Abstract
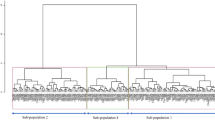
Genetic and fatty acid variability in four datasets of accessions and prebreeding lines of Jatropha curcas was analyzed. The datasets were comprised of J. curcas accessions (13), BC1 (28), BC1F2 (12) and single seeds (12). The BC1, BC1F2 and single seed dataset were derived from an interspecific cross between J. curcas and J. integerrima. The average within-group (within dataset) polymorphism revealed by AFLP markers was 28.5 %. The average genetic similarity within the four datasets, namely, J. curcas accessions, BC1, BC1F2 and single seeds was 0.92, 0.82, 0.90 and 0.92 respectively with an overall average genetic similarity of 0.80. The values of percent polymorphism, Shannon’s information index and expected heterozygosity were highest for BC1 dataset. The total unsaturated fatty acids in the oil varied from 63.64 to 89.10 %. Levene’s test showed that within-group variance was significantly different for three fatty acids, namely, palmitic acid, oleic acid and linoleic acid. BC1F2 dataset had the highest variance for palmitic and oleic acid, whereas BC1 dataset had the highest variance for linoleic acid. The results revealed that both genetic as well as fatty acid variability was significantly higher in the newly created prebreeding lines and the level of fatty acid variability in these datasets was different and highly correlated to the level of their genetic variability as revealed by AFLP markers. This new prebreeding material, especially, the BC1 and BC1F2 lines, will be useful for development of Jatropha varieties with specific fatty acid composition for specific industrial applications.


Similar content being viewed by others
Abbreviations
- AFLP:
-
Amplified fragment length polymorphism
- F1 :
-
Filial 1
- BC1 :
-
Backcross 1
- GC-MS:
-
Gas chromatography–mass spectrometry
- CTAB:
-
Cetyltrimethylammonium bromide
References
Basha SD, George F, Makkar HPS, Becker K, Sujatha M (2009) A comparative study of biochemical traits and molecular markers for assessment of genetic relationships between Jatropha curcas (L.) germplasm from different countries. Plant Sci 176:812–823
Bressan EA, Sebbenn AM, Ferreira RR, Lee TSG, Figueira A (2013) Jatropha curcas L. (Euphorbiaceae) exhibits a mixed mating system, high correlated mating and apomixis. Tree Genet Genomics 9:1089–1097
Carvalho CR, Clarindo WR, Praca MM, Araujo FS, Carels N (2008) Genome size, base composition and karyotype of Jatropha curcas L., an important biofuel plant. Plant Sci 174(6):613–617. doi:10.1016/j.plantsci.2008.03.010
Graef G, LaVallee BJ, Tenopir P, Tat M, Schweiger B, Kinney AJ, Van Gerpen JH, Clemente TE (2009) A high-oleic-acid and low-palmitic-acid soybean: agronomic performance and evaluation as a feedstock for biodiesel. Plant Biotechnol J 7:411–421
Knothe G (2005) Dependence of biodiesel fuel properties on the structure of fatty acid alkyl esters. Fuel Process Technol 86:1059–1070
Kureel RS, Singh CB, Gupta AK, Pandey A (2008) 3rd R&D Report on Tree Borne Oilseeds. National Oilseeds and Vegetable Oils Development Board, Gurgaon, India
Liu P, Wang CM, Li L, Sun F, Yue GH (2011) Mapping QTLs for oil traits and eQTLs for oleosin genes in Jatropha. BMC Plant Biol 11:132. doi:10.1186/1471-2229-11-132
Makkar HPS, Martinez-Herrera J, Becker K (2008) Variations in seed number per fruit, seed physical parameters and contents of oil, protein and phorbol ester in toxic and non-toxic genotypes of Jatropha curcas. J Plant Sci 3:260–265
Meru G, McGregor C (2014) Quantitative trait loci and candidate genes associated with fatty acid content of watermelon seed. J Am Soc Hortic Sci 139(4):433–441
Openshaw K (2000) A review of Jatropha curcas L: an oil plant of unfulfilled promise. Biomass Bioenerg 19:1–15
Ovando-Medina I, Espinosa-García FJ, Núñez-Farfán J, Salvador-Figueroa M (2011) Genetic variation in Mexican Jatropha curcas L. estimated with seed oil fatty acids. J Oleo Sci 60(6):301–311
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Perrier X, Jacquemoud-Collet JP (2006) DARwin software, Version 5.0.158, http://darwin.cirad.fr/darwin
Qu J, Mao HZ, Chen W, Gao SQ, Bai YN, Sun YW, Geng YF, Ye J (2012) Development of marker-free transgenic Jatropha plants with increased levels of seed oleic acid. Biotechnol Biofuels 5(1):10. doi:10.1186/1754-6834-5-10
Rohlf FJ (2000) NTSYS-pc: Numerical Taxonomy and Multivariate Analysis System, Version 2.2, Exeter Software, Setauket, NY
Sharma SS, Negi MS, Sinha P, Kumar K, Tripathi SB (2011) Assessment of genetic diversity of biodiesel species Pongamia pinnata accessions using AFLP and three endonuclease -AFLP. Plant Mol Biol Rep 29(1):12–18
Singh A, Negi MS, Rajagopal J, Bhatia S, Tomar UK, Srivastava PS, Lakshmikumaran M (1999) Assessment of genetic diversity in Azadirachta indica using AFLP markers. Theor Appl Genet 99(1):272–279
Sinha P, Negi MS, Sharma SS, Md Islam A, Tripathi SB (2013) Analysis of genome-wide homozygosity in Jatropha curcas accessions using AFLP markers. Int J Res Pharm Sci 3:191–201
Sinha P, Md Islam A, Negi MS, Tripathi SB (2015) First identification of core accessions of Jatropha curcas from India based on molecular genetic diversity. Plant Genetic Resources (In press)
Sudheer PDVN, Mastan SG, Rahman H, Reddy MP (2010) Molecular characterization and genetic diversity analysis of Jatropha curcas L. in India using RAPD and AFLP analysis. Mol Biol Rep 37:2249–2257
Thies W (1971) Rapid and easy analysis of fatty acid composition in individual rapeseed cotyledons. Z Pflanzenzhchtg 65:181–202
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Friters A, Pot J, Paleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucl Acids Res 23(21):4407–4414. doi:10.1093/nar/23.21.4407
Wang CM, Liu P, Yi C, Gu K, Sun F, Li L, Lo LC, Liu X, Feng F, Lin G, Cao S, Hong Y, Yin Z, Yue GH (2011) A first generation microsatellite- and SNP-based linkage map of Jatropha. PLoS One 6(8):e23632
Acknowledgments
The authors sincerely acknowledge the funding received from the Department of Biotechnology, Government of India. Pratima Sinha thanks Ministry of New and Renewable Energy, Government of India, for providing fellowship for this work. We thank Dr. Y. S. Sodhi of The Centre for Genetic Manipulation of Crop Plants, University of Delhi, for his kind help in fatty acid profiling.
Conflict of interest
Authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. S1
Morphological features of J. curcas
ESM 2
(DOCX 21 kb)
ESM 3
(DOCX 17 kb)
Rights and permissions
About this article
Cite this article
Sinha, P., Islam, M.A., Negi, M.S. et al. Analysis of genetic diversity and fatty acid composition in a prebreeding material of Jatropha. J. Plant Biochem. Biotechnol. 25, 111–116 (2016). https://doi.org/10.1007/s13562-015-0301-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-015-0301-2