Abstract
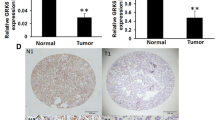
Golgi phosphoprotein 3 (GOLPH3) has been reported to be involved in the development of several human tumours. However, little is known about GOLPH3 expression and its clinical significance in non-small cell lung cancer (NSCLC). The present study was conducted to investigate the expression of GOLPH3 and its prognostic significance in NSCLC. Immunohistochemical analysis was used to determine the expression of GOLPH3 in 145 cases NSCLC tissue and their corresponding adjacent normal tissues. The results are the following: (1) The GOLPH3 protein was mainly located in the cytoplasm of NSCLC tissue; (2) GOLPH3 was highly expressed in 71.7 % of NSCLC tissues versus 22.8 % of adjacent tissues (P < 0.01); (3) GOLPH3 expression was significantly associated with tumour-node-metastasis (TNM) stage (P = 0.001), lymph node status (P = 0.014), and degree of differentiation (P = 0.018); (4) The Kaplan–Meier curve indicated that the patients with high GOLPH3 expression had significantly shorter survival than those with low GOLPH3 expression; and (5) Cox regression analysis demonstrated that the expression of GOLPH3, lymph node status, and TNM stage were independent prognostic predictors for NSCLC patients. High GOLPH3 expression is predictive of poor prognosis of NSCLC, implying that GOLPH3 may be a promising new target for targeted therapies for NSCLC.



Similar content being viewed by others
References
Barlési F, Giaccone G, Gallegos-Ruiz MI, et al. Global histone modifications predict prognosis of resected non small-cell lung cancer. J Clin Oncol. 2007;25(28):4358–64.
Hung JJ, Jeng WJ, Hsu WH, et al. Time trends of overall survival and survival after recurrence in completely resected stage I non-small cell lung cancer. J Thorac Oncol. 2012;7(2):397–405.
Zhu CQ, Shih W, Ling CH, et al. Immunohistochemical markers of prognosis in non-small cell lung cancer: a review and proposal for a multiphase approach to marker evaluation. J Clin Pathol. 2006;59(8):790–800.
Sechi S, Colotti G, Belloni G, et al. GOLPH3 is essential for contractile ring formation and Rab11 localization to the cleavage site during cytokinesis in Drosophila melanogaster. PLoS Genet. 2014;10(5):e1004305.
Bell AW, Ward MA, Blackstock WP, et al. Proteomics characterization of abundant Golgi membrane proteins. J Biol Chem. 2001;276(7):5152–65.
Scott KL, Kabbarah O, Liang MC, et al. GOLPH3 modulates mTOR signalling and rapamycin sensitivity in cancer. Nature. 2009;459(7250):1085–90.
Zeng Z, Lin H, Zhao X, et al. Overexpression of GOLPH3 promotes proliferation and tumorigenicity in breast cancer via suppression of the FOXO1 transcription factor. Clin Cancer Res. 2012;18(15):4059–69.
Hua X, Yu L, Pan W, et al. Increased expression of Golgi phosphoprotein-3 is associated with tumor aggressiveness and poor prognosis of prostate cancer. Diagn Pathol. 2012;7:127.
Wang JH, Chen XT, Wen ZS, et al. High expression of GOLPH3 in esophageal squamous cell carcinoma correlates with poor prognosis. PLoS One. 2012;7(10):e45622.
Hu BS, Hu H, Zhu CY, et al. Overexpression of GOLPH3 is associated with poor clinical outcome in gastric cancer. Tumour Biol. 2013;34(1):515–20.
Wang Z, Li Z, Wu C, et al. MACC1 overexpression predicts a poor prognosis for non-small cell lung cancer. Med Oncol. 2014;31(1):790.
Pereira NA, Pu HX, Goh H, et al. Golgi phosphoprotein 3 mediates the Golgi localization and function of protein O-linked mannose β-1,2-N-acetlyglucosaminyltransferase 1. J Biol Chem. 2014;289(21):14762–770.
Ng MM, Dippold HC, Buschman MD, et al. GOLPH3L antagonizes GOLPH3 to determine Golgi morphology. Mol Biol Cell. 2013;24(6):796–808.
Ali MF, Chachadi VB, Petrosyan A, et al. Golgi phosphoprotein 3 determines cell binding properties under dynamic flow by controlling Golgi localization of core 2 N-acetylglucosaminyltransferase 1. J Biol Chem. 2012;287(47):39564–77.
Kunigou O, Nagao H, Kawabata N, et al. Role of GOLPH3 and GOLPH3L in the proliferation of human rhabdomyosarcoma. Oncol Rep. 2011;26(5):1337–42.
Graham TR, Burd CG. Coordination of Golgi functions by phosphatidylinositol 4-kinases. Trends Cell Biol. 2011;21(2):113–21.
Xue Y, Wu G, Liao Y, et al. GOLPH3 is a novel marker of poor prognosis and a potential therapeutic target in human renal cell carcinoma. Br J Cancer. 2014;110(9):2250–60.
Ma Y, Ren Y, Zhang X, et al. High GOLPH3 expression is associated with a more aggressive behavior of epithelial ovarian carcinoma. Virchows Arch. 2014;464(4):443–52.
Cizkova M, Vacher S, Meseure D, et al. PIK3R1 underexpression is an independent prognostic marker in breast cancer. BMC Cancer. 2013;13:545.
Kapoor S. Promising, new prognostic markers of esophageal carcinomas. APMIS. 2013;121(10):1011.
Foiani M, Bartek J. Golgi feels DNA’s pain. Cell. 2014;156(3):392–3.
Farber-Katz SE, Dippold HC, Buschman MD, et al. DNA damage triggers Golgi dispersal via DNA-PK and GOLPH3. Cell. 2014;156(3):413–27.
Tu L, Chen L, Banfield DK. A conserved N-terminal arginine-motif in GOLPH3- family proteins mediates binding to coatomer. Traffic. 2012;13(11):1496–507.
Dippold HC, Ng MM, Farber-Katz SE, et al. GOLPH3 bridges phosphatidylinositol-4- phosphate and actomyosin to stretch and shape the Golgi to promote budding. Cell. 2009;139(2):337–51.
Scott KL, Chin L. Signaling from the Golgi: mechanisms and models for Golgi phosphoprotein 3-mediated oncogenesis. Clin Cancer Res. 2010;16(8):2229–34.
Abraham RT. GOLPH3 links the Golgi network to mTOR signaling and human cancer. Pigment Cell Melanoma Res. 2009;22(4):378–9.
Bugarcic A, Zhe Y, Kerr MC, et al. Vps26A and Vps26B subunits define distinct retromer complexes. Traffic. 2011;12(12):1759–73.
Zhou X, Zhan W, Bian W, et al. GOLPH3 regulates the migration and invasion of glioma cells though RhoA. Biochem Biophys Res Commun. 2013;433(3):338–44.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, Y., Ma, M. & Han, B. GOLPH3 high expression predicts poor prognosis in patients with resected non-small cell lung cancer: an immunohistochemical analysis. Tumor Biol. 35, 10833–10839 (2014). https://doi.org/10.1007/s13277-014-2357-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2357-3