Abstract
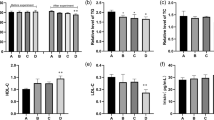
Herein, we described the physicochemical properties of deproteinized extract of calf blood(DECB) and established a hypoxia model treated with or without DECB to detect the sugar, lactic acid, protein, and ATP contents of mice and then identified and analyzed the differentially expressed genes between two groups using mRNA expression chip. According to the results of the airtight hypoxia experiment, mice in the model+DECB group had a significantly prolonged time of hypoxia tolerance compared with the model group. The biochemical test indicated that DECB could significantly increase the level of sugar, ATP and proteins and reduce the amount of lactic acid in mice. It also revealed that Hmgcs2, Cpt1a, Angptl4, Cyp8b1, and Ehhadh genes were involved in mice energy metabolism, and were closely associated with metabolic signaling pathway. These results suggest that DECB might be a potential drug to treat metabolic diseases. Among the genes with differential expression under hypoxia, Angptl4, Cyp8b1, and Ehhadh were critical factors for sugar metabolism. Hmgcs2 provided energy directly, and Cptla regulated cellular inflammatory responses, promoting energy metabolism.
Similar content being viewed by others
References
Wu W., Zeng L. N., Peng Y. Y., Lu X. H., Li C. Y., Wang Z. C., European Review for Medical and Pharmacological Sciences, 2014, 18(22), 3406
Xu G. Y., Xu J. H., Han X., Li H. Y., Yuan G. X., An L. P., Du P. G., International Immunopharmacology, 2018, 56, 212
Xu G. Y., Han X., Yuan G. X., An L. P., Du P. G., PLoS One, 2017, 12(7), e0180899
Chen M. J., Gong L., Qiu X. D., Chinese Journal of Ophthalmology, 2012, 48(12), 1083
Fan C., Wang Y., Zhang Y., Lang L., Deng X., Cheng Y., Chinese Journal of Industrial Hygiene and Occupational Diseases, 2014, 32(12), 924
Huang D. W., Sherman B. T., Tan Q., Collins J. R., Alvord W. G., Roayaei J., Stephens R., Baseler M. W., Lane H. C., Lempicki R. A., Genome Biology, 2007, 8(9), R183
Ashburner M., Ball C. A., Blake J. A., Botstein D., Butler H., Cherry J. M., Davis A. P., Dolinski K., Dwight S. S., Eppig J. T., Harris M. A., Hill D. P., Issel-Tarver L., Kasarskis A., Lewis S., Matese J. C., Richardson J. E., Ringwald M., Rubin G. M., Sherlock G., The Gene Ontology Consortium Nature Genetics, 2000, 25(1), 25
Shi Q., Zhou W., Chen C., Zhang B. Y., Xiao K., Zhang X. C., Shen X. J., Li Q., Deng L. Q., Dong J. H., Lin W. Q., Huang P., Jiang W. J., Lv J., Han J., Dong X. P., PLoS One, 2015, 10(10), e0139552
Irwin D. M., Tan H., Molecular Phylogenetics and Evolution, 2015, 70, 195
Zhang T. T., Zhang G. M., Jin Y. H., Guo Y. X., Wang Z., Fan Y. X., El-Samahy M. A., Wang F., Tissue & Cell, 2017, 49(5), 603
Helenius T. O., Misiorek J. O., Nystrom J. H., Fortelius L. E., Habte-zion A., Liao J., Asghar M. N., Zhang H., Azhar S., Omary M. B., Toivola D. M., Molecular Biology of the Cell, 2015, 26(12), 2298
Kang D. Y., Nipin S. P., Darvin P., Joung Y. H., Byun H. J., Do C. H., Park K. D., Park M. N., Cho K. H., Yang Y. M., Animal Biotechnology, 2017, 28(3), 189
Gobin S., Bonnefont J. P., Prip-Buus C., Mugnier C., Ferrec M., Demaugre F., Saudubray J. M., Rostane H., Djouadi F., Wilcox W., Cederbaum S., Haas R., Nyhan W. L., Green A., Gray G., Girard J., Thuillier L., Human Genetics, 2002, 111(2), 179
Diaz-Rua R., Palou A., Oliver P., Food & Nutrition Research, 2016, 60, 33554
Napal L., Marrero P. F., Haro D., Journal of Molecular Biology, 2005, 354(4), 751
Gao X., Li K., Hui X., Kong X., Sweeney G., Wang Y., Xu A., Teng M., Liu P., Wu D., The Biochemical Journal, 2011, 435(3), 723
Kim L., Kim H. G., Kim H., Kim H. H., Park S. K., Uhm C. S., Lee Z. H., Koh G. Y., Biochem. J., 2000, 346, 603
Ito Y., Oike Y., Yasunaga K., Hamada K., Miyata K., Matsumoto S., Sugano S., Tanihara H., Masuho Y., Suda T., Cancer Research, 2003, 63(20), 6651
Janssen A. W. F., Dijk W., Boekhorst J., Kuipers F., Groen A. K., Lukovac S., Hooiveld G., Kersten S., Biochimica et Biophysica Acta, 2017, 1862(10), 1056
Bertaggia E., Jensen K. K., Castro-Perez J., Xu Y., di Paolo G., Chan R. B., Wang L., Haeusler R. A., American Journal of Physiology Endocrinology and Metabolism, 2017, 313(2), E121
Mork L. M., Strom S. C., Mode A., Ellis E. C., Journal of Clinical and Experimental Hepatology, 2016, 6(2), 87
Watanabe M., Morimoto K., Houten S. M., Kaneko-Iwasaki N., Sugizaki T., Horai Y., Mataki C., Sato H., Murahashi K., Arita E., Schoonjans K., Suzuki T., Itoh H., Auwerx J., PLoS One, 2012, 7(8), e38286
Houten S. M., Denis S., Argmann C. A., Jia Y., Ferdinandusse S., Reddy J. K., Wanders R. J., Journal of Lipid Research, 2012, 53(7), 1296
Rosen M. B., Das K. P., Wood C. R., Wolf C. J., Abbott B. D., Lau C., Toxicology, 2013, 308, 129
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supported by the Natural Science Foundation of Jilin Province, China(No.20180101099JC) and the “Thirteen Five” Science and Technology Project of the Department of Education of Jilin Province, China (No. JJKH20180372KJ).
Rights and permissions
About this article
Cite this article
Zhou, T., Xu, G., Sun, L. et al. Improving Energy Metabolism of Deproteinized Extract of Calf Blood Through Regulation of Hmgcs2, Cpt1a, Angptl4, Cyp8b1, and Ehhadh Genes in Mice. Chem. Res. Chin. Univ. 35, 427–433 (2019). https://doi.org/10.1007/s40242-019-9021-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40242-019-9021-9