Abstract
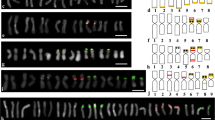
Representatives of the Cactaceae subfamilies Pereskioideae and Opuntioideae from northeastern Brazil were studied using banding with the fluorochromes, CMA3 and DAPI, as well as with fluorescent in situ hybridization using 45S and 5S rDNA probes to identify the distributions of their heterochromatin and rDNA sites. Pereskia aculeata, P. bahiensis, P. grandifolia (Pereskioideae), Brasilopuntia brasiliensis, Tacinga funalis, and T. palmadora showed 2n = 22, while Opuntia dillenii showed 2n = 44, and O. ficus-indica 2n = 88. The karyotypes of all of the species were symmetric, with average chromosome lengths varying from 1.94 µm in O. dillenii to 3.17 µm in P. aculeata. One pair of terminal CMA+ bands corresponding to NORs occurred in all of the diploid cytotypes (except O. ficus-indica, which has two pairs of terminal CMA+ bands) as well as in O. dillenii (tetraploid). CMA+ bands were also observed in the interstitial region of the long arm of a chromosome pair in B. brasiliensis, while a number of variable proximal bands were observed on three chromosome pairs in O. dillenii and on most of the chromosomes of O. ficus-indica. The 45S rDNA sites corresponded to the terminal CMA+ bands, while the 5S rDNA sites were located in the interstitial regions of the long arms of the chromosome pairs of P. aculeata, P. bahiensis, P. grandifolia, and B. brasiliensis. Our data, and earlier publications, suggest that the subfamily Opuntioideae can be characterized as having proximal/interstitial CMA+ heterochromatin in at least one chromosome pair (except in Tacinga). The absence of proximal heterochromatic bands, however, appears to be a synapomorphy of the basal lineages of Cactaceae (subfamily Pereskioideae + Maihuenioideae), suggesting that karyotypes with heterochromatin restricted to the terminal region of a chromosome pair (45S rDNA) represent a plesiomorphic character of the family.



Similar content being viewed by others
References
Anderson EF (2001) The Cactus family. Timber Press, Portland
Ansari HA, Ellison NW, Reader SM, Badaeva ED, Friebe B, Miller TE, Williams WM (1999) Molecular cytogenetic organization of 5S and 18S-26S rDNA loci in white clover (Trifolium repens L.) and related species. Ann Bot 83:199–206
Baker MA, Rebman JP, Parfitt BD, Pinkava DJ, Zimmerman AD (2009) Chromosome numbers in some cacti of Western North America-VIII. Haseltonia 15:117–134
Bandyopadhyay B, Sharma A (2000) The use of multivariate analysis of karyotypes to determine relationships between species of Opuntia. Caryologia 53:121–126
Bárcenas RT, Yesson C, Hawkins JA (2011) Molecular systematics of the Cactaceae. Cladistics 27:470–489
Barros e Silva AE, Guerra M (2010) The meaning of DAPI bands observed after C-banding and FISH procedures. Biotech Histochem 85:115–125
Carvalho R, Soares Filho WS, Brasileiro-Vidal AC, Guerra M (2005) The relationships among lemons, limes and citron: a chromosomal comparison. Cytogenet Genome Res 109:276–282
Castro JP, Souza LGR, Alves LF, Silva AEB, Guerra M, P. Felix LP (2013) In Marhold K (ed) IAPT/IOPB chromosome data 15. Taxon 62:1073
Cota JH, Philbrick CT (1994) Chromosome number variation and polyploidy in the genus Echinocereus (Cactaceae). Am J Bot 81:1054–1062
Cota JH, Wallace RS (1995) Karyotypic studies in the Echinocerus (Cactaceae) and their taxonomic significance. Caryologia 48:105–122
Das AB, Mohanty S (2006) Karyotype analysis and in situ nuclear DNA content in seven species of Echinopsis Zucc. of the family Cactaceae. Cytologia 71:75–79
Edwards EJ, Donoghue MJ (2005) Pereskia and the origin of the cactus life-form. Am Nat 167:777–793
Feitoza LL, Felix LP, Castro AAJL, Carvalho R (2009) Cytogenetics of Alismatales s.s.: chromosomal evolution and C-banding. Plant Syst Evol 280:119–131
Goldblatt P, Johnson ED (2006) Index to plant chromosomes numbers 2001–2003. Monographs, Missouri Botanical Garden
Hubbell HR (1985) Silver staining as an indicator of active ribosomal genes. Stain Technol 60:285–294
Hunt D, Taylor N, Charles G (2006) The new cactus lexicon, vols I, II reprint. DH Books, Melborne
Lan T, Albert VA (2011) Dynamic distribution patterns of ribosomal DNA and chromosomal evolution in Paphiopedilum, a lady’s slipper orchid. BMC Plant Biol. doi:10.1186/1471-2229-11-126
Las Peñas ML, Bernardello G, Kiesling R (2008) Karyotypes and fluorescent chromosome banding in Pyrrhocactus (Cactaceae). Plant Syst Evol 272:211–222
Las Peñas ML, Urdampilleta JD, Bernardello G, Forni-Martins ER (2009) Karyotypes, heterochromatin, and physical mapping of 18S-26S rDNA in Cactaceae. Cytogenet Genome Res 124:72–80
Las Peñas ML, Kiesling R, Bernardello G (2011) Karyotype, heterochromatin, and physical mapping of 5S and 18-5.8-26S rDNA genes in Setiechinopsis (Cactaceae), an Argentine endemic genus. Haseltonia 16:83–90
Las Peñas ML, Urdampilleta JD, López-Carro B, Santiñaque F, Kiesling R, Bernardello G (2014) Classical and molecular cytogenetics and DNA content in Maihuenia and Pereskia (Cactaceae). Plant Syst Evol 300:549–558
Lombello RA, Forni-Martins ER (1998) Cytological studies in climbers of Brazilian forest reserve. Cytologia 63:415–420
Moreno NC, Amarilla LD, Las Peñas ML, Bernardello G (2015) Molecular cytogenetic insights into the evolution of the epiphytic genus Lepismium (Cactaceae) and related genera. Bot J Linn Soc 177:263–277
Moscone EA, Loidl J, Ehrendorfer F, Hunziker AT (1995) Analysis of active nucleolus organizing regions in Capsicum (Solanaceae) by silver staining. Am J Bot 82:276–287
Nyffeler R (2002) Phylogenetic relationships in the Cactus family (Cactaceae) based on evidence from trnK/matK and trnL-trnF sequences. Am J Bot 89:312–326
Oliveira IG, Moraes AP, Almeida EM, Assis FNM, Cabral JS, Barros F, Felix LP (2015) Chromosomal evolution in Pleurothallidinae (Orchidaceae: Epidendroideae) with an emphasis on the genus Acianthera: chromosome numbers and heterochromatin. Bot J Linn Soc 178:102–120
Palomino G, Socorro ZL, Scheinvar L (1988) Estudios citogenéticos de dos especies y una variedad del género Nyctocereus (Cactaceae). Bol Soc Bot Mex 48:75–80
Parfitt BD (1987) Echinocereus nicholii (L. Benson) Cactaceae. Phytologia 63:157–158
Pedrosa A, Guerra M, Soares-Filho WS (1997) An hierarchy of activation of nucleolar organizer regions in Citrus sinensis (L.) Osbeck. Cytobios 92:43–51
Pedrosa A, Sandal N, Stougaard J, Schweizer D, Bachmair A (2002) Chromosomal map of the model legume Lotus japonicus. Genetics 161:1661–1672
Pedrosa-Harand A, de Almeida CC, Mosiolek M, Blair MW, Schweizer D, Guerra M (2006) Extensive ribosomal DNA amplification during Andean common bean (Phaseolus vulgaris L.) evolution. Theor Appl Genet 112:924–933
Peruzzi L, Eroğlu HE (2013) Karyotype asymmetry: again, how to measure and what to measure? Comp Cytogenet 7:1–9
Pinkava DJ, Mcleod MG (1971) Chromosome numbers in some cacti of western North America. Brittonia 23:171–176
Pinkava DJ, McGill LA, Reeves T (1977) Chromosome number in some cacti of western North America. J Torrey Bot Soc 104:105–110
Pinkava DJ, Baker MA, Parfitt BD (1985) Chromosome numbers in some cacti of western North America. V. Syst Bot 10:471–483
Pinkava DJ, Parfitt BD, Baker MA, Worthington RD (1992) Chromosome numbers in some cacti of western North America. VI. Madroño 39:98–113
Romero Zarco C (1986) A new method for estimating karyotype asymmetry. Taxon 35:526–530
Ross R (1981) Chromosome counts, cytology, and reproduction in the Cactaceae. Am J Bot 68:463–470
Stebbins GL (1971) Chromosomal evolution in higher plants. E. Arnold, London
Stevens PF (2015) Angiosperm phylogeny website. http://www.mobot.org/MOBOT/research/APweb/. Accessed 07 July 2015
Sumner AT (2003) Chromosomes: organization and function. Blackwell, North Berwick
Taylor N, Santos MR, Larocca J, Zappi D (2015) Lista de Espécies da Flora do Brasil—Cactaceae. Jardim Botânico do Rio de Janeiro. http://floradobrasil.jbrj.gov.br/jabot/floradobrasil/FB70. Accessed 27 Feb 2015
Wallace RS (1995) Molecular systematic study of the Cactaceae: using chloroplast DNA variation to elucidate cactus phylogeny. Bradleya 13:1–12
Zurita F, Jiménez R, Guardia RD, Burgos M (1999) The relative rDNA content of a NOR determines its level of expression and its probability of becoming active. A sequential silver staining and in situ hybridization study. Chromosome Res 7:563–570
Acknowledgments
The authors thank CNPQ (Conselho Nacional de Desenvolvimento Científico e Tecnológico) for the grant awarded to the last author; Professor Marcelo Guerra for allowing us to use the installations of the Laboratório de Citogenética e Evolução Vegetal at Universidade Federal de Pernambuco, which was critical to the realization of the present work; and the anonymous referee whose suggestions helped improve this paper.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Castro, J.P., Medeiros-Neto, E., Souza, G. et al. CMA band variability and physical mapping of 5S and 45S rDNA sites in Brazilian Cactaceae: Pereskioideae and Opuntioideae. Braz. J. Bot 39, 613–620 (2016). https://doi.org/10.1007/s40415-015-0248-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-015-0248-5