Abstract
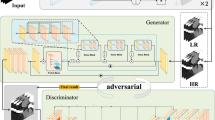
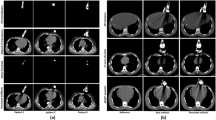
Ultrasound computed tomography (USCT) is an emerging technology that offers a noninvasive and radiation-free imaging approach with high sensitivity, making it promising for the early detection and diagnosis of breast cancer. The speed-of-sound (SOS) parameter plays a crucial role in distinguishing between benign masses and breast cancer. However, traditional SOS reconstruction methods face challenges in achieving a balance between resolution and computational efficiency, which hinders their clinical applications due to high computational complexity and long reconstruction times. In this paper, we propose a novel and efficient approach for direct SOS image reconstruction based on an improved conditional generative adversarial network. The generator directly reconstructs SOS images from time-of-flight information, eliminating the need for intermediate steps. Residual spatial-channel attention blocks are integrated into the generator to adaptively determine the relevance of arrival time from the transducer pair corresponding to each pixel in the SOS image. An ablation study verified the effectiveness of this module. Qualitative and quantitative evaluation results on breast phantom datasets demonstrate that this method is capable of rapidly reconstructing high-quality SOS images, achieving better generation results and image quality. Therefore, we believe that the proposed algorithm represents a new direction in the research area of USCT SOS reconstruction.









Similar content being viewed by others
References
Giaquinto AN, Sung H, Miller KD, Kramer JL, Newman LA, Minihan A, et al. Breast cancer statistics, 2022. CA Cancer J Clin. 2022;72(6):524–41.
Weiss A, Chavez-MacGregor M, Lichtensztajn DY, Yi M, Tadros A, Hortobagyi GN, et al. Validation study of the American Joint Committee on Cancer eighth edition prognostic stage compared with the anatomic stage in breast cancer. JAMA Oncol. 2018;4(2):203–209.
Boyd NF, Guo H, Martin LJ, Sun LM, Stone J, Fishell E, et al. Mammographic density and the risk and detection of breast cancer. N Engl J Med. 2007;356(3):227–36.
O’Flynn EAM, Fromageau J, Ledger AE, Messa A, D’Aquino A, Schoemaker MJ, et al. Ultrasound tomography evaluation of breast density: a comparison with noncontrast magnetic resonance imaging. Invest Radiol. 2017;52(6):343–8.
Wiskin J, Malik B, Pirshafiey N, Klock J. Limited view reconstructions with transmission ultrasound tomography: clinical implications and quantitative accuracy. In: Medical imaging 2020: ultrasonic imaging and tomography. vol. 11319. Bellingham: SPIE; 2020. p. 167–174.
Andre MP, Barker C, Sekhon N, Wiskin J, Borup DT, Callahan K. Pre-clinical experience with full-wave inverse-scattering for breast imaging. In: Acoustical Imaging. vol. 29. Dordrecht: Springer; 2008. p. 73–80.
Ali R, Hsieh S, Dahl J. Open-source gauss-newton-based methods for refraction-corrected ultrasound computed tomography. In: Medical imaging 2019: ultrasonic imaging and tomography. vol. 10955. Bellingham: SPIE; 2019. p. 39–52.
Huthwaite P, Simonetti F. High-resolution imaging without iteration: a fast and robust method for breast ultrasound tomography. J Acoust Soc Am. 2011;130(3):1721–34.
Lucka F, Perez-Liva M, Treeby BE, Cox BT. High resolution 3D ultrasonic breast imaging by time-domain full waveform inversion. Inverse Probl. 2022;38(2):025008.
Vishnevskiy V, Rau R, Goksel O. Deep variational networks with exponential weighting for learning computed tomography. In: Proceeding of MICCAI. vol. 11769. Cham: Springer; 2019. p. 310–318.
Vishnevskiy V, Sanabria SJ, Goksel O. Image reconstruction via variational network for real-time hand-held sound-speed imaging. In: Proceeding of MLMIR. vol. 11074. Cham: Springer; 2018. p. 120–128.
Bernhardt M, Vishnevskiy V, Rau R, Goksel O. Training variational networks with multidomain simulations: speed-of-sound image reconstruction. IEEE Trans Ultrason Ferroelectr Freq Control. 2020;67(12):2584–94.
Fan Y, Ying L. Solving traveltime tomography with deep learning. Commun Math Stat. 2023;11:3–19.
Qu X, Ren C, Yan G, Zheng D, Tang W, Wang S, et al. Deep-learning-based ultrasound sound-speed tomography reconstruction with Tikhonov pseudo-inverse priori. Ultrasound Med Biol. 2022;48(10):2079–94.
Fan Y, Wang H, Gemmeke H, Hesser J. MI-Net: a deep network for non-linear ultrasound computed tomography reconstruction. In: Proceeding of IEEE International Ultrasonics Symposium. Piscataway: IEEE 2020;1–3.
Fan Y, Wang H, Gemmeke H, Hopp T, Hesser J. DDN: dual domain network architecture for non-linear ultrasound transmission tomography reconstruction. In: Medical imaging 2021: ultrasonic imaging and tomography. vol. 11602. Bellingham: SPIE 2021;40–45.
Zhao W, Wang H, Gemmeke H, van Dongen KWA, Hopp T, Hesser J. Ultrasound transmission tomography image reconstruction with a fully convolutional neural network. Phys Med Biol. 2020;65(23):235021.
Fan Y, Wang H, Gemmeke H, Hopp T, Dongen KV, Hesser J. Memory-Efficient Neural Network For Non-Linear Ultrasound Computed Tomography Reconstruction. In: Proceeding of IEEE international symposium on biology image. Piscataway: IEEE 2021;429–432.
Prasad S, Almekkawy M. DeepUCT: Complex cascaded deep learning network for improved ultrasound tomography. Phys Med Biol. 2022;67(6):065008.
Ronneberger O, Fischer P, Brox T. U-Net: Convolutional networks for biomedical image segmentation. In: Proceeding of MICCAI. vol. 9351. Cham: Springer 2015; 234–241.
Fan YL, Wang HJ, Gemmeke H, Hopp T, Hesser J. Model-data-driven image reconstruction with neural networks for ultrasound computed tomography breast imaging. Neurocomputing. 2022;467:10–21.
Zhu B, Liu JZ, Cauley SF, Rosen BR, Rosen MS. Image reconstruction by domain-transform manifold learning. Nature. 2018;555(7697):487–92.
Goodfellow IJ, Pouget-Abadie J, Mirza M, Xu B, Warde-Farley D, Ozair S, et al. Generative adversarial nets. In: Proceeding of NIPS. vol. 2. New York: Curran Associates 2014;2672–2680.
Arjovsky M, Chintala S, Bottou L. Wasserstein generative adversarial networks. In: Proceeding of ICML. New York: PMLR 2017;214–223.
Gulrajani I, Ahmed F, Arjovsky M, Dumoulin V, Courville A. Improved Training of Wasserstein GANs. In: Proceeding of NIPS. vol. 30. New York: Curran associates 2017;5769–5779.
Isola P, Zhu JY, Zhou TH, Efros AA. Image-to-image translation with conditional adversarial networks. In: Proceeding of CVPR. Piscataway: IEEE 2017;1125–1134.
Mirza M, Osindero S. Conditional generative adversarial nets. Preprint at https://arxiv.org/abs/1411.1784.
Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. Preprint at https://arxiv.org/abs/1409.1556.
Hinton GE, Salakhutdinov RR. Reducing the dimensionality of data with neural networks. Science. 2006;313(5786):504–7.
Hu J, Shen L, Albanie S, Sun G, Wu E. Squeeze-and-excitation networks. IEEE Trans Pattern Anal Mach Intell. 2020;42:2011–23.
Roy AG, Navab N, Wachinger C. Recalibrating fully convolutional networks with spatial and channel “squeeze and excitation” blocks. IEEE Trans Med Imaging. 2019;38:540–9.
Maas AL, Hannun AY, Ng AY. Rectifier nonlinearities improve neural network acoustic models. In: Proceeding of ICML. vol. 28. New York: PMLR 2013;p. 3.
Kak AC, Slaney M. Principles of computerized tomographic imaging. Philadelphia: SIAM; 2001.
Lou Y, Zhou W, Matthews T, Appleton C, Anastasio M. Generation of anatomically realistic numerical phantoms for photoacoustic and ultrasonic breast imaging. J Biomed Opt. 2017;4:041015.
Lou Y. Optical and acoustic breast phantoms. Harvard Dataverse https://doi.org/10.7910/DVN/NZBJOC.
Mettivier G, Sarno A, Franco Fd, Bliznakova K, Bliznakov Z, Hernandez AM, et al. The Napoli-Varna-Davis project for virtual clinical trials in X-ray breast imaging. In: Proceeding of IEEE nuclear science symposium and medicine image conference. Piscataway: IEEE 2019;1–5.
Sarno A, Mettivier G, di Franco F, Varallo A, Bliznakova K, Hernandez AM, et al. Dataset of patient-derived digital breast phantoms for in silico studies in breast computed tomography, digital breast tomosynthesis, and digital mammography. Med Phys. 2021;48(5):2682–93.
Bliznakova K, Dukov N, Feradov F, Gospodinova G, Bliznakov Z, Russo P, et al. Development of breast lesions models database. Phys Med. 2019;64:293–303.
Dukov N, Bliznakova K, Feradov F, Ridlev I, Bosmans H, Mettivier G, et al. Models of breast lesions based on three-dimensional X-ray breast images. Phys Med. 2019;57:80–7.
Li C, Duric N, Littrup P, Huang L. In vivo breast sound-speed imaging with ultrasound tomography. Ultrasound Med Biol. 2009;35(10):1615–28.
Funding
This work was supported by the National Natural Science Foundation of China (Grant Nos. 62122072, 12174368, and 61705216), the National Key R & D Program of China (Grant No. 2022YFA1404400), the Anhui Provincial Science and Technology Department (Grant Nos. 202203a07020020 and 18030801138), and the Research Fund of the University of Science and Technology of China (YD2090002015).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Long, X., Tian, C. Spatial and channel attention-based conditional Wasserstein GAN for direct and rapid image reconstruction in ultrasound computed tomography. Biomed. Eng. Lett. 14, 57–68 (2024). https://doi.org/10.1007/s13534-023-00310-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13534-023-00310-x