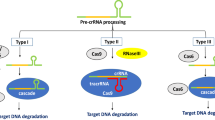
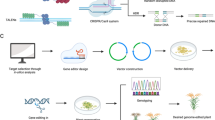
Abstract
Targeted genome editing by Clustered Regularly Interspaced Short Palindromic Repeat- CRISPR-associated (CRISPR-Cas) system has revolutionized basic and translational plant research. There is widespread use of CRISPR-Cas technology which has the potential to address challenges like food insecurity and climate crisis. Crops with improved traits (e.g., higher yield, drought tolerant) that would take several years to generate can now be developed at a much reduced time, drastically expediting the availability of the crops for release in the market. However, several factors are involved in successfully applying the CRISPR-Cas system in agriculture and the widespread adoption and acceptability of genome-edited products that involve multiple institutions and people from different spheres of society. Besides the scientific and legal intricacies of releasing CRISPR-edited crops, “public perception” equally matters in successfully deploying the technology and its products. “Lack of” or “overwhelming” information can both affect the success of the CRISPR-Cas system in translational agriculture research. A bird’s-eye-view of the CRISPR-Cas genome editing tool for people from different strata of society is essential for the wide acceptability of genome-edited crops. This review provides a general overview of the CRISPR-Cas system, the concept of technology development, challenges, and regulations involved in translational research.
Graphical abstract

Similar content being viewed by others
Introduction
“Molecular structure of DNA”, the groundbreaking research work that reported the structure of DNA, was published in Nature on April 25, 1953 [55]. In the past 70 years, the scientific community has made discoveries and innovations by which we can read (use of sequencing technologies), cut, copy, paste DNA from one source of DNA to the other (use of recombinant DNA technology) and transfer foreign DNA into various kinds of cells [51]. Besides, researchers have also studied, identified, and engineered proteins (Zinc Finger Nuclease (ZFN) and Transcription Activator Like Effector Nuclease (TALEN)) that can bind to any particular DNA sequence in a sequence-dependent manner [14]. These seminal discoveries and inventions have created approaches to modify the DNA sequence of a genome in a targeted manner, broadly known as genome editing. The discovery of the use of the CRISPR in association with the CRISPR-associated endonuclease to edit any sequence of DNA in a targeted manner has catapulted the adoption of genome editing by CRISPR-Cas systems for translational research, especially in the areas of medicine and agriculture [8, 23, 24, 53].
Food insecurity and climate crisis are the two critical challenges of the human population [9, 10]. The 2023 report on food security published by FAO, IFAD, UNICEF, WFP, or WHO estimated that in the year 2022, between 691 and 783 million people faced hunger [10]. The emissions gap report 2022, published by the United Nations Environment Program, states that there is a need for “rapid transformation of societies” to address the climate crisis, and “agricultural production” remains the largest contributor to greenhouse gases in the food systems [50]. Several efforts have been initiated to address food insecurity and climate crisis, including improving agricultural practices, developing newer varieties of crops with higher yields, requiring reduced amounts of fertilizers, being resistant to diseases, and increasing resiliency against climate change. However, there is a need to implement technologies or products that can be rapidly deployed in the agri-food ecosystem to accelerate the efforts toward addressing these challenges.
New plant breeding techniques such as genome editing by CRISPR-Cas system have demonstrated tremendous potential in expediting crop improvement through their use in basic and translational research [4, 9, 41, 59]. Since the first use of CRISPRs 10 years ago in plants, genome-edited crops have been commercialized and are available in the market, supporting the efforts toward accelerating the development of crops with novel traits (Table 1) [9, 29, 33, 43, 49]. In addition, multiple.
crops are at different stages of the research and development pipelines [9]. Furthermore, the combination of genomics and CRISPR-Cas system provides the opportunity to discover novel targets for crop improvement that have remained unearthed in the past because of the high cost and lack of tools [25, 42, 48, 59].
The widespread use of the CRISPR-Cas system calls for a “birds-eye-view” of the CRISPR-Cas genome editing tool that is comprehensive and simple. In this review, an attempt is made to provide a generalized overview of the complex journey (CRISPR biology, genome-edited crop development, regulations, associated risks, and stakeholders involved) of the CRISPR-Cas system from lab to field to market. To provide a birds-eye-view this review does not attempt to provide a detailed list of genome-edited crops and the current status of regulation for genome-edited crops in different countries, which can be found in other excellent review articles [7, 9, 25].
CRISPR-Cas system—overview of its use as genome editing tool in agriculture
The basic process of genome editing
Broadly, at the molecular and biochemical level, genome editing tools comprise a DNA targeting component and nucleases (Fig. 1a) [13, 14, 28]. On the expression of these two components in a cell, the targeting component binds to a specific sequence of DNA, while the nuclease cleaves the double strand DNA near the proximity of the binding site, which then gets repaired by the cellular DNA repair machinery [13]. During the repair process of the targeted cleaved site, DNA bases can get inserted, deleted, and replaced, which changes the DNA sequence of this site, thus editing the genome.
Basics of genome editing tools. a Genome editing tools comprise a sequence-specific targeting component and an endonuclease capable of introducing a double strand break in the DNA. They can be classified into two categories based on the targeting component. b The first category comprises genome editing tools with protein domains acting as the DNA binding component (e.g., Transcription Activator Like Nucleases (top) and Zinc Finger Nucleases (bottom)). c, d The second category comprises an RNA molecule (guide RNA) that acts as the targeting component (e.g., CRISPR-Cas system). d The CRISPR-associated nuclease can be engineered to generate a catalytically inactive dead Cas (dCas), which is fused to functional proteins or protein domains (called effectors) to develop novel editing technologies. For example, fusing DNA methyltransferases to dCAS systems generates tools for methylating the DNA cytosine, a type of epigenome editing. The (asterisk*) denotes a mutated Cas. The figure depicts only one strategy of targeting DNA modifiers, several other strategies have been developed to target multiple effectors. (1c and 1d were created by using Biorender.com)
Types of genome editing tools
Based on the nature of the targeting component, genome editing tools can be classified into two large categories: 1) Zinc Finger Nucleases and Transcription Activator Like Nucleases—In this category, a sequence specific DNA binding protein guides the endonuclease to the targeted DNA sequence (Fig. 1b); 2) CRISPR-Cas systems—A nucleic acid (called single guide RNA) acts as a guide to target an endonuclease to the DNA (Fig. 1c) [11]. The most common type of CRISPR-Cas system is the CRISPR-Cas9 system. The system comprises of a single guide RNA molecule and the Cas9 endonuclease. The Cas9 endonuclease and the guide RNA forms a complex which binds to the target sequence by the guide RNA. On binding to the target site, Cas9 introduces a double strand break in the DNA.
Both categories of genome editing tools may require testing of several variants of the targeting components (guide RNA or protein sequences) to identify a variant that is highly specific. Designing proteins for category 1 of the genome editing tool that binds to DNA in a sequence-specific manner is tedious, extremely difficult, and expensive compared to creating a RNA sequences that can guide the CRISPR-Cas system to its target, which is one of the primary reasons for the success of CRISPR-Cas systems in genome editing.
CRISPR-Cas as a targeting tool for other types of editing
Besides using the CRISPR-Cas system for genome editing by introducing double-strand breaks, the system has been adopted to target a wide range of effectors (e.g., DNA modifying enzymes, transcriptional activators and repressor) (Fig. 1d). To target effectors, a catalytically inactive Cas (called dead Cas or dCas—generated by introducing mutations at the catalytic site of the endonuclease) with guide RNAs is harnessed for site-specific manipulation of DNA without introducing double-strand breaks [40]. Depending on the type of effector attached to the dCas system, novel editing technologies, such as base editing, prime editing, and epigenome editing has been developed [12, 14, 28]. Furthermore, dCas systems has been fused to transcriptional activators or repressors to turn on or off gene expression [14]. In addition to targeting DNA, CRISPR-Cas variants have been identified that target RNA, thus increasing the repertoire of application of CRISPR [26].
An overview of the concept of scientific process of discovery to its application for the betterment of humanity with reference to CRISPR-Cas use in agriculture
In the present era of the use of CRISPR-Cas genome editing tool for crop improvement, the distinction between basic biology and translational research is getting blurred. Therefore, it becomes apparent that a comprehensive overview of the discovery of CRISPR (“basic biology for knowledge generation”) to its application (“translational research”) in agriculture is necessary. Here I have divided the process of “scientific discovery” to its “availability in the market” into four major phases (Fig. 2): Phase 1) Knowledge generation—Primarily aimed to understand the principles of life with long-term goals of improvement of humanity; Phase 2) Invention—Development of a useful tool/process/product based on the data available from exploratory research; Phase 3) Innovation and upscaling—Using the invention to solve a problem of humankind, developing prototypes, examining risks, and improving reliability; Phase 4) Release of product to market—Providing the final products to the end-consumers. Although 4 distinct phases are mentioned here, it is to be noted that the activities among the different phases overlap and may take several years to decades. Furthermore, legal processes, e.g., invention disclosure, patents, and licensing, also come into play from phase 2, which are out of the scope of this review. Readers are encouraged to refer to the following resources for more information [16] (https://www.techtransfer.harvard.edu/master-the-technology-transfer-process-with-this-step-by-step-guide/.; https://tlo.mit.edu/learn-about-intellectual-property/technology-transfer-process.; https://research.utoronto.ca/inventions-commercialization-entrepreneurship/commercialize-invention.)
Conceptual roadmap of scientific discovery from basic biology to translational research. The journey from basic biology to translational research can be divided into four phases 1) knowledge generation, 2) invention, 3) innovation and upscaling, and 4) release of end product. An example is provided in parentheses depicting the stages of CRISPR development
In the context of using CRISPR-Cas technology in agriculture, studying CRISPR and examining its role in prokaryotes encompasses the knowledge generation phase. The presence of the CRISPR in the bacterial genome was first reported in 1987 [21]. Only in early 2000 identification of CRISPR-associated (Cas) genes and the similarity between spacers and bacteriophage sequences were noticed, providing the first hints of CRISPR in bacterial immunity [22, 30]. Finally, indications of CRISPR playing a role against bacteriophage were demonstrated in 2007 [5]. Knowledge generation (Phase 1) for the CRISPR-Cas system is still undergoing with many novel discoveries being made [2, 54]. The seminal work (invention) demonstrating the sequence-specific double-stranded DNA cleavage by combining Cas9 protein with a single guide RNA by Dr. Jennifer Doudna and Dr. Emanuele Charpentier in 2012 initiated the phase 2 [23]. This was followed by the demonstration of successful genome editing by the CRISPR-Cas system in eukaryotic cells in the laboratory of Dr. Feng Zhang and Dr. Jennifer Doudna [8, 24]. In the same year, three different groups simultaneously demonstrated gene editing by CRISPR in model plants [29, 33, 43], which opened the avenue for application of CRISPR in agriculture leading to the initiation of the innovation phase. Several labs around the world initiated experiments to edit the genome of various crops to improve traits such as disease resistance, drought tolerance [59]. Since the first demonstration of using CRISPR in agriculture in 2013, multiple crops with improved characteristics have been developed to address specific problems (Table 1) [9]. For example, bacterial blight-resistant rice and powdery mildew-resistant wheat have been developed [36, 56]. The innovation phase also includes validation, quality control, and risk assessment of the genome-edited crops. Finally, the end products are released to the market, i.e., to the end consumers, which is phase 4. The purview of the term “consumers” for a genome-edited crop can vary depending on the end product and the trait improved. For example, consumers of genome-edited products aimed at increasing food and feed quality, e.g., “tomatoes with high content of Gaba amino butyric acid,” are the buyer consuming the fruit [35]. While the “consumers” for a genome-edited crop with improved agronomic traits, e.g., “bacterial speck resistant tomatoes”, are the growers themselves [37]. The first CRISPR-edited crop available in the market is GABA-enriched tomatoes [52].
People involved in the 4 phases of discovery to application
In the four phases of basic to translational research, researchers (scientists, professors, students, and postdoctoral fellows) are highly involved in the first three phases of the process, which include the basic understanding of the CRISPR biology, development of CRISPR as a genome editing tool, examining and assessing the associated risks and optimizing the genome-edited products (Fig. 3). The third phase can also include agricultural corporations and growers. The fourth phase includes consumers of the genome-edited products, which usually includes people from all strata of society (anyone purchasing/consuming the end product). Besides, lawyers and policymakers are also heavily involved in this process. It has been suggested that a high engagement among all the stakeholders will facilitate research and innovation [27]. Therefore, knowledge about the stakeholders is key to developing strategies for wider acceptability of the CRISPR-Cas system.
Schematic diagram showing people involved at different phases of the development of CRISPR-technology. The diagram shows that consumers of CRISPR-edited crops can be broadly divided into two groups (Individuals/entities directly involved and not- directly involved) based on their association with the development of the technology and its products. Examples of stakeholders involved at different phases is mentioned to provide a broader picture of the society
Challenges associated with the use of CRISPR-Cas system in agriculture
Any new technology faces several challenges before it is successfully adopted [34]. Several reports and reviews are available on the impacts, hazards, and challenges of CRISPR-Cas technology use in agriculture [3, 18, 38, 46]. Here we highlight a few key challenges associated with the different phases:
-
1.
Challenges in phases 2 & 3—(a) Off-target or non-specific edits –A major concern in the use of CRISPR is the introduction of unwanted edits by non-specific binding of the CRISPR-Cas system to other parts of the genome when expressed in a plant [58]. These off-target effects may introduce changes that might be lethal for a genome edited crop, thus posing a significant challenge for the use of CRISPR-Cas tool in translational research [47, 57]. Researchers involved at phase 1, 2 and 3 have developed different strategies to reduce off-targets which includes identifying or generating CRISPR-Cas variants with higher specificity and developing computational tools that aid in designing guide RNAs with lesser chances of binding to unwanted regions of the genome [58]. In addition, tools to identify genome-wide off-target effects have also been developed [44]. (b) Stable insertion of CRISPR-Cas constructs—The most popular method of expressing the CRISPR-Cas system for agricultural purposes is by agrobacterium-mediated gene transfer, which inserts the CRISPR-Cas system into the plant’s genome, hence, involving the generation of transgenic plants [15]. The transgene is eventually segregated out [19]. Several novel methods using nanotechnology, plant viral vectors, and protein delivery have been innovated to avoid inserting the CRISPR-Cas system into the plant’s genome [1, 17, 28].
-
2.
Challenges in Phase 4: This step is critical as they involve the direct involvement of growers and consumers. An important consideration at this stage is the “perception” of the technology and the product by its grower [46]. A recent study in Germany indicated that “public perception” towards technology is critical [31, 32]. If the consumer or the grower does not perceive it well, then the risks of rejection in an open market are very high. For example, if consumers perceive genome-edited crops as harmful to health and the environment, even though scientifically proven otherwise, growers would not use genome-edited crops because of the risk of not getting any consumers. Therefore, it is crucial to develop outreach programs to make people aware of the technology’s pros and cons and be transparent to consumers [6]. For example, an effective outreach approach might be to explain the genome editing tools to school and university students by organizing seminars and by incorporating discussion about the genome editing tools in university courses. These courses should be mandatory for all students.
-
3.
Regulatory Challenges—The controversy on genetically modified crops is longstanding and remains unresolved [45]. CRISPR-Cas technology has re-ignited the discussion on the regulating genetically modified crops [39]. Whether genome-edited crops should fall under the regulations of crops developed by genetic engineering or the traditional breeding practices have been debated in many countries in the last few years. To facilitate the process of regulation for genome-edited crops, several experts have recommended protocols for the regulation of genome-edited crops [20]. Currently, it seems that several countries are inclined to have similar regulations for genome-edited crops as conventional breeding crops if no foreign DNA is present in the crop that was used to introduce the edit [7]. However, countries that are relaxing the regulations, the level of relaxation may vary among countries and the genome edited products being considered.
Conclusion
This review provides a birds-eve-view of the scientific process of CRISPRs, from being a research topic to its use in translational research for improving agricultural traits. The field of CRISPR-Cas system and its use for genome editing has exponentially grown in the past 10 years. Therefore, it is to be noted that many complexities have not been discussed in this review to give a broad picture of the field.
The CRISPR-Cas technology and CRISPR-edited products are already under scrutiny by people from different strata of society, which will rapidly increase as more genome-edited products get released into the market. A critical component in accepting, adopting, and deploying genome-edited crops is the “public (consumer) perception” of CRISPR-Cas technology. The “consumers” include a broad range of people who can be categorized—(1) people directly involved in developing the technology and its products and (2) people not directly involved in developing the technology (Fig. 3). It is onerous for the proponents of the CRISPR-Cas systems to be transparent to the other communities in explaining the pros and cons of the technology [27].
With the ease of the use of the CRISPR-Cas system, its benefits to crop improvement, and the ease of regulations on genome-edited crops, it is apparent that there will be an increase in the commercialization and availability of the number of genome-edited crops and their products. However, the success of the genome-edited crop will largely depend on its acceptability by consumers. A birds-eye-view provides an overview of the whole process of CRISPR-technology development that can help consumers make better-informed choices about CRISPR-edited products.
References
Ahmar S, Mahmood T, Fiaz S, Mora-Poblete F, Shafique MS, Chattha MS, Jung KH. Advantage of nanotechnology-based genome editing system and its application in crop improvement. Front Plant Sci. 2021;12:663849. https://doi.org/10.3389/fpls.2021.663849.
Al-Shayeb B, Skopintsev P, Soczek KM, Stahl EC, Li Z, Groover E, Smock D, Eggers AR, Pausch P, Cress BF, Huang CJ, Staskawicz B, Savage DF, Jacobsen SE, Banfield JF, Doudna JA. Diverse virus-encoded CRISPR-Cas systems include streamlined genome editors. Cell. 2022;185(24):4574–86. https://doi.org/10.1016/j.cell.2022.10.020.
Baltes NJ, Gil-Humanes J, Voytas DF. Genome engineering and agriculture: opportunities and challenges. Prog Mol Biol Transl Sci. 2017;149:1–26. https://doi.org/10.1016/bs.pmbts.2017.03.011.
Bansal KC, Molla KA, Chinusamy V. Genome editing: a boon for plant biologists, breeders and farmers. Curr Sci. 2022;123(1):5.
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315(5819):1709–12. https://doi.org/10.1126/science.1138140.
Baur P. When farmers are pulled in too many directions: comparing institutional drivers of food safety and environmental sustainability in california agriculture. Agric Hum Values Hum Values. 2020;37:1175–94. https://doi.org/10.1007/s10460-020-10123-8.
Buchholzer M, Frommer WB. An increasing number of countries regulate genome editing in crops. New Phytol. 2023;237(1):12–5. https://doi.org/10.1111/nph.18333.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339(6121):819–23. https://doi.org/10.1126/science.1231143.
FAO. Gene editing and agrifood systems. Rome: FAO; 2022. p. 86.
FAO I, UNICEF, WFP, WHO. The state of food security and nutrition in the world 2023 Urbanization, agrifood systems transformation and healthy diets across the rural–urban continuum. Rome: FAO; 2023. p. 316.
Gaj T, Gersbach CA, Barbas CF 3rd. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013;31(7):397–405. https://doi.org/10.1016/j.tibtech.2013.04.004.
Gallego-Bartolome J. DNA methylation in plants: mechanisms and tools for targeted manipulation. New Phytol. 2020. https://doi.org/10.1111/nph.16529.
Gao C. Precision plant breeding using genome editing technologies. Transgenic Res. 2019;28(Suppl 2):53–5. https://doi.org/10.1007/s11248-019-00132-7.
Gardiner J, Ghoshal B, Wang M, Jacobsen SE. CRISPR-Cas-mediated transcriptional control and epi-mutagenesis. Plant Physiol. 2022;188(4):1811–24. https://doi.org/10.1093/plphys/kiac033.
Gelvin SB. Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol Mol Biol Rev. 2003;67(1):16–37. https://doi.org/10.1128/MMBR.67.1.16-37.2003.
Graff GD, Sherkow JS. Models of technology transfer for genome-editing technologies. Annu Rev Genomics Hum Genet. 2020;21:509–34. https://doi.org/10.1146/annurev-genom-121119-100145.
Gu X, Liu L, Zhang H. Transgene-free genome editing in plants. Front Genome Ed. 2021;3:805317. https://doi.org/10.3389/fgeed.2021.805317.
Hartung F, Schiemann J. Precise plant breeding using new genome editing techniques: opportunities, safety and regulation in the EU. Plant J. 2014;78(5):742–52. https://doi.org/10.1111/tpj.12413.
He Y, Zhao Y. Technological breakthroughs in generating transgene-free and genetically stable CRISPR-edited plants. aBIOTECH. 2020;1(1):88–96. https://doi.org/10.1007/s42994-019-00013-x.
Huang S, Weigel D, Beachy RN, Li J. A proposed regulatory framework for genome-edited crops. Nat Genet. 2016;48(2):109–11. https://doi.org/10.1038/ng.3484.
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol. 1987;169(12):5429–33. https://doi.org/10.1128/jb.169.12.5429-5433.1987.
Jansen R, Embden JD, Gaastra W, Schouls LW. Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol. 2002;43:1565–75. https://doi.org/10.1046/j.1365-2958.2002.02839.x.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337(6096):816–21. https://doi.org/10.1126/science.1225829.
Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J. RNA-programmed genome editing in human cells. Elife. 2013;2:e00471. https://doi.org/10.7554/eLife.00471.
Karmakar S, Das P, Panda D, Xie K, Baig MJ, Molla KA. A detailed landscape of CRISPR-Cas-mediated plant disease and pest management. Plant Sci. 2022;323:111376. https://doi.org/10.1016/j.plantsci.2022.111376.
Koonin EV, Makarova KS, Zhang F. Diversity, classification and evolution of CRISPR-Cas systems. Curr Opin Microbiol. 2017;37:67–78. https://doi.org/10.1016/j.mib.2017.05.008.
Kuiken T, Barrangou R, Grieger K. (Broken) Promises of sustainable food and agriculture through new biotechnologies: the CRISPR case. The CRISPR J. 2021. https://doi.org/10.1089/crispr.2020.0098.
Kumar M, Prusty MR, Pandey MK, Singh PK, Bohra A, Guo B, Varshney RK. Application of CRISPR/Cas9-mediated gene editing for abiotic stress management in crop plants. Front Plant Sci. 2023;14:1157678. https://doi.org/10.3389/fpls.2023.1157678.
Li JF, Norville JE, Aach J, McCormack M, Zhang D, Bush J, Church GM, Sheen J. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol. 2013;31(8):688–91. https://doi.org/10.1038/nbt.2654.
Mojica FJ, Diez-Villasenor C, Garcia-Martinez J, Soria E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol. 2005;60(2):174–82. https://doi.org/10.1007/s00239-004-0046-3.
Muller R, Clare A, Feiler J, Marco N. Between a rock and a hard place. EMBO Rep. 2021;22(7):e53205.
Muller R, Feiler J, Clare A. A doomed technology? On gene editing in bavarian livestock agriculture, policy field conflicts and responsible research and innovation. Front Polit Sci. 2022. https://doi.org/10.3389/fpos.2022.800211.
Nekrasov V, Staskawicz B, Weigel D, Jones JD, Kamoun S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol. 2013;31(8):691–3. https://doi.org/10.1038/nbt.2655.
Nelissen H, Moloney M, Inze D. Translational research: from pot to plot. Plant Biotechnol J. 2014;12(3):277–85. https://doi.org/10.1111/pbi.12176.
Nonaka S, Arai C, Takayama M, Matsukura C, Ezura H. Efficient increase of ɣ-aminobutyric acid (GABA) content in tomato fruits by targeted mutagenesis. Sci Rep. 2017;7(1):7057. https://doi.org/10.1038/s41598-017-06400-y.
Oliva R, Ji C, Atienza-Grande G, Huguet-Tapia JC, Perez-Quintero A, Li T, Eom JS, Li C, Nguyen H, Liu B, Auguy F, Sciallano C, Luu VT, Dossa GS, Cunnac S, Schmidt SM, Slamet-Loedin IH, Vera Cruz C, Szurek B, Frommer WB, White FF, Yang B. Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat Biotechnol. 2019;37(11):1344–50. https://doi.org/10.1038/s41587-019-0267-z.
Ortigosa A, Gimenez-Ibanez S, Leonhardt N, Solano R. Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol J. 2019;17(3):665–73. https://doi.org/10.1111/pbi.13006.
Paul JW 3rd, Qi Y. CRISPR/Cas9 for plant genome editing: accomplishments, problems and prospects. Plant Cell Rep. 2016;35(7):1417–27. https://doi.org/10.1007/s00299-016-1985-z.
Pixley KV, Falck-Zepeda JB, Paarlberg RL, Phillips PWB, Slamet-Loedin IH, Dhugga KS, Campos H, Gutterson N. Genome-edited crops for improved food security of smallholder farmers. Nat Genet. 2022;54(4):364–7. https://doi.org/10.1038/s41588-022-01046-7.
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell. 2013;152(5):1173–83. https://doi.org/10.1016/j.cell.2013.02.022.
Santosh Kumar VV, Verma RK, Yadav SK, Yadav P, Watts A, Rao MV, Chinnusamy V. CRISPR-Cas9 mediated genome editing of drought and salt tolerance (OsDST) gene in indica mega rice cultivar MTU1010. Physiol Mol Biol Plants. 2020;26(6):1099–110. https://doi.org/10.1007/s12298-020-00819-w.
Sathee L, Jagadhesan B, Pandesha PH, Barman D, Adavi BS, Nagar S, Krishna GK, Tripathi S, Jha SK, Chinnusamy V. Genome editing targets for improving nutrient use efficiency and nutrient stress adaptation. Front Genet. 2022;13:900897. https://doi.org/10.3389/fgene.2022.900897.
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu JL, Gao C. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol. 2013;31(8):686–8. https://doi.org/10.1038/nbt.2650.
Shillito RD, Whitt S, Ross M, Ghavami F, Vleesschauwer DD, D’Halluin K, Hoecke AV, Meulewaeter F. Detection of genome edits in plants—from editing to seed. In Vitro Cell Dev Biol Plant Spec Issue Gen Edit (Spec Issue Gen Edit). 2021. https://doi.org/10.1007/s11627-021-10214-z.
Smyth SJ. Genetically modified crops, regulatory delays, and international trade. Food Energy Secur. 2016;6(2):9. https://doi.org/10.1002/fes3.100.
Strobbe S, Wesana J, Van Der Straeten D, De Steur H. Public acceptance and stakeholder views of gene edited foods: a global overview. Trends Biotechnol. 2023;41(6):736–40. https://doi.org/10.1016/j.tibtech.2022.12.011.
Sturme MHJ, van der Berg JP, Bouwman LMS, De Schrijver A, de Maagd RA, Kleter GA, Battaglia-de Wilde E. Occurrence and nature of off-target modifications by CRISPR-Cas genome editing in plants. ACS Agric Sci Technol. 2022;2(2):192–201. https://doi.org/10.1021/acsagscitech.1c00270.
Sun Y, Shang L, Zhu QH, Fan L, Guo L. Twenty years of plant genome sequencing: achievements and challenges. Trends Plant Sci. 2022;27(4):391–401. https://doi.org/10.1016/j.tplants.2021.10.006.
Tuncel A, Pan C, Sprink T, Wilhelm R, Barrangou R, Li L, Shih PM, Varshney RK, Tripath L, Eck JV, Mandadi K, Yiping Q. Genome-edited foods. Nature Reviews Bioengineering. 2023;1:799–816.
UNEP (2022) Emissions gap report 2022: the closing window—climate crisis calls for rapid transformation of societies. Nairobi. http://www.unep.org/resources/emissions-gap-report-2022
van der Oost J, Patinios C. The genome editing revolution. Trends Biotechnol. 2023;41(3):396–409. https://doi.org/10.1016/j.tibtech.2022.12.022.
Waltz E. GABA-enriched tomato is first CRISPR-edited food to enter market. Nat Biotechnol. 2022;40(1):9–11. https://doi.org/10.1038/d41587-021-00026-2.
Wang JY, Doudna JA. CRISPR technology: a decade of genome editing is only the beginning. Science. 2023;379(6629):eadd8643. https://doi.org/10.1126/science.add8643.
Wang JY, Pausch P, Doudna JA. Structural biology of CRISPR-Cas immunity and genome editing enzymes. Nat Rev Microbiol. 2022;20(11):641–56. https://doi.org/10.1038/s41579-022-00739-4.
Watson JD, Crick FH. Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature. 1953;171(4356):737–8. https://doi.org/10.1038/171737a0.
Zhang Y, Bai Y, Wu G, Zou S, Chen Y, Gao C, Tang D. Simultaneous modification of three homoeologs of TaEDR1 by genome editing enhances powdery mildew resistance in wheat. Plant J. 2017;91(4):714–24. https://doi.org/10.1111/tpj.13599.
Zhang Y, Wu Y, Li G, Qi A, Zhang Y, Zhang T, Qi Y. Genome-wide investigation of multiplexed CRISPR-Cas12a-mediated editing in rice. Plant Genome. 2023;16(2):e20266. https://doi.org/10.1002/tpg2.20266.
Zhao H, Wolt JD. Risk associated with off-target plant genome editing and methods for its limitation. Emerg Top Life Sci. 2017;1(2):231–40. https://doi.org/10.1042/ETLS20170037.
Zhu H, Li C, Gao C. Applications of CRISPR-Cas in agriculture and plant biotechnology. Nat Rev Mol Cell Biol. 2020;21(11):661–77. https://doi.org/10.1038/s41580-020-00288-9.
Acknowledgements
I apologize to colleagues whose work has not been included in the manuscript.
Funding
Open Access funding provided by Agriculture & Agri-Food Canada. No funding was associated with this work. Basudev Ghoshal is supported by core funding from Agriculture and Agri-Food Canada.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no conflict of interest and no financial interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Corresponding Editor: Kailash C. Bansal; Reviewers: Ajay Singh, Ashwani Pareek.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ghoshal, B. A birds-eye-view on CRISPR-Cas system in agriculture. Nucleus (2024). https://doi.org/10.1007/s13237-023-00462-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13237-023-00462-2