Abstract
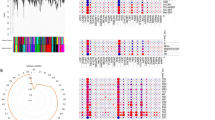
In this study, we integrated the gene expression data of sepsis to reveal more precise genome-wide expression signature to shed light on the pathological mechanism of sepsis. Differentially expressed genes via integrating five microarray datasets from the Gene Expression Omnibus database were obtained. The gene function and involved pathways of differentially expressed genes (DEGs) were detected by GeneCodis3. Transcription factors (TFs) targeting top 20 dysregulated DEGs (including up- and downregulated genes) were found based on the TRANSFAC. A total of 1339 DEGs were detected including 788 upregulated and 551 downregulated genes. These genes were mostly involved in DNA-dependent transcription regulation, blood coagulation, and innate immune response, pathogenic escherichia coli infection, epithelial cell signaling in helicobacter pylori infection, and chemokine signaling pathway. TFs bioinformatic analysis of 20 DEGs generated 374 pairs of TF-target gene involving 47 TFs. At last, we found that five top ten upregulated DEGs (S100A8, S100A9, S100A12, PGLYRP1 and MMP9) and three downregulated DEGs (ZNF84, CYB561A3 and BST1) were under the regulation of three hub TFs of Pax-4, POU2F1, and Nkx2–5. The identified eight DEGs may be regarded as the diagnosis marker and drug target for sepsis.

Similar content being viewed by others
References
Angus DC, Linde-Zwirble WT, Lidicker J, Clermont G, Carcillo J, Pinsky MR (2001) Epidemiology of severe sepsis in the United States: analysis of incidence, outcome, and associated costs of care. Crit Care Med 29:1303–1310
Annane D, Aegerter P, Jars-Guincestre MC, Guidet B (2003) Current epidemiology of septic shock: the CUB-Rea network. Am J Respir Crit Care Med 168:165–172. doi:10.1164/rccm.2201087
Assou S et al (2009) A gene expression signature shared by human mature oocytes and embryonic stem cells. BMC Genom 10:10. doi:10.1186/1471-2164-10-10
Brunkhorst FM, Reinhart K (2009) Diagnosis and causal treatment of sepsis. Der Internist 50:810–816. doi:10.1007/s00108-008-2287-5
Calvano SE et al (2005) A network-based analysis of systemic inflammation in humans. Nature 437:1032–1037. doi:10.1038/nature03985
Cazalis MA et al (2014) Early and dynamic changes in gene expression in septic shock patients: a genome-wide approach. Intensive Care Med Exp 2:20. doi:10.1186/s40635-014-0020-3
Chang KH, Wu YR, Chen YC, Fung HC, Lee-Chen GJ, Chen CM (2015) STK39, But Not BST1, HLA-DQB1, and SPPL2B polymorphism, is associated with Han-Chinese Parkinson’s disease in Taiwan. Medicine 94:e1690. doi:10.1097/md.0000000000001690
de Montmollin E, Annane D (2011) Year in review 2010: critical care-multiple organ dysfunction and sepsis. Crit care 15:236. doi:10.1186/cc10359
Diboun I, Wernisch L, Orengo CA, Koltzenburg M (2006) Microarray analysis after RNA amplification can detect pronounced differences in gene expression using limma. BMC Genom 7:252. doi:10.1186/1471-2164-7-252
Dziarski R, Platt KA, Gelius E, Steiner H, Gupta D (2003) Defect in neutrophil killing and increased susceptibility to infection with nonpathogenic gram-positive bacteria in peptidoglycan recognition protein-S (PGRP-S)-deficient mice. Blood 102:689–697. doi:10.1182/blood-2002-12-3853
Foell D et al (2013) Proinflammatory S100A12 can activate human monocytes via Toll-like receptor 4. Am J Respir Crit Care Med 187:1324–1334. doi:10.1164/rccm.201209-1602OC
Galliera E, Tacchini L, Corsi Romanelli MM (2015) Matrix metalloproteinases as biomarkers of disease: updates and new insights. Clin Chem Lab Med 53:349–355. doi:10.1515/cclm-2014-0520
Hoesel B, Schmid JA (2013) The complexity of NF-kappaB signaling in inflammation and cancer. Mol cancer 12:86. doi:10.1186/1476-4598-12-86
Inoue H, Nomiyama J, Nakai K, Matsutani A, Tanizawa Y, Oka Y (1998) Isolation of full-length cDNA of mouse PAX4 gene and identification of its human homologue. Biochem Biophys Res Commun 243:628–633. doi:10.1006/bbrc.1998.8144
Ishihara K, Hirano T (2000) BST-1/CD157 regulates the humoral immune responses in vivo. Chem Immunol 75:235–255
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kang D, Liu G, Lundstrom A, Gelius E, Steiner H (1998) A peptidoglycan recognition protein in innate immunity conserved from insects to humans. Proc Natl Acad Sci USA 95:10078–10082
Kesh K et al (2015) Association of MMP7-181A→G promoter polymorphism with gastric cancer risk: influence of nicotine in differential allele-specific transcription via increased phosphorylation of cAMP-response element-binding protein (CREB). J Biol Chem 290:14391–14406. doi:10.1074/jbc.M114.630129
Komuro I, Izumo S (1993) Csx: a murine homeobox-containing gene specifically expressed in the developing heart. Proc Natl Acad Sci USA 90:8145–8149
Kurz T, Gustafsson B, Brunk UT (2006) Intralysosomal iron chelation protects against oxidative stress-induced cellular damage. FEBS J 273:3106–3117. doi:10.1111/j.1742-4658.2006.05321.x
Lints TJ, Parsons LM, Hartley L, Lyons I, Harvey RP (1993) Nkx-2.5: a novel murine homeobox gene expressed in early heart progenitor cells and their myogenic descendants. Development 119:419–431
Liu C, Gelius E, Liu G, Steiner H, Dziarski R (2000) Mammalian peptidoglycan recognition protein binds peptidoglycan with high affinity, is expressed in neutrophils, and inhibits bacterial growth. J Biol Chem 275:24490–24499. doi:10.1074/jbc.M001239200
Liu C, Xu Z, Gupta D, Dziarski R (2001) Peptidoglycan recognition proteins: a novel family of four human innate immunity pattern recognition molecules. J Biol Chem 276:34686–34694. doi:10.1074/jbc.M105566200
Lopez-Bergami P, Lau E, Ronai Z (2010) Emerging roles of ATF2 and the dynamic AP1 network in cancer. Nat Rev Cancer 10:65–76. doi:10.1038/nrc2681
McKie AT et al (2001) An iron-regulated ferric reductase associated with the absorption of dietary iron. Science 291:1755–1759. doi:10.1126/science.1057206 (New York, NY)
Mordvinov VA et al (1999) Binding of YY1 and Oct1 to a novel element that downregulates expression of IL-5 in human T cells. J Allergy Clin Immunol 103:1125–1135
Morone S et al (2012) Overexpression of CD157 contributes to epithelial ovarian cancer progression by promoting mesenchymal differentiation. PLoS One 7:e43649. doi:10.1371/journal.pone.0043649
Mullen M, Gonzalez-Perez RR (2016) Leptin-Induced JAK/STAT signaling and cancer growth. Vaccines 4:26. doi:10.3390/vaccines4030026
Nakamura T, Ebihara I, Shimada N, Shoji H, Koide H (1998) Modulation of plasma metalloproteinase-9 concentrations and peripheral blood monocyte mRNA levels in patients with septic shock: effect of fiber-immobilized polymyxin B treatment. Am J Med Sci 316:355–360
Pal S, Bhattacharjee A, Ali A, Mandal NC, Mandal SC, Pal M (2014) Chronic inflammation and cancer: potential chemoprevention through nuclear factor kappa B and p53 mutual antagonism. J Immunol 11:23. doi:10.1186/1476-9255-11-23
Park SY, Jing X, Gupta D, Dziarski R (2013) Peptidoglycan recognition protein 1 enhances experimental asthma by promoting Th2 and Th17 and limiting regulatory T cell and plasmacytoid dendritic cell responses. J Immunol 190:3480–3492. doi:10.4049/jimmunol.1202675
Pellegrina DV, Severino P, Machado MC, Pinheiro da Silva F, Reis EM (2015) Microarray gene expression analysis of neutrophils from elderly septic patients. Genom Data 6:51–53. doi:10.1016/j.gdata.2015.08.013
Phillips K, Luisi B (2000) The virtuoso of versatility: POU proteins that flex to fit. J Mol Biol 302:1023–1039. doi:10.1006/jmbi.2000.4107
Ramana KV, Tammali R, Srivastava SK (2010) Inhibition of aldose reductase prevents growth factor-induced G1-S phase transition through the AKT/phosphoinositide 3-kinase/E2F-1 pathway in human colon cancer cells. Mol Cancer Ther 9:813–824. doi:10.1158/1535-7163.mct-09-0795
Roth J, Vogl T, Sorg C, Sunderkotter C (2003) Phagocyte-specific S100 proteins: a novel group of proinflammatory molecules. Trends Immunol 24:155–158
Ryan AK, Rosenfeld MG (1997) POU domain family values: flexibility, partnerships, and developmental codes. Genes Dev 11:1207–1225
Smoot ME, Ono K, Ruscheinski J, Wang PL, Ideker T (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27:431–432. doi:10.1093/bioinformatics/btq675
Speake C et al (2015) An interactive web application for the dissemination of human systems immunology data. J Trans Med 13:196. doi:10.1186/s12967-015-0541-x
Standage SW, Wong HR (2011) Biomarkers for pediatric sepsis and septic shock. Expert Rev Anti Infect Ther 9:71–79. doi:10.1586/eri.10.154
Tabas-Madrid D, Nogales-Cadenas R, Pascual-Montano A (2012) GeneCodis3: a non-redundant and modular enrichment analysis tool for functional genomics. Nucleic Acids Res 40:W478–W483. doi:10.1093/nar/gks402
Terrin G et al (2011) Serum calprotectin: an antimicrobial peptide as a new marker for the diagnosis of sepsis in very low birth weight newborns. Clin Dev Immunol 2011:291085. doi:10.1155/2011/291085
Tydell CC, Yount N, Tran D, Yuan J, Selsted ME (2002) Isolation, characterization, and antimicrobial properties of bovine oligosaccharide-binding protein. A microbicidal granule protein of eosinophils and neutrophils. J Biol Chem 277:19658–19664. doi:10.1074/jbc.M200659200
Vandooren J, Van den Steen PE, Opdenakker G (2013) Biochemistry and molecular biology of gelatinase B or matrix metalloproteinase-9 (MMP-9): the next decade. Crit Rev Biochem Mol Biol 48:222–272. doi:10.3109/10409238.2013.770819
Walther C et al (1991) Pax: a murine multigene family of paired box-containing genes. Genomics 11:424–434
Wang D, DuBois RN (2010) Therapeutic potential of peroxisome proliferator-activated receptors in chronic inflammation and colorectal cancer. Gastroenterol Clin North Am 39:697–707. doi:10.1016/j.gtc.2010.08.014
Wingender E (2008) The TRANSFAC project as an example of framework technology that supports the analysis of genomic regulation. Brief Bioinform 9:326–332. doi:10.1093/bib/bbn016
Yepes S, Torres MM, Andrade RE (2015) Clustering of expression data in chronic lymphocytic leukemia reveals new molecular subdivisions. PLoS One 10:e0137132. doi:10.1371/journal.pone.0137132
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14. doi:10.1186/gb-2010-11-2-r14
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, J., Cheng, Y., Duan, M. et al. Unveiling differentially expressed genes upon regulation of transcription factors in sepsis. 3 Biotech 7, 46 (2017). https://doi.org/10.1007/s13205-017-0713-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-0713-x