Abstract
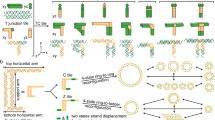
The science of DNA nanotechnology has led to the synthesis of size-controllable materials with uniformed geometries at nano- and micro-scales. Herein, we proposed the double-crossover (DX), antiparallel, and even half-turns perimeter tiles (DAE tiles) to synthesize intact, mono-crystalline, and giant 2D DNA micro-assemblies. The DNA tiles with 10-half-turns perimeter were synthesized via self-assembly of 106 nucleotides (NT) circular scaffold along with the complimentary staple strands. The DAE DNA tiles were successfully polymerized to achieve stable lattices by adjusting the inter-tile distances of certain lengths for attaining torsional (or twisting) chirality. We determined that the inter-tile connections affected the degree of coiling (or super-coiling) and twisting forces (right- or left-handed twists) in the DNA helix. While the degree of polymerization of DNA tiles was also tune-able by controlling the lengths and structural designs of the circular core of the DNA tiles. Furthermore, the number of half-turns in the core and on the connection arms (4 or 5) with even “E” or odd “O” half-turns was crucial. It affected the direction of winding of DNA duplexes to alter the overall stiffness and sturdiness of DNA lattices. The number of half-turns in the connections were either “4 or Even (21 NT); E with 5 NT sticky ends” or “5 Odd (26 NT); O with 6 NT sticky ends” (DAE-E or DAE-O tile systems). The AFM results revealed that the above tile systems (DAE-E or DAE-O) together with the locations of crossovers, and holiday junctions along the DNA tiles controlled the left- or right-handed coiling of DNA double strands. This phenomenon affected the compactness of resulting DNA motifs, the overall intrinsic curvatures, double-strand packing, and the geometry of DNA lattices.
Graphical abstract









Similar content being viewed by others
References
Aghebat Rafat A, Pirzer T, Scheible MB et al (2014) Surface-assisted large-scale ordering of DNA origami tiles. Angew Chemie Int Ed 53:7665–7668. https://doi.org/10.1002/anie.201403965
Ali MM, Li F, Zhang Z et al (2014) Rolling circle amplification: a versatile tool for chemical biology, materials science and medicine. Chem Soc Rev 43:3324–3341
Baig MMFA, Dissanayaka WL, Zhang C (2021a) 2D DNA nanoporous scaffold promotes osteogenic differentiation of pre-osteoblasts. Int J Biol Macromol 188:657–669. https://doi.org/10.1016/j.ijbiomac.2021.07.198
Baig MMFA, Zhang C, Akhtar MF et al (2021b) Treatment of Wilms’ nephroblastoma cancer cells via EGFR targeting of dactinomycin loaded DNA-nanowires. J Pharm Investig 51:233–242. https://doi.org/10.1007/s40005-020-00509-5
Barua R, Das S (2003) Finite field arithmetic using self-assembly of DNA tilings. In: 2003 Congress on Evolutionary Computation, CEC 2003—Proceedings. pp 2529–2536
Zion MY Ben, He X, Maass CC, et al (2017) Self-assembled three-dimensional chiral colloidal architecture
Brady RA, Brooks NJ, Foderà V et al (2018) Amphiphilic-DNA platform for the design of crystalline frameworks with programmable structure and functionality. J Am Chem Soc 140:15384–15392. https://doi.org/10.1021/jacs.8b09143
Dohno C, Makishi S, Nakatani K, Contera S (2017) Amphiphilic DNA tiles for controlled insertion and 2D assembly on fluid lipid membranes: the effect on mechanical properties. Nanoscale 9:3051–3058. https://doi.org/10.1039/c6nr07084a
Hong F, Jiang S, Lan X et al (2018) Layered-crossover tiles with precisely tunable angles for 2D and 3D DNA crystal engineering. J Am Chem Soc 140:14670–14676. https://doi.org/10.1021/jacs.8b07180
Kempter S, Khmelinskaia A, Strauss MT et al (2019) Single particle tracking and super-resolution imaging of membrane-assisted stop-and-go diffusion and lattice assembly of DNA origami. ACS Nano 13:996–1002. https://doi.org/10.1021/acsnano.8b04631
Li M, Zuo H, Yu J et al (2017) One DNA strand homo-polymerizes into defined nanostructures. Nanoscale 9:10601–10605. https://doi.org/10.1039/c7nr03640j
Lopez-Gomollon S, Nicolas FE (2013) Purification of DNA oligos by denaturing polyacrylamide gel electrophoresis (PAGE). Methods Enzymol 529:65–83. https://doi.org/10.1016/B978-0-12-418687-3.00006-9
Lubbe AS, Liu Q, Smith SJ et al (2018) Photoswitching of DNA hybridization using a molecular motor. J Am Chem Soc 140:5069–5076. https://doi.org/10.1021/jacs.7b09476
Mao X, Li K, Liu M et al (2019) Directing curli polymerization with DNA origami nucleators. Nat Commun 10:1–5. https://doi.org/10.1038/s41467-019-09369-6
Marin-Gonzalez A, Vilhena JG, Moreno-Herrero F, Perez R (2019) DNA crookedness regulates DNA mechanical properties at short length scales. Phys Rev Lett 122:1–8. https://doi.org/10.1103/PhysRevLett.122.048102
Marras AE, Zhou L, Su HJ, Castro CE (2015) Programmable motion of DNA origami mechanisms. Proc Natl Acad Sci U S A 112:713–718. https://doi.org/10.1073/pnas.1408869112
Martens K, Binkowski F, Nguyen L et al (2021) Long- and short-ranged chiral interactions in DNA-assembled plasmonic chains. Nat Commun 12:1–9. https://doi.org/10.1038/s41467-021-22289-8
Mitchell JC, Harris JR, Malo J et al (2004) Self-assembly of chiral DNA nanotubes. J Am Chem Soc 126:16342–16343. https://doi.org/10.1021/ja043890h
Nakayama Y, Yamaguchi H, Einaga N, Esumi M (2016) Pitfalls of DNA quantification using dnabinding fluorescent dyes and suggested solutions. PLoS ONE 11:e0150528. https://doi.org/10.1371/journal.pone.0150528
O’Neill P, Rothemund PWK, Kumar A, Fygenson DK (2006) Sturdier DNA nanotubes via ligation. Nano Lett 6:1379–1383. https://doi.org/10.1021/nl0603505
Ohya Y, Miyoshi N, Hashizume M et al (2012) Formation of 1D and 2D gold nanoparticle arrays by divalent DNA-gold nanoparticle conjugates. Small 8:2335–2340. https://doi.org/10.1002/smll.201200092
Rangnekar A, Labean TH (2014) Building DNA nanostructures for molecular computation, templated assembly, and biological applications. Acc Chem Res 47:1778–1788. https://doi.org/10.1021/ar500023b
Sato Y, Endo M, Morita M et al (2018) Environment-dependent self-assembly of DNA origami lattices on phase-separated lipid membranes. Adv Mater Interfaces 5:1–9. https://doi.org/10.1002/admi.201800437
Shi X, Lu W, Wang Z et al (2014) Programmable DNA tile self-assembly using a hierarchical sub-tile strategy. Nanotechnology 25:1–10. https://doi.org/10.1088/0957-4484/25/7/075602
Shin J, Kim J, Park SH, Ha TH (2018) Kinetic trans-assembly of DNA nanostructures. ACS Nano 12:9423–9432. https://doi.org/10.1021/acsnano.8b04639
Sung HP, Finkelstein G, LaBean TH (2008) Stepwise self-assembly of DNA tile lattices using dsDNA bridges. J Am Chem Soc 130:40–41. https://doi.org/10.1021/ja078122f
Tagawa M, Shohda KI, Fujimoto K, et al (2007) Heat-resistant DNA tile arrays constructed by template-directed photoligation through 5-carboxyvinyl-2′-deoxyuridine. Nucleic Acids Res 35:. https://doi.org/10.1093/nar/gkm872
Valdés A, Martínez-García B, Segura J et al (2019) Quantitative disclosure of DNA knot chirality by high-resolution 2D-gel electrophoresis. Nucleic Acids Res 47:e29. https://doi.org/10.1093/nar/gkz015
Wang P, Gaitanaros S, Lee S et al (2016) Programming self-assembly of DNA origami honeycomb two-dimensional lattices and plasmonic metamaterials. J Am Chem Soc 138:7733–7740. https://doi.org/10.1021/jacs.6b03966
Winfree E, Liu F, Wenzler LA, Seeman NC (1998) Design and self-assembly of two-dimensional DNA crystals. Nature 394:539–544. https://doi.org/10.1038/28998
Zhang C, Shao PG, Van Kan JA, Van Der Maarel JRC (2009) Macromolecular crowding induced elongation and compaction of single DNA molecules confined in a nanochannel. Proc Natl Acad Sci U S A 106:16651–16656. https://doi.org/10.1073/pnas.0904741106
Zhang B, Qin X, Zhou M et al (2021) Tetrahedral DNA nanostructure improves transport efficiency and anti-fungal effect of histatin 5 against Candida albicans. Cell Prolif 54:1–9. https://doi.org/10.1111/cpr.13020
Zhao Z, Liu Y, Yan H (2011) Organizing DNA origami tiles into larger structures using preformed scaffold frames. Nano Lett 11:2997–3002. https://doi.org/10.1021/nl201603a
Acknowledgements
The work was supported by Hong Kong Research Grant Council (RGC) funding projects (GRF#16308818, GRF#16309920, and GRF#16309421), Hong Kong Innovation and Technology Commission (HKITC) funding project (MHP/003/19) and the Social Development Project of Guizhou Department of Science and Technology ([No.2020]4Y214).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Baig, M.M.F.A., Gao, X., Farid, A. et al. Synthesis of stable 2D micro-assemblies of DNA tiles achieved via intrinsic curvatures in the skeleton of DNA duplexes coupled with the flexible support of the twisted side-arms. Appl Nanosci 13, 4279–4289 (2023). https://doi.org/10.1007/s13204-022-02616-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13204-022-02616-1