Abstract
Purpose
The pathogenesis of diabetes is considered polygenic as a result of complex interactions between genetic/epigenetic and environmental factors. This review intended to evaluate the scientometric and knowledge gap of diabetes genetics researches conducted in Iran as a case of developing countries, and drawn up a roadmap for future studies.
Methods
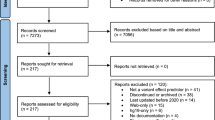
We searched Scopus and PubMed databases from January 2015 until December 2019 using the keywords: (diabetes OR diabetic) AND (Iran). All publications were reviewed by two experts and after choosing relevant articles, they were categorized based on the subject, level of evidence, study design, publication year, and type of genetic studies.
Results
Of 10,540 records, 428 articles were met the inclusion criteria. Generally, the number of researches about diabetes genetics rose since 2015. Case–control/cross-sectional and animal studies were the common types of study design and based on the subject, the most frequent researches were about genetic factors involved in diabetes development (38%). Briefly, the top seven genes that were evaluated for T2DM were TCF7L2, APOAII, FTO, PON1, ADIPOQ, MTHFR, and PPARG respectively, and also, CTL4 for T1DM. miR-21, miR-155, and miR-375 respectively were the most micro-RNAs that were evaluated. Furthermore, there were six studies about lncRNAs.
Discussion and Conclusion
Investigation about the genetic of diabetes is progressed although there are some limitations like non-homogenous data from Iran, heterogeneity of ethnicity, and rationale of studies. Compared to the previous analysis in Iran, still, GWAS and large-scale studies are required to achieve better policies for manage and control of diabetes disease.



Similar content being viewed by others
Data availability
The datasets used and analyzed during the current study are available from the corresponding author.
References
Zheng Y, Ley SH, Hu FB. Global aetiology and epidemiology of type 2 diabetes mellitus and its complications. Nat Rev Endocrinol. 2018;14(2):88–98.
Peykari N, Djalalinia S, Kasaeian A, Naderimagham S, Hasannia T, Larijani B, et al. Diabetes research in Middle East countries; a scientometrics study from 1990 to 2012. J Res Med Sci. 2015;20(3):253–62.
King H, Aubert RE, Herman WH. Global burden of diabetes, 1995–2025: prevalence, numerical estimates, and projections. Diabetes Care. 1998;21(9):1414–31.
International Diabetes Federation. IDF Diabetes Atlas Ninth. Dunia: IDF (2019).
Javanbakht M, Mashayekhi A, Baradaran HR, Haghdoost A, Afshin A. Projection of diabetes population size and associated economic burden through 2030 in Iran: evidence from micro-simulation Markov model and Bayesian meta-analysis. PLoS one. 2015;10(7):e0132505.
Esteghamati A, Larijani B, Aghajani MH, Ghaemi F, Kermanchi J, Shahrami A, et al. Diabetes in Iran: prospective analysis from first nationwide diabetes report of National Program for Prevention and Control of Diabetes (NPPCD-2016). Sci Rep. 2017;7(1):13461.
American Diabetes Association. 2. Classification and diagnosis of diabetes: standards of medical care in diabetes—2020. Diabetes care. 2020 Jan 1;43(Supplement 1):S14–31.
Fuchsberger C, Flannick J, Teslovich TM, Mahajan A, Agarwala V, Gaulton KJ, et al. The genetic architecture of type 2 diabetes. Nature. 2016;536(7614):41–7.
Imamura M, Maeda S. Genetics of type 2 diabetes: the GWAS era and future perspectives. Endocrine journal. 2011;1107190592.
Imamura M, Horikoshi M, Maeda S. Genome-wide association study for type 2 diabetes. In Genome-Wide Association Studies 2019 (pp. 49-86). Springer, Singapore.
Bakay M, Pandey R, Grant SFA, Hakonarson H. The genetic contribution to type 1 diabetes. Curr DiabRep. 2019;19(11):116.
Suzuki K, Akiyama M, Ishigaki K, Kanai M, Hosoe J, Shojima N, et al. Identification of 28 new susceptibility loci for type 2 diabetes in the Japanese population. Nat Genet. 2019;51(3):379–86.
Kulkarni M, Leszczynska A, Wei G, Winkler MA, Tang J, Funari VA, et al. Genome-wide analysis suggests a differential microRNA signature associated with normal and diabetic human corneal limbus. Sci Rep. 2017;7(1):1–15.
Takematsu E, Spencer A, Auster J, Chen P-C, Graham A, Martin P, et al. Genome wide analysis of gene expression changes in skin from patients with type 2 diabetes. PLoS One. 2020;15(2):e0225267.
Staiger H, Schaeffeler E, Schwab M, Häring H-U. Pharmacogenetics: implications for modern type 2 diabetes therapy. Rev Diabet Stud. 2015;12(3–4):363.
Pearson E. Personalized medicine in diabetes: the role of ‘omics’ and biomarkers. Diabet Med. 2016;33(6):712–7.
Nair AK, Baier LJ. Complex genetics of type 2 diabetes and effect size: what have we learned from isolated populations? Rev Diabet Stud. 2015;12(3–4):299.
Goto A, Noda M, Goto M, Yasuda K, Mizoue T, Yamaji T, et al. Predictive performance of a genetic risk score using 11 susceptibility alleles for the incidence of Type 2 diabetes in a general Japanese population: a nested case–control study. Diabet Med. 2018;35(5):602–11.
Chan JC, Malik V, Jia W, Kadowaki T, Yajnik CS, Yoon K-H, et al. Diabetes in Asia: epidemiology, risk factors, and pathophysiology. JAMA. 2009;301(20):2129–40.
Fatemeh B, Maryam O, Farideh R, Ensieh N-E, Saeedeh S, Bagher L. Iran Diabetes Research Roadmap (IDRR) study knowledge gap in ge-netic research on diabetes mellitus in Iran: a Review Article. Iran J Public Health. 2017;46(Supple 1):53–9.
Saeedi P, Petersohn I, Salpea P, Malanda B, Karuranga S, Unwin N, et al. IDF Diabetes Atlas Committee Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: results from the International Diabetes Federation Diabetes Atlas. Diabetes Res Clin Pract. 2019;157:107843.
WHO. Global report on diabetes 2016. Geneva: World Health Organization; 2016.
Johnson GC, Todd JA. Strategies in complex disease mapping. Curr Opin Genet Dev. 2000;10(3):330–4.
Risch NJ. Searching for genetic determinants in the new millennium. Nature. 2000;405(6788):847–56.
Collins FS, Guyer MS, Chakravarti A. Variations on a theme: cataloging human DNA sequence variation. Science. 1997;278(5343):1580–1.
Abuhendi N, Qush A, Naji F, Abunada H, Al Buainain R, Shi Z, et al. Genetic polymorphisms associated with type 2 diabetes in the Arab world: a systematic review and meta-analysis. Diabetes Res Clin Pract. 2019;151:198–208.
Yako YY, Guewo-Fokeng M, Balti EV, Bouatia-Naji N, Matsha TE, Sobngwi E, et al. Genetic risk of type 2 diabetes in populations of the African continent: a systematic review and meta-analyses. Diabetes Res Clin Pract. 2016;114:136–50.
Asemi-Esfahani Z, Alian F, Fazilati M, Nazem H, Palizban A. Investigating the relationship between rs7903146 polymorphism genotypes of TCF7L2 gene and visfatin serum level in patients with type 2 diabetes. J Biol Today’s World. 2017;6(4):67–74.
Burton PR, Clayton DG, Cardon LR, Craddock N, Deloukas P, Duncanson A, et al. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature. 2007;447(7145):661–78.
Go MJ, Lee Y, Park S, Kwak SH, Kim B-J, Lee J. Genetic-risk assessment of GWAS-derived susceptibility loci for type 2 diabetes in a 10 year follow-up of a population-based cohort study. J Hum Genet. 2016;61(12):1009–12.
Khodaeian M, Enayati S, Tabatabaei-Malazy O, Amoli MM. Association between genetic variants and diabetes mellitus in Iranian populations: a systematic review of observational studies. J Diabet Res. 2015;2015:585917.
He Y, Ding Y, Liang B, Lin J, Kim T-K, Yu H, et al. A systematic study of dysregulated microRNA in type 2 diabetes mellitus. Int J Mol Sci. 2017;18(3):456.
Kaur P, Kotru S, Singh S, Behera BS, Munshi A. Role of miRNAs in the pathogenesis of T2DM, insulin secretion, insulin resistance, and β cell dysfunction: the story so far. J Physiol Biochem. 2020;76(4):485–502.
Leung A, Natarajan R. Long noncoding RNAs in diabetes and diabetic complications. Antioxid Redox Signal. 2018;29(11):1064–73.
Zhang W, Zheng J, Hu X, Chen L. Dysregulated expression of long noncoding RNAs serves as diagnostic biomarkers of type 2 diabetes mellitus. Endocrine. 2019;65(3):494–503.
Zhu H, Bian X, Gong J, Yu P, Lu H. Long noncoding RNAs as novel biomarkers for Type 2 diabetes. Biomark Med. 2020;14(15):1501–11.
Taheri M, Eghtedarian R, Dinger ME, Ghafouri-Fard S. Emerging roles of non-coding RNAs in the pathogenesis of type 1 diabetes mellitus. Biomed Pharmacother. 2020;129:110509.
Fodor A, Cozma A, Suharoschi R, Sitar-Taut A, Roman G. Clinical and genetic predictors of diabetes drug’s response. Drug Metab Rev. 2019;51(4):408–27.
Author information
Authors and Affiliations
Contributions
All authors contributed and approved the paper before submission. FR and EN planned the study, ShE did the databases search, SE and FE screened and classified the result which approved by FR and FB. MA contributed in supervision of review, SE and FE prepared the manuscript that approved by all named authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fana, S.E., Esmaeili, F., Esmaeili, S. et al. Knowledge discovery in genetics of diabetes in Iran, a roadmap for future researches. J Diabetes Metab Disord 20, 1785–1791 (2021). https://doi.org/10.1007/s40200-021-00838-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40200-021-00838-8