs resulted in 194 alleles with an average 3 alleles/loci. With an average of 0.54, the Polymorphism Information Content (PIC) varied from 0.34 to 0.75, reflecting higher allelic variation. The average gene diversity, heterozygosity, major allele frequency, and minor allele frequency were 0.61, 0.31, 0.48, and 0.52, respectively. Cluster analysis grouped 31 genetic stocks into 3 clusters. The AMOVA revealed that within population variation was higher than between them (76% vs. 24%). Clustering was further supported by the structure and Principal Coordinate Analysis (PCoA). Structure analysis grouped the genetic stocks into three sub-populations. These findings will help in suggesting different cross combinations for wheat rust resistance breeding and pyramiding of multiple rust resistance genes.



Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Abbasov M, Akparov Z, Gross T, Babayeva S, Izzatullayeva V, Hajiyev E, Rustamov K, Gross P, Tekin M, Akar T, Chao S (2018) Genetic relationship of diploid wheat (Triticum spp.) species assessed by SSR markers. Genetic Resources and Crop Evolution 65:1441–1453
Aboukhaddour R, Fetch T, McCallum BD, Harding MW, Beres BL, Graf RJ (2020) Wheat diseases on the prairies: A Canadian story. Plant Pathology 69:418–432
Adhikari S, Kumari J, Jacob SR, Prasad P, Gangwar OP, Lata C, Thakur R, Singh AK, Bansal R, Kumar S, Bhardwaj SC (2022) Landraces-potential treasure for sustainable wheat improvement. Genet Resour Crop Evol 69(2):499–523
Asmamaw M, Keneni G, Tesfaye K (2019) Genetic diversity of Ethiopian durum wheat (Triticum durum Desf) landrace collections as revelled by SSR markers. Adv Crop Sci Tech 7(1):413
Babu P, Baranwal DK, Harikrishna Pal D, Bharti H, Joshi P, Thiyagarajan B, Gaikwad KB, Bhardwaj SC, Singh GP, Singh A (2020) Application of genomics tools in wheat breeding to attain durable rust resistance. Frontiers in Plant Science 11:567147
Beddow JM, Pardey PG, Chai Y, Hurley TM, Kriticos DJ, Braun H-J, Park RF, Cuddy WS, Yonow T (2015) Research investment implications of shifts in the global geography of wheat stripe rust. Nature Plants 1:1–5
Belete Y, Shimelis H, Laing M, Mathew I (2020) Genetic diversity and population structure of bread wheat genotypes determined via phenotypic and SSR marker analyses under drought-stress conditions. Journal of Crop Improvement 35:303–325
Bhardwaj SC, Singh GP, Gangwar OP, Prasad P, Kumar S (2019) Status of wheat rust research and progress in rust management-Indian context. Agronomy 9:892
Bhavani S, Singh RP, Hodson DP, Huerta-Espino J, Randhawa MS (2022) Wheat rusts: current status, prospects of genetic control and integrated approaches to enhance resistance durability. In: Reynolds MP, Braun HJ (eds) Wheat Improvement: Food Security in a Changing Climate. Springer International Publishing, Cham, pp 125–141
Bhawar KB, Sharma JB, Singh AK, Sivasamy M, Singh M, Prabhu KV, Tomar SM, Sharma TR, Singh B (2011) Molecular marker assisted pyramiding of leaf rust resistance genes Lr19 and Lr28 in bread wheat (Triticum aestivum L.) variety HD2687. Indian J Genet Plant Breed 71:304–311
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American journal of human genetics 32:314–331
Chen X, Min D, Yasir TA, Hu YG (2012) Genetic diversity, population structure and linkage disequilibrium in elite Chinese winter wheat investigated with SSR markers. PLoS One 7:e44510
Dagnaw T, Mulugeta B, Haileselassie T, Geleta M, Ortiz R, Tesfaye K (2023) genetic diversity of durum wheat (Triticum turgidum L. ssp. durum, Desf.) germplasm as revealed by morphological and SSR markers. Genes 14:1155
Demirel S, Demirel F (2023) Molecular identification and population structure of emmer and einkorn wheat lines with different ploidy levels using SSR markers. Genetic Resources and Crop Evolution 11:1–10
Devi S, Singh V, Yashveer S, Dalal MS, Chawla R, Akbarzai DK, Chaurasia H (2023) Molecular characterization of bread wheat (Triticum aestivum) genotypes using SSR markers. Indian J Agric Sci (9):948–953
Eltaher S, Sallam A, Belamkar V, Emara HA, Nower AA, Salem KF, Poland J, Baenziger PS (2018) Genetic diversity and population structure of F3:6 Nebraska winter wheat genotypes using genotyping-by-sequencing. Frontiers in genetics 9:76
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular ecology 14:2611–2620
Farhangian-Kashani S, Azadi A, Khaghani S, Changizi M, Gomarian M (2021) Association analysis and evaluation of genetic diversity in wheat genotypes using SSR markers. BiologiaFutura 72(4):441–452
Hao C, Wang L, Ge H, Dong Y, Zhang X (2011) Genetic diversity and linkage disequilibrium in Chinese bread wheat (Triticum aestivum L.) revealed by SSR markers. PLoS One 6:e17279
Haque MS, Saha NR, Islam MT (2021) Screening for drought tolerance in wheat genotypes by morphological and SSR markers. Journal of Crop Science and Biotechnology 24:27–39
Iqbal N, Tabasum A, Sayed H, Hameed A (2009) Evaluation of genetic diversity among bread wheat varieties and landraces of Pakistan by SSR markers. Cereal Res Commun 37(4):489–498
Jabari M, Golparvar A, Sorkhilalehloo B, Shams M (2023) Investigation of genetic diversity of Iranian wild relatives of bread wheat using ISSR and SSR markers. Journal of Genetic Engineering and Biotechnology 21:73
Jaccard P (1908) NouvellesRecherches Sue la Distribution Florale. Bulletin De La Socie’te’ Vaundoise Des Sciences Naturelles 44:223–0270
Jiang J, Friebe B, Gill BS (1994) Recent advances in alien gene transfer in wheat. Euphytica 73:199–212
Jlassi I, Bnejdi F, Saadoun M, Hajji A, Mansouri D, Ben-Attia M, El-Gazzah M, El-Bok S (2021) SSR markers and seed quality traits revealed genetic diversity in durum wheat (Triticum durum Desf.). Molecular Biology Reports 48:3185–3193
Kaur S, Kaur J, Mavi GS, Dhillon GS, Sharma A, Singh R, Devi U, Chhuneja P (2020) Pyramiding of high grain weight with stripe rust and leaf rust resistance in elite Indian wheat cultivar using a combination of marker assisted and phenotypic selection. Frontiers in genetics 11:593426
Klymiuk V, Yaniv E, Huang L, Raats D, Fatiukha A, Chen S, Feng L, Frenkel Z, Krugman T, Lidzbarsky G, Chang W (2018) Cloning of the wheat Yr15 resistance gene sheds light on the plant tandem kinase-pseudokinase family. Nature communications 9:3735
Kolmer JA (1996) Genetics of resistance to wheat leaf rust. Annual Review of Phytopathology 34:435–455
Kumar P, Yadava RK, Kumar S, Kumar P (2016) Molecular diversity analysis in Wheat genotypes using SSR markers. Electronic Journal of Plant Breeding 7:464–468
Liu K, Muse SV (2005) Power Marker: integrated analysis environment for genetic marker data. Bioinformatics 21:2128–2129
McIntosh RA, Dubcovsky J, Rogers WJ, Morris CF, Appels R, Xia XC (2016) Catalogue of Gene Symbols for Wheat: 2015-2016 (Supplement). Available at: https://shigen.nig.ac.jp/wheat/komugi/genes/macgene/supplement2015.pdf. Accessed 6 May 2024
Mohi-Ud-Din M, Hossain MA, Rohman MM, Uddin MN, Haque MS, Dessoky ES, Alqurashi M, Aloufi S (2022) Assessment of genetic diversity of bread wheat genotypes for drought tolerance using canopy reflectance-based phenotyping and SSR marker-based genotyping. Sustainability 14:9818
Moradkhani H, Mehrabi AA, Etminan A, Pour-Aboughadareh A (2015) Molecular diversity and phylogeny of Triticum-Aegilops species possessing D genome revealed by SSR and ISSR markers. Plant Breeding and Seed Science 71:81–95
Mundt CC (2018) Pyramiding for resistance durability: theory and practice. Phytopathology 108:792–802
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Research 8:4321–4326
Nei M (1972) Analysis of gene diversity in subdivided populations. Proceedings of National Academy of Science of the United States of America 70:3321–3323
Oliver RP (2014) A reassessment of the risk of rust fungi developing resistance to fungicides. Pest Management Science 70:1641–1645
Pardey PG, Beddow J, Kriticos D, Hurley T, Park R, Duveiller E, Sutherst R, Burdon J, Hodson D (2013) Right-sizing stem-rust research. Science 340:147–148
Peakall R, Smouse PE (2012) Gen AlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539
Pour-Aboughadareh A, Mohmoudi M, Ahmadi J, Moghaddam M, Mehrabi AA, Alavikia SS (2017) Agro-morphological and molecular variability in Triticum boeoticum accessions from Zagros Mountains, Iran. Genetic Resources and Crop Evolution 64:545–556
Pour-Aboughadareh A, Poczai P, Etminan A, Jadidi O, Kianersi F, Shooshtari L (2022) An analysis of genetic variability and population structure in wheat germplasm using microsatellite and gene-based markers. Plants 11:1205
Prasad P, Bhardwaj SC, Khan H, Gangwar OP, Kumar S, Singh SB (2016) Ug99: saga, reality and status. Current Science 110:1614–1616
Prasad P, Savadi S, Bhardwaj SC, Gupta PK (2020) The progress of leaf rust research in wheat. Fungal Biology 124:537–550
Prasad P, Bhardwaj SC, Thakur RK, Adhikari S, Gangwar OP, Lata C, Kumar S (2021a) Prospects of climate change effects on crop diseases with particular reference to wheat. Journal of Cereal Research 13:117–134
Prasad P, Khan H, Bhardwaj SC, Savadi S, Gangwar OP, Kumar S (2021b) Practical manual on protocols and methodologies in wheat rust research. Directorate of knowledge management in agriculture. ICAR, New Delhi, p 76
Prasad P, Thakur R, Bhardwaj SC, Savadi S, Gangwar OP, Lata C, Adhikari S, Kumar S, Kundu S, Sharma Manjul A, Prakasha TL, Navathe S, Hegde GM, Game BC, Mishra KK, Khan H, Gupta V, Mishra CN, Kumar S, Kumar S, Singh G (2023) Virulence and genetic analysis of Puccinia graminis tritici in the Indian sub-continent from 2016 to 2022 and evaluation of wheat varieties for stem rust resistance. Frontiers in Plant Science 14:1196808
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Randhawa MS, Bains NS, Sohu VS, Chhuneja P, Trethowan RM, Bariana HS, Bansal U (2019) Marker assisted transfer of stripe rust and stem rust resistance genes into four wheat cultivars. Agronomy 9(9):497
Roelfs AP (1988) Genetic control of phenotypes in wheat stem rust. Annual Review of Phytopathology 26:351–367
Roelfs AP, Singh RP, Saari EE (1992) Rust disease of wheat: concepts and methods of disease management. CIMMYT, Mexico, pp 1–81
Roussel V, Koenig J, Beckert M, Balfourier F (2004) Molecular diversity in French bread wheat accessions related to temporal trends and breeding programmes. Theoretical and Applied Genetics 108:920–930
Salem KF, Röder MS, Börner A (2015) Assessing genetic diversity of Egyptian hexaploid wheat (Triticum aestivum L.) using microsatellite markers. Genetic Resources and Crop Evolution 62:377–385
Stakman EC, Stewart DM, Loegering WQ (1962) Identification of physiologic races of Puccinia graminis var tritici United States Department of Agriculture. Agricultural Research Service E617. Bull 53:1–53
Tahir O, Bangash SA, Ibrahim M, Shahab S, Khattak SH, Ud Din I, Khan MN, Hafeez A, Wahab S, Ali B, Makki RM (2023) Evaluation of agronomic performance and genetic diversity analysis using simple sequence repeats markers in selected wheat lines. Sustainability 15:293
Türkoğlu A, Haliloğlu K, Mohammadi SA, Öztürk A, Bolouri P, Özkan G, Bocianowski J, Pour-Aboughadareh A, Jamshidi B (2023) Genetic Diversity and Population Structure in Türkiye Bread Wheat Genotypes Revealed by Simple Sequence Repeats (SSR) Markers. Genes 14:1182
Tyagi S, Kumar A, Gautam T, Pandey R, Rustgi S, Mir RR (2021) Development and use of miRNA-derived SSR markers for the study of genetic diversity, population structure, and characterization of genotypes for breeding heat tolerant wheat varieties. PLoS ONE 16:e0231063
Ya N, Raveendar S, Bayarsukh N, Ya M, Lee JR, Lee KJ, Shin MJ, Cho GT, Ma KH, Lee G (2017) Genetic diversity and population structure of Mongolian wheat based on SSR markers: implications for conservation and management. Plant Breed Biotechnol 5(3):213–20
ZareiAbbasabad E, Mohammadi SA, Moghaddam M, Jalal Kamali MR (2016) Analysis of genetic diversity, population structure and linkage disequilibrium in Iranian wheat landraces using SSR markers. Plant Genetic Resources 1:1–8
Zheng S, Li Y, Lu L, Liu Z, Zhang C, Ao D, Li L, Zhang C, Liu R, Luo C, Wu Y (2017) Evaluating the contribution of Yr genes to stripe rust resistance breeding through marker-assisted detection in wheat. Euphytica 213:1–6
Acknowledgments
The authors wish to acknowledge the Director, ICAR-Indian Institute of Wheat and Barley Research, Karnal, Haryana for extending infrastructure facility for carrying out the research work.
Author information
Authors and Affiliations
Contributions
Conception or design of the work (SA, SCB, OPG, PP, and CL), Data collection (SA, PP, SB), Data analysis and interpretation (SA, SCB, OPG, PP, CL, SB, and GC), Drafting the article (SA, SCB, OPG, PP, CL, and GC). All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
There is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Adhikari, S., Bhardwaj, S.C., Gangwar, O.P. et al. Morphological characterization and molecular diversity assessment of rust resistant genetic stocks of wheat. Trop. plant pathol. (2024). https://doi.org/10.1007/s40858-024-00650-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40858-024-00650-8