Abstract
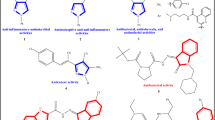
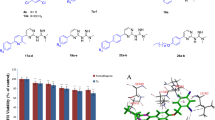
Thiazoles are an important class of heterocyclic compounds that possess a sulfur and nitrogen containing five-membered ring, which acts as a pharmacophore, and show a wide range of complex biological activities. A series of sixteen 2-arylidenehydrazinyl-4-arylthiazole analogues (3a–p) were evaluated for cytotoxic activity against brine shrimp (Artemia salina) nauplii and their minimum inhibitory concentrations were determined against two Gram-positive (Listeria monocytogenes and Enterococcus faecalis) and two Gram-negative bacterial strains (C. sakazakii and E. coli). Of the tested compounds, 3g demonstrated highest cytotoxicity with a LC50 value of 54 ppm followed by compound 3h (LC50 = 85 ppm), in a short-term bioassay using A. salina, whereas compound 3i exhibited the most potent antibacterial activities against L. monocytogenes, E. faecalis, and C. sakazakii with MIC values ranging from 50 to 100 μg mL−1. Compound 3g showed highest antibacterial activity against E. coli (MIC = 50 μg mL−1). In silico drug-likeness, pharmacokinetic (ADME) properties, toxicity effects, and drug scores were also evaluated, and none of the sixteen compounds were found to violate Lipiniski’s rule of five or Veber’s rule, indicating potential for development as oral drug candidates. In addition, a docking study of compound 3i into the active site of E. coli FabH receptor, an attractive target for the development of new antibacterial agents, showed it has good binding properties.








Similar content being viewed by others
References
Alam MS, Liu L, Lee YE, Lee DU (2011) Synthesis, antibacterial activity and quantum-chemical studies of novel 2-arylidenehydrazinyl-4-arylthiazole analogues. Chem Pharm Bull 59:568–573
Alam MS, Ahmed JU, Lee DU (2014a) Synthesis, antibacterial, antioxidant activity and QSAR studies of novel 2-arylidenehydrazinyl-4-arylthiazole analogues. Chem Pharm Bull 62:1259–1268
Alam MS, Lee DU, Bari ML (2014b) Antibacterial and cytotoxic activities of Schiff base analogues of 4-aminoantipyrine. J Korean Soc Appl Biol Chem 57:613–619
Ban M, Taguchi H, Katsushima T, Takahashi M, Shinoda K, Watanabe A, Tominaga T (1998) Novel antiallergic and antiinflammatory agents. Part I: synthesis and pharmacology of glycolic amide derivatives. Bioorg Med Chem 6:1069–1076
Cheng K, Xue JY, Zhu HL (2013) Design, synthesis and antibacterial activity studies of thiazole derivatives as potent ecKAS III inhibitors. Bioorg Med Chem Lett 23:4235–4238
Chimenti F, Maccioni E, Secci D, Bolasco A, Chimenti P, Granese A, Befani O, Turini P, Alcaro S, Ortuso F, Cardia MC, Distinto S (2007) Selective inhibitory activity against MAO and molecular modeling studies of 2-thiazolylhydrazone derivatives. J Med Chem 50:707–712
Clark RF, Zhang T, Wang X, Wang R, Zhang X, Camp HS, Beutel BA, Sham HL, Gu YJ (2007) Phenoxy thiazole derivatives as potent and selective acetyl-CoA carboxylase 2 inhibitors: modulation of isozyme selectivity by incorporation of phenyl ring substituents. Bioorg Med Chem Lett 17:1961–1965
Cohen A, Verhaeghe P, Crozet MD, Hutter S, Rathelot P, Vanelle P, Azas N (2012) Tandem synthesis and in vitro antiplasmodial evaluation of new naphtho[2,1-d] thiazole derivatives. Eur J Med Chem 55:315–324
Deb PK, Kaur R, Chandrasekaran B, Bala M, Gill D, Kaki VR, Akkinepalli RR, Mailavaram R (2014) Synthesis, anti-inflammatory evaluation, and docking studies of some new thiazole derivatives. Med Chem Res 23:2780–2792
Dewar MJS, Zoebisch EG, Healy EF, Stewart JJP (1985) Development and use of quantum molecular models 75. Comparative tests of theoretical procedures for studying chemical reactions. J Am Chem Soc 107:3902–3909
El-Sabbagh OI, Baraka MM, Ibrahim SM, Pannecouque C, Andrei G, Snoeck R, Balzarini J, Rashad AA (2009) Synthesis and antiviral activity of new pyrazole and thiazole derivatives. Eur J Med Chem 44:3746–3753
Ertl P, Rohde B, Selzer P (2000) Fast calculation of molecular polar surface area (PSA) as a sum of fragment-based contributions and its application to the prediction of drug transport properties. J Med Chem 43:3714–3717
Finney DJ (1978) Statistical method in biological assay, 3rd edn. Charles Griffin and Company, London, p 661
Hartl M, Humpf HU (2000) Toxicity assessment of fumonisins using the brine shrimp (Artemia salina) bioassay. Food Chem Toxicol 38:1097–1102
Heath RJ, Rock CO (1996) Regulation of fatty acid elongation and initiation by acyl-acyl carrier protein in Escherichia coli. J Biol Chem 271:1833–1836
Helal MHM, Salem MA, El-Gaby MSA, Aljahdali M (2013) Synthesis and biological evaluation of some novel thiazole compounds as potential anti-inflammatory agents. Eur J Med Chem 65:517–526
Iino T, Hashimoto N, Sasaki K, Ohyama S, Yoshimoto R, Hosaka H, Hasegawa T, Chiba M, Nagata Y, Nishimura JET (2009) Structure–activity relationships of 3,5-disubstituted benzamides as glucokinase activators with potent in vivo efficacy. Bioorg Med Chem 17:3800–3809
Jain A, Pandey V, Ganeshpurkar A, Dubey N, Bansal D (2015) Formulation and characterization of floating microballoons of nizatidine for effective treatment of gastric ulcers in murine model. Drug Deliv 22:306–311
Khandekar SS, Daines RA, Lonsdale JT (2003) Bacterial β-ketoacyl-Acyl carrier protein synthases as targets for antibacterial agents. Curr Protein Pept Sci 4:21–29
Lee JY, Jeong KW, Shin S, Lee JU, Kim Y (2012) Discovery of novel selective inhibitors of Staphylococcus aureus β-ketoacyl acyl carrier protein synthase III. Eur J Med Chem 47:261–269
Li JR, Li DD, Wang RR, Sun J, Dong JJ, Du QR, Fang F, Zhang WM, Zhu HL (2014) Design and synthesis of thiazole derivatives as potent FabH inhibitors with antibacterial activity. Eur J Med Chem 75:438–447
Lipinski CA, Lombardo L, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26
Lv PC, Wang KR, Yang Y, Mao WJ, Chen J, Xiong J, Zhu HL (2009) Design, synthesis and biological evaluation of novel thiazole derivatives as potent FabH inhibitors. Bioorg Med Chem Lett 19:6750–6754
Mayer BN, Ferrigni NR, Putnam JE, Jacobsen LB, Nichols DE, McLaughlin JL (1982) Brine shrimp: a convenient general bioassay for active plant constituents. Planta Med 45:31–34
Mayo SL, Olafson BD, Goddard WA (1990) DREIDING: a generic force field for molecular simulations. J Phys Chem 94:8897–8909
Mjambili F, Njoroge M, Naran K, Kock CD, Smith PJ, Mizrahi V, Warner D, Chibale K (2014) Synthesis and biological evaluation of 2-aminothiazole derivatives as antimycobacterial and antiplasmodial agents. Bioorg Med Chem Lett 24:560–564
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Netalkar PP, Netalkar SP, Budagumpi S, Revankar VK (2014) Synthesis, crystal structures and characterization of late first row transition metal complexes derived from benzothiazole core: anti-tuberculosis activity and special emphasis on DNA binding and cleavage property. Eur J Med Chem 79:47–56
Nie Z, Perretta C, Lu J, Su Y, Margosiak S, Gajiwala KS, Cortez J, Nikulin V, Yager KM, Appelt K, Chu S (2005) Structure-based design, synthesis, and study of potent inhibitors of β-ketoacyl-acyl carrier protein synthase III as potential antimicrobial agents. J Med Chem 48:1596–1606
Nishikaku F, Koga Y (1993) Suppression of murine collagen-induced arthritis by treatment with a novel thiazole derivative, SM-8849. Immunopharmacology 25:65–74
Nishina C, Enoki N, Tawata S, Mori A, Kobayashi K, Fukushima M (1987) Antibacterial activity of flavonoids against Staphylococcus epidermidis, a skin bacterium. Agric Biol Chem 51:139–143
Novakova I, Subileau EA, Toegel S, Gruber D, Lachmann B, Urban E, Chesne C, Noe CR, Neuhaus W (2014) Transport rankings of nonsteroidal antiinflammatory drugs across blood–brain barrier in vitro models. PLoS ONE 9:e86806
Popsavin M, Torovic L, Svircev M, Kojic V, Bogdanovic G, Popsavin V (2006) Synthesis and antiproliferative activity of two new tiazofurin analogues with 2′-amido functionalities. Bioorg Med Chem Lett 16:2773–2776
Price AC, Choi KH, Heath RJ, Li Z, White SW, Rock CO (2001) Inhibition of β-ketoacyl-acyl carrier protein synthases by thiolactomycin and cerulenin. J Biol Chem 276:6551–6559
Rahmutulla B, Matsushita K, Satoh M, Seimiya M, Tsuchida S, Kubo S, Shimada H, Ohtsuka M, Miyazaki M, Nomura F (2014) Alternative splicing of FBP-interacting repressor coordinates c-Myc, P27Kip1/cyclinE and Ku86/XRCC5 expression as a molecular sensor for bleomycininduced DNA damage pathway. Oncotarget 15:2404–2417
Romagnoli R, Baraldi PG, Brancale A, Ricci A, Hamel E, Bortolozzi R, Basso G, Viola G (2011) Convergent synthesis and biological evaluation of 2-amino-4-(3′,4′,5′-trimethoxyphenyl)-5-aryl thiazoles as microtubule targeting agents. J Med Chem 54:5144–5153
Sadowski J, Kubinyi H (1998) A scoring scheme for discriminating between drugs and nondrugs. J Med Chem 41:3325–3329
Sarojini BK, Krishna BG, Darshanraj CG, Bharath BR, Manjunatha H (2010) Synthesis, characterization, in vitro and molecular docking studies of new 2,5-dichlorothienyl substituted thiazole derivatives for antimicrobial properties. Eur J Med Chem 45:3490–3496
Sevrioukova IF, Poulos TL (2013) Dissecting cytochrome P450 3A4-ligand interactions using ritonavir analogues. Biochemistry 52:4474–4481
Shih MH, Ying KF (2004) Syntheses and evaluation of antioxidant activity of sydnonyl substituted thiazolidinone and thiazoline derivatives. Bioorg Med Chem 12:4633–4643
Shiradkar MR, Akula KC, Dasari V, Baru V, Chiningiri B, Gandhi S, Kaur R (2007) Clubbed thiazoles by MAOS: a novel approach to cyclin-dependent kinase 5/p25 inhibitors as a potential treatment for Alzheimer’s disease. Bioorg Med Chem 15:2601–2610
Shoichet BK, McGovern SL, Wei B, Irwin JJ (2002) Lead discovery using molecular docking. Curr Opin Chem Bio 6:439–446
Soares MI, Brito AF, Laranjo M, Paixao JA, Botelho MF, Melo TMVDP (2013) Chiral 6,7-bis(hydroxymethyl)-1H,3H-pyrrolo[1,2-c]thiazoles with anti-breast cancer properties. Eur J Med Chem 60:254–262
Styrt B, Rocklin RE, Klempner MS (1985) Inhibition of neutrophil superoxide production by fanetizole. Inflammation 9:233–244
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Turan-Zitouni G, Ozdemir A, Kaplancikli ZA (2011) Synthesis and antiviral activity of some (3,4-diaryl-3H-thiazol-2-ylidene)pyrimidin-2-yl amine derivatives. Phosphorus–Sulphur–Silicon 186:233–239
Wei L, Cheng J, Meng Y, Ren Y, Deng H, Guo Y (2014) A novel formulation of thiamine dilaurylsulphate and its preservative effect on apple juice and sterilised milk. Food Chem 1:415–422
Weidenbörner M, Chandra Jha H (1993) Antifungal activity of flavonoids and their mixtures against different fungi occurring on grain. Pestic Sci 38:347–351
Yang JM, Chen CC (2004) GEMDOCK: a generic evolutionary method for molecular docking. Proteins 55:288–304
Yang YS, Zhang F, Gao C, Zhang YB, Wang XL, Tang JF, Sun J, Gong HB, Zhu HL (2012) Discovery and modification of sulfur-containing heterocyclic pyrazolone derivatives as potential novel class of β -ketoacyl-acyl carrier protein synthase III (FabH) inhibitors. Bioorg Med Chem Lett 22:4619–4624
Ye J, Liu Q, Wang C, Meng Q, Sun H, Peng J, Ma X, Liu K (2013) Benzylpenicillin inhibits the renal excretion of acyclovir by OAT1 and OAT3. Pharmacol Rep 65:505–512
Zablotskaya A, Segal I, Geronikaki A, Tatiana E, Belyakov S, Petrova M, Shestakova I, Zvejniece L, Nikolajeva V (2013) Synthesis, physicochemical characterization, cytotoxicity, antimicrobial, antiinflammatory and psychotropic activity of new N-[1,3-(benzo)thiazol-2-yl]-ω-[3,4-dihydroisoquinolin-2(1H)-yl]alkanamides. Eur J Med Chem 70:846–856
Zhang MQ, Wilkinson B (2007) Drug discovery beyond the ‘rule-of-five’. Curr Opin Biotechnol 18:478–488
Zhang WT, Ruan JL, Wu PF, Jiang FC, Zhang LN, Fang W, Chen XL, Wang Y, Cao BS, Chen GY, Zhu YJ, Gu J, Chen JG (2009) Design, synthesis, and cytoprotective effect of 2-aminothiazole analogues as potent poly(ADP-Ribose) polymerase-1 inhibitors. J Med Chem 52:718–725
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Alam, M.S., Ahmed, J.U. & Lee, DU. Biological features, drug-likeness, pharmacokinetic properties, and docking of 2-arylidenehydrazinyl-4-arylthiazole analogues. Appl Biol Chem 59, 181–192 (2016). https://doi.org/10.1007/s13765-016-0148-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13765-016-0148-9