Abstract
Marine karst ecosystems exist at the land-sea interface and are characterised by underwater formations sculpted over time by the action of seawater. Submerged caves and crevices of these ecosystems host a rich array of marine life of which sponges are among the most abundant and diverse components. In the present study, we describe elements of the sponge fauna sampled from a unique karst ecosystem at a remote island, Orchid Island, off the southeastern coast of Taiwan. The present study includes several understudied sponge taxa, including sclerosponges (Acanthochaetetes wellsi, and Astrosclera willeyana) and several lithistid species from dark, shallow-water caves. Prokaryotic communities were obtained from a total of 22 demosponge species, of which 11 are potentially new to science. The tetracladinid, lithistids harboured prokaryotic communities, which clustered separately from all other sponge species, contrasting with the non-tetracladinid, lithistid Vetulina incrustans. The tetracladinid, lithistids, furthermore, formed two distinct clusters with species of the Spirophorina suborder clustering apart from those of the Astrophorina suborder. The sclerosponge A. wellsi also harboured a distinct prokaryotic community in terms of composition including five unique, abundant OTUs with relatively low sequence similarities to organisms in GenBank. All cave sponges were enriched with SAR202 members, a group of bacteria known for their role in the degradation of recalcitrant compounds. The highest relative abundance of SAR202 was found in A. wellsi. We propose that the cave sponges of Orchid Island may play an as-yet uncharted role in nutrient dynamics at the land-sea interface.
Similar content being viewed by others
Introduction
Marine karst areas are unique ecosystems of spellbinding beauty and considerable potential scientific importance. They are often renowned for their striking geological formations, adorned with intricate rock structures. Clinging to the coast, they consist of intricate networks of pools, crevices, and caves with a plethora of life forms inhabiting these varying habitats. Situated along rocky coastlines, they are subject to ebb and flow such that the flora and fauna have adapted to fluctuations in temperature, light intensity, and salinity (Meroz-Fine et al. 2005; Sket 1996). Sought-after destinations for diving and ecotourism, among other activities, they have come under increasing scientific scrutiny although much of this focus has been heavily skewed to the Mediterranean (Gerovasileiou and Voultsiadou 2012; Harmelin 1997; Sket 1996; but see Ise et al. 2023; Schuster et al. 2021).
Sponges are among the most abundant and species-rich components of marine karst ecosystems, particularly in light-deprived, karst caves (Gerovasileiou et al. 2016; Pisera and Gerovasileiou 2021). Karst caves can also host taxa typically found in the tenebrous depths of the ocean, as exemplified by recent discoveries of carnivorous and glass sponges in Mediterranean caves (Bakran-Petricioli et al. 2007; Vacelet 1996; Vacelet et al. 1994). Another compelling aspect of marine karst ecosystems lies in their potential for unearthing rare and previously undocumented species, including a significant number of species unique to these environments (Gerovasileiou et al. 2015; Iliffe and Kornicker 2009; Manconi and Serusi 2008).
In August 2018, we encountered a highly interesting marine karst ecosystem while snorkeling in a large tidal pool at a small island called Orchid Island (also known as Lanyu). Orchid Island is located in the Pacific Ocean about 68 km off the East coast of Taiwan and about 140 km from the northern Philippines. The island harbours many endemic terrestrial species (Hsieh 2002). Furthermore, despite several research efforts focusing on the coral reefs of Orchid Island, the karst pool and cave systems have remained neglected and understudied (Chao 2002; Chao and Lee 2002; Chen et al. 2022a, b; Kao et al. 2007; Lee 1980; Lee and Lee 1981; Reigle 1963; Yu 1985).
The subtidal cave system consists of several, very shallow tunnels connected via intertidal pools. We only explored the tunnels close to the pools, and these were all very shallow. The tunnels and caves were decorated by an abundance of small calcareous sponges, in addition to lithistid sponges and sclerosponges with specimens of the sclerosponge Acanthochaetetes wellsi up to 20 cm in length. Several potentially new lithistid species were sampled, which belonged to relatively rare genera including Manihinea (family Theonellidae), Neophrissospongia (family Corallistidae), and Scleritoderma (family Scleritodermidae).
With respect to their associated prokaryotic communities, the serendipitous discovery of the sponges inhabiting the karst ecosystem of Orchid Island enabled us to compare several sponge species free from the confounding effects of large-scale spatial processes and differences in local environmental conditions. Despite initial findings, which suggested that sponge composition was temporally and spatially ‘stable’ (Cárdenas et al. 2018; Hentschel et al. 2002; Hill et al. 2006; Pita et al. 2013; Reveillaud et al. 2014; Taylor et al. 2005;), a number of studies have now shown that the prokaryotic compositions of several sponge species differ among localities and as a function of geographic distance. Cleary et al. (2022), for example, showed that 55% of the variation in prokaryotic composition of the sponge species Hyrtios erectus could be explained by geographic distance after controlling for variation due to environmental conditions.
The primary goal of the present study was to provide a preliminary description of the sponges of the karst ecosystem of Orchid Island and their associated prokaryotic communities. For the first time, the prokaryotic communities of two, sympatrically occurring sclerosponge species, Acanthochaetetes wellsi and Astrosclera willeyana, were compared. We also assessed prokaryotic communities of a range of lithistid sponges in order to explore the species-level variation in the microbiomes of this understudied group, in addition to comparing these species to the other sampled sponge taxa. Lithistid sponges have been generally neglected with the notable exception of the species Theonella swinhoei, which has been extensively studied and characterised as a high microbial abundance (HMA) species (Lurgi et al. 2019). Kuo et al. (2019) also studied the bacterial composition of T. swinhoei using pyrosequencing. We also examined two phototrophic sponge species and determined their dominant symbionts. For all studied species, we identified abundant OTUs and compared these with homologous sequences in the GenBank database.
Material and methods
Study area
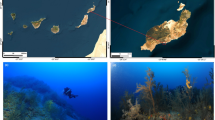
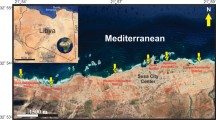
Orchid Island, with an area of 48.4 km2 and a coastline of 38.5 km, and the adjacent islet (Hsiao Lanyu) lie 70 km to the east of the southern tip of mainland Taiwan in the western Pacific Ocean (Fig. 1). Sponge samples were collected across a total of nine different sites. These two islands are the northward extension of the Luzon volcanic arch formed by volcanic eruptions between 3.5–1.4 and 0.04–0.02 Ma, respectively, which is driven by the convergence boundary between the Eurasian and the Philippine Sea plates (Chen et al. 19901992; Wang and Burnett 1990; Yang et al. 1996). Geologically, Orchid Island mainly consists of igneous andesite intermingled with basalt, foraminifera limestone, laterite platforms, and coral rocks uplifted at a rate of 3.2 mm annually during the Pleistocene and Holocene (Chen 1993; Richard et al. 1986; Wang and Burnett 1990). The outermost coral rock and fringing reefs, which have been exposed to intense wave erosion, have sculpted an intricate network of caves and tunnels interconnected to the open sea (Fig. 2a).
a Drone image of submarine caves taken by JKZ Cleary; b photograph of pool habitat and entrance to submerged caves taken by DFR Cleary; c photograph of the sclerosponge Acanthochaetetes wellsi taken by DFR Cleary; d photograph of Petrosia corticata (the green-grey sponge) and Manihinea sp. (the red sponge) taken by DFR Cleary
Part of the subtidal and wave-cut groove habitats, the karst ecosystem of Orchid Island is prominently featured along its coastline. Pools connected to the open sea via underwater tunnels and canyons experience regular tidal fluctuations (Fig. 2b). In addition to this, subsurface freshwater flows into the system from the island resulting in pronounced salinity fluctuations close to land. The surface areas of the pools range from several to hundreds of square meters and reach depths of 3 to 12 meters. Caves and tunnels, varying in diameter from less than a meter to a few meters, form complex mazes and encompass environments with limited to no light penetration.
Sampling
All sponge specimens were collected using snorkeling and SCUBA diving. All specimens were photographed in their natural environment, and a fragment was collected using a scalpel. Sponge specimens were initially identified as morphospecies in the field, and between 1 and 5 specimens were collected per morphospecies; in addition to sponge samples, sediment samples were collected from the top 5 cm surface layer using a sterile Falcon tube (Supplementary data 1). Upon removal from the water, a section from each sponge specimen was carefully excised and promptly placed in a vial containing 96% alcohol. Care was taken to include surface and interior tissue of each specimen. The vials were shipped to the Netherlands and stored in a − 20 °C freezer until subsequent DNA extraction. Sponge species identification was carried out using traditional taxonomic characters with a Leica microscope; samples from all species have been deposited at the Naturalis Biodiversity Center (as RMNH POR), Leiden, the Netherlands.
DNA extraction
DNA was extracted using the Qiagen DNeasy Powersoil extraction kit (Qiagen, Venlo, the Netherlands). A maximum of 250 mg of sponge tissue was used per sample; tissue was taken from all sides of the specimen (outside to core, and if applicable top, middle, and bottom of the sample). Sediment DNA extraction utilised 250 mg of wet sediment. The manufacturer’s protocol was followed with the exception of the initial vortexing step, which was carried out using the Qiagen TissueLyser II (Qiagen NV, Hilden, Germany). Sponge tissue was cut into small pieces using sterilised tweezers and scalpel blades and transferred to PowerBead Pro tubes containing ceramic and silica beads of different sizes. An extraction blank, in which no tissue was added to the PowerBead Pro tubes, was also included. Library preparation involved a two-step PCR protocol for all samples in addition to two negative controls (mQ water instead of template DNA) and the extraction blank. For the first PCR, the V3-V4 regions of the 16S rRNA gene were targeted and amplified using the primers 314F/ S-D-Bact-0785-a-A-21 (5′-CCTACGGGNGGCWGCAG-3′/5′-GACTACHVGGGTATCTAATCC-3′; Klindworth et al. 2013) with added 5′ Nextera transposase adaptors using the KAPA HiFi HotStart Ready Mix PCR Kit with a T100 Thermal Cycler (Bio-Rad, Hercules, CA, USA). The following PCR conditions were used: initial denaturation at 95 °C for 3 min, 30 cycles of denaturation at 98 °C for 20 s, annealing at 55 °C for 30 s, followed by extension at 72 °C for 30 s. The final extension was carried out at 72 °C for 1 min. PCR success was confirmed on an E-Gel™ (agarose gels at 2%), and the absence of amplification was validated for the negative controls and the extraction blank. PCR products were then cleaned with NucleoMag NGS-Beads (bead volume at 0.9 times the total volume of the sample, Macherey Nagel, Düren, Germany) using the VP 407AM-N 96 Pin Magnetic Bead Extractor stamp (V&P Scientific, San Diego, CA, USA). For the second PCR, the cleaned PCR products (1 µL each) were amplified and labelled using the MiSeq Nextera XT DNA library preparation kit (Illumina, San Diego, CA, USA) with the same thermal cycling scheme limited to 8 cycles. PCR products were then analysed with the Fragment Analyser Agilent 5300 using the DNF-910–33 dsDNA Reagent Kit (35–1500 bp) protocol (Agilent Technologies, Santa Clara, CA, USA) to confirm successful labelling of the DNA fragments. Negative controls and extraction blanks remained negative after this step. Pooling at equimolar concentration was performed with QIAgility 2 (Qiagen, Hilden, Germany). The final pool was then cleaned with NucleoMag NGSBeads, eluted in Milli-Q, and the DNA concentration was verified using Tapestation 4150 (Kit HSD 5000, Agilent Technologies, Santa Clara, CA, USA). Paired-end sequence reads were generated with an Illumina MiSeq v3 PE300 platform at BaseClear B.V. (Leiden, the Netherlands). FASTQ read sequence files were generated using bcl2fastq version 2.20 (Illumina). Initial quality assessment was based on data passing the Illumina Chastity filtering. Subsequently, reads containing PhiX control signal were removed using an in-house filtering protocol. In addition to this, reads containing (partial) adapters were clipped (up to a minimum read length of 50 bps). The second quality assessment was based on the remaining reads using the FASTQC quality control tool version 0.11.8.
Sequencing analysis
The 16S rRNA amplicon libraries were analysed using QIIME2 (version 2019.7; Bolyen et al. 2019). Raw data were imported yielding a demultiplexed ‘qza’ data file (artifact). The DADA2 plugin (Callahan et al. 2016) in QIIME 2 was subsequently used to trim sequences (final length 400 nt). The DADA2 analysis yielded output archives containing an OTU (at a 100% similarity threshold, also known as amplicon sequence variant or ‘ASV’) table, denoising stats, and a fasta file of representative sequences. The feature-classifier plugin with the extract-reads method was then used with the i-sequences argument set to silva-138–99-seqs.qza. This was followed by the feature-classifier plugin with the fit-classifier-naive-bayes method, and the i-reference-taxonomy method set to silva-138–99-tax.qza. Both silva-138 files can be obtained from https://docs.qiime2.org/2020.8/data-resources/?highlight=silva. The feature-classifier plugin was then used with the classify-sklearn method, and the i-reads argument was set to the representative sequences file generated by the DADA2 analysis to produce a table with taxonomic classifications for all OTUs. Finally, mitochondria, chloroplasts, and Eukaryota were filtered out using the qiime taxa plugin with the filter-table method. All OTUs unclassified at domain and phylum level were also removed. The OTU and taxonomy tables were subsequently merged in R (R Core Team 2022). Accession numbers of closely related organisms to selected OTUs were obtained using NCBI Basic Local Alignment Search Tool (BLAST) (Altschul et al. 1990). Sequences used in this study have been uploaded to the NCBI ShortRead Archive (BioProject nr: PRJNA865713 and BioSample nr: SUB11909922).
Statistical analyses
A table containing the OTU counts was imported into R. Supplementary data 2 contains all operational taxonomic unit (OTU) counts per sample and taxonomic classifications of all OTUs. The 50 most abundant OTUs are summarised in Supplementary data 3 including the results of the BLAST analyses. The raw OTU counts matrix was used to compare diversity and higher taxon abundance among groups. Diversity indices, namely, rarefied richness, evenness, Shannon’s H′, and Fisher’s alpha, were obtained using the rarefy and diversity functions from the vegan (Oksanen et al. 2020) package in R. Evenness (Pielou’s J) was calculated by dividing Shannon’s H′ by the number of OTUs in each sample. Variation in prokaryotic composition among groups (sponge species and sediment) was visualised with Principal Coordinates Analysis (PCO), and we tested for significant differences among groups with an adonis analysis from the vegan package. Subsequent statistical analyses were limited to groups with at least three samples. For the PCO, a Bray–Curtis distance matrix was first obtained using the phyloseq package (McMurdie and Holmes 2013) whereby the count data was rarefied using the rarefy_even_depth function with the sample.size argument set to the minimum sample size (n = 6978) and subsequently log10 transformed.
Results
A preliminary description of several sponge species from Orchid Island’s marine karst ecosystem
Prokaryotic communities were obtained from a total of 22 sponge species representing a total of nine orders and 17 families. All were members of the class Demospongiae. The majority of species were restricted to low-light, karst cave habitats. The following sponge species were sampled from submarine caves and canyons, Acanthochaetetes wellsi Hartman & Goreau, 1975, Acanthostylotella cornuta (Topsent, 1897), Aciculites ciliata Wilson, 1925, Asteropus cf. simplex, Astrosclera willeyana Lister, 1900, Petrosia corticata (Wilson, 1925), Sollasipelta ornata (Sollas, 1888), and Vetulina incrustans Schuster et al., 2018. Several species could not be conclusively identified to species level and are potentially new species. These are the following: Theonella aff. timmi, Svenzea aff. devoogdae, Discodermia sp., Manihinea sp., Neophrissospongia sp., Penares sp., Scleritoderma sp., Xestospongia sp., Polymastia sp., and Theonella sp.. Two species, Lamellodysidea herbacea (Keller, 1889) and Lamellodysidea sp., were sampled from pool habitat, while Xestospongia testudinaria Lamarck, 1815 and Stylissa carteri Dendy, 1889 were sampled from nearby coral reef habitat (Supplementary data 1). Formal descriptions of all potentially new species identified in this study are underway.
Only three specimens of the sclerosponge Astrosclera willeyana were observed from three different sites, and they all had the usual pyriform, half-spherical growth form. Acanthochaetetes wellsi was found in a wide range of shapes, and formed large, stalactite-like formations and was highly abundant and dominant in certain caves (Fig. 2c).
The larger sclerosponges coexisted within their cave and canyon habitats with a large number of smaller species, including a diverse array of calcareous and lithistid sponge species. Lithistid demosponges emerged as dominant and species-rich components of the cave fauna. The specimens sampled encompassed species belonging to the orders Tetractinellida and Sphaercladina representing five different families. Aside from the tetractinellid, lithistid sponges, we also sampled a single non-tetractinellid, lithistid sponge species, Vetulina incrustans.
To date, we have successfully identified nine tetractinellid species of which five are potentially novel. Of the order Tetractinellida, the most species-rich group belonged to the family Theonellidae with four potentially new species in the genera Theonella, Maninihea, and Discodermia. Within the family Scleritodermidae, two species were observed, the large flabelliform Aciculites ciliata and a potentially new species of Scleritoderma with an unusual tube-forming morphology. A small globular specimen of Sollasipelta ornata (Neopeltidae) and a potentially new species of Neophrissospongia (Corallistidae) were sampled. The latter species was abundant at many different sites and formed cylindrical, fused masses and had a brownish colour. Apart from the lithistid tetractinellids, two non-lithistid sponge species belonging to the order Tetractinellida were sampled, namely, Asteropus cf. simplex and a potentially novel species of Penares.
In addition to the sclerosponges and tetractinellids, an array of other sponge taxa across a range of orders were identified including potentially novel species. Three species belonging to the order Haplosclerida (Petrosiidae) were observed. The giant barrel sponge, Xestospongia testudinaria, was found at water depths exceeding 20 meters in coral reefs nearby the karst ecosystem. The green/grey Petrosia (Strongylophora) corticata (Wilson, 1925) was relatively abundant in overhangs and caves (Fig. 2d), and formed large cylindrical branches and unlike other Petrosiidae was parchment-like and soft. An unusual, potentially new Petrosiidae species was observed from crevices and caves; it had a pink colouration and turned dark brown/black upon preservation and has been tentatively identified as Xestospongia sp.. Two species belonging to the Scopalinidae family were observed, the very common, orange reef species Stylissa carteri from nearby coral reefs and a potentially new species of Svenzea. One species belonging to the order Polymastiida was observed, namely, Polymastia sp.. The cyanobacterial sponge Lamellodysidea herbacea (Dictyoceratida: Dysideidae) was found in shallow-water pools together with another, as-yet-unidentified, species of Lamellodysidea.
Sponge-associated prokaryotic communities
Prokaryotic communities were retrieved from 68 samples belonging to 23 groups, namely 22 sponge species and sediment (Supplementary data 1). After quality control, the dataset consisted of 727,262 sequences and 6636 OTUs. In terms of sequences the most abundant phyla were Chloroflexi (198,432 sequences, 1209 OTUs accounting for 18.2% of all OTUs), Proteobacteria (172,986 sequences, 1878 OTUs accounting for 28.3% of all OTUs), Actinobacteriota (81,928 sequences, 484 OTUs accounting for 7.3% of all OTUs), and Cyanobacteria (63,630 sequences, 166 OTUs accounting for 2.5% of all OTUs). The 50 most abundant OTUs are presented in Supplementary data 3. Biodiversity indices exhibited significant variations among groups (Fig. 3 and Supplementary data 4). Evenness was highest in sediment followed by species of tetractinellid sponges, all of which had high evenness. Stylissa carteri, A. willeyana and the phototrophic sponges L. herbacea and Lamellodysidea sp. had significantly lower evenness than all other sponge species (Supplementary data 4). As with evenness, richness was highest in the tetractinellid sponge species. Richness was significantly higher in the tetractinellid Aciculites ciliata than in all non-tetractinellid species (Supplementary data 4).
Diversity values and results of glm analyses for selected indices. a Evenness: F17,43 = 35.9, P < 0.001; b Richness: F17,43 = 10.49, P < 0.001; c Shannon’s H': F17,43 = 25.54, P < 0.001; d Fisher’s alpha: F17,43 = 11.55, P < 0.001. The groups sampled were as follows: Ac, Acanthostylotella cornuta; Ai, Astrosclera willeyana; Aw, Acanthochaetetes wellsi; Lh, Lamellodysidea herbacea; Ls, Lamellodysidea sp.; Pc, Petrosia corticata; Xs, Xestospongia sp.; Xt, Xestospongia testudinaria; Pa, Polymastia sp.; Sc, Stylissa carteri; Sv, Svenzea aff. devoogdae; Vi, Vetulina incrustans; Aa, Aciculites ciliata; Ax, Asteropus cf. simplex; Ds, Discodermia sp.; Ma, Manihinea sp.; Ns, Neophrissospongia sp.; Pn, Penares sp.; Sl, Scleritoderma sp.; So, Sollasipelta ornuta; Tt, Theonella aff. timmi; Ta, Theonella sp.; and Sd, sediment. Bonferroni corrected P values < 0.0125 indicate significant variation among groups
Proteobacterial abundance was significantly higher in S. carteri and A. wellsi than in all other sponge species except for the highly variable L. herbacea (Fig. 4). Aciculites ciliata was the only tetractinellid cave species in which cyanobacterial sequences were recorded. Cyanobacterial abundance was, furthermore, significantly higher in L. herbacea and Lamellodysidea sp. than in all other sponge species. The relative abundances of Chloroflexi and Actinobacteriota were significantly lower in L. herbacea, Lamellodysidea sp., and S. carteri than all other sponge species (Supplementary data 5). No Acidobacteriota members were recorded in S. carteri. In addition to this, the relative abundance of this phylum was significantly lower in Lamellodysidea herbacea, Lamellodysidea sp., Acanthochaetetes wellsi, and Vetulina incrustans than in the remaining sponge species. No Gemmatimonadota members were recorded in S. carteri or Lamellodysidea sp. The relative abundance of Gemmatimonadota was also significantly lower in A. wellsi, V. incrustans, and Xestospongia sp. than all other sponge species.
Relative abundances and results of glm analyses for the most abundant phyla: a Chloroflexi: F17,43 = 34.71, P < 0.001; b Proteobacteria: F17,43 = 22.18, P < 0.001; c Actinobacteriota: F17,43 = 20.47, P < 0.001; d Cyanobacteria: F17,43 = 65.55, P < 0.001; e Acidobacteriota: F17,43 = 31.92, P < 0.001, and f Gemmatimonadota: F17,43 = 19.77, P < 0.001. The groups sampled were as follows: Ac, Acanthostylotella cornuta; Ai, Astrosclera willeyana; Aw, Acanthochaetetes wellsi; Lh, Lamellodysidea herbacea; Ls, Lamellodysidea sp.; Pc, Petrosia corticata; Xs, Xestospongia sp.; Xt, Xestospongia testudinaria; Pa, Polymastia sp.; Sc, Stylissa carteri; Sv, Svenzea aff. devoogdae; Vi, Vetulina incrustans; Aa, Aciculites ciliata; Ax, Asteropus cf. simplex; Ds, Discodermia sp.; Ma, Manihinea sp.; Ns, Neophrissospongia sp.; Pn, Penares sp.; Sl, Scleritoderma sp.; So, Sollasipelta ornuta; Tt, Theonella aff. timmi; Ta, Theonella sp.; and Sd: sediment
The most abundant class, the Dehalococcoidia (Chloroflexota), was highly variable among sponge species ranging from no recorded sequences in L. herbacea and Lamellodysidea sp. to its greatest abundance in the sponge A. wellsi, which was significantly greater than in all other sponge species where the taxon was recorded (Fig. 5 and Supplementary data 6). The class Alphaproteobacteria was most abundant in the sponges A. wellsi and L. herbacea. For A. wellsi, the difference was significant for all sponge species except L. herbacea. The class Gammaproteobacteria was significantly more abundant in S. carteri than in all remaining sponge species. Results for the Acidimicrobiia and Cyanobacteria were basically the same as those at phylum level considering their overwhelming dominance. The Anaerolineae differed markedly from the Dehalococcoidia with Anaerolineae members reaching greatest abundance in X. testudinaria, significantly greater than in all other sponge species where the taxon was recorded. No members of the Anaerolineae were recorded in the species A. wellsi, L. herbacea, Lamellodysidea sp., or S. carteri.
Relative abundances and results of glm analyses for the most abundant classes: a Dehalococcoidia: F17,43 = 23.39, P < 0.001; b Alphaproteobacteria: F17,43 = 24.11, P < 0.001; c Gammaproteobacteria: F17,43 = 16.82, P < 0.001; d Acidimicrobiia: F17,43 = 14.91, P < 0.001; e Cyanobacteria: F17,43 = 65.55, P < 0.001; and f Anaerolineae: F17,43 = 32.79, P < 0.001. The groups sampled were as follows: Ac, Acanthostylotella cornuta; Ai, Astrosclera willeyana; Aw, Acanthochaetetes wellsi; Lh, Lamellodysidea herbacea; Ls, Lamellodysidea sp.; Pc, Petrosia corticata; Xs, Xestospongia sp.; Xt, Xestospongia testudinaria; Pa, Polymastia sp.; Sc, Stylissa carteri; Sv, Svenzea aff. devoogdae; Vi, Vetulina incrustans; Aa, Aciculites ciliata; Ax, Asteropus cf. simplex; Ds, Discodermia sp.; Ma, Manihinea sp.; Ns, Neophrissospongia sp.; Pn, Penares sp.; Sl, Scleritoderma sp.; So, Sollasipelta ornuta; Tt, Theonella aff. timmi; Ta, Theonella sp.; and Sd, sediment
PCO ordinations of the first and second axes including all samples are shown in Fig. 6. The first axis separated tetractinellid species belonging to the Ancorinidae, Corallistidae, Geodiidae, and Theonellidae families from all other sponge species, while the second axis separated tetractinellid species belonging to the Sclerotidermidae family from all other species. There was a highly significant difference in composition among groups (sponge species and sediment) (adonis, F17,43 = 7.87, P < 0.001, R2 = 0.757). Species identity/group, thus explained more than 75% of the variation in composition. The eigenvalues for the first and second axes of the full dataset were 4.48 and 2.36, respectively, and accounted for 15.03% and 7.90% of the variation in the data set or 22.93% of the total variation.
Ordination showing the first two axes of the principal coordinates analysis (PCO) of OTU composition. Symbols are colour coded and represent samples belonging to different groups as shown in the legend on the right side of the figure. Grey symbols denote weighted averages scores for OTUs. The symbol size corresponds to group abundance (number of sequence reads). The legend symbols represent the following groups: Ac, Acanthostylotella cornuta; Ai, Astrosclera willeyana; Aw, Acanthochaetetes wellsi; Lh, Lamellodysidea herbacea; Ls, Lamellodysidea sp.; Pc, Petrosia corticata; Xs, Xestospongia sp.; Xt, Xestospongia testudinaria; Pa, Polymastia sp.; Sc, Stylissa carteri; Sv, Svenzea aff. devoogdae; Vi, Vetulina incrustans; Aa, Aciculites ciliata; Ax, Asteropus cf. simplex; Ds, Discodermia sp.; Ma, Manihinea sp.; Ns, Neophrissospongia sp.; Pn, Penares sp.; Sl, Scleritoderma sp.; So, Sollasipelta ornuta; Tt, Theonella aff. timmi; Ta, Theonella sp.; and Sd, sediment
Excluding the tetractinellid species, the major axis of variation in the PCO analysis separated the known low microbial abundance (LMA) sponge species S. carteri from the known HMA sponge species X. testudinaria with the other sponge species intermediate (Fig. 7). There, however, appeared to be two clusters of samples of these intermediate species with one, closer to X. testudinaria, consisting of the species P. corticata, Polymastia sp., A. willeyana, Xestospongia sp., and Svenzea aff. devoogdae and the other, closer to S. carteri, consisting of the species A. cornuta, V. incrustans, L. herbacea, and Lamellodysidea sp. in addition to sediment. The second axis of variation separated the sclerosponge A. wellsi from all other sponge species.
Ordination showing the first and second axes of the principal coordinates analysis (PCO) of OTU composition excluding tetractinellid host species. Symbols are colour coded and represent samples belonging to different groups as shown in the legend on the right side of the figure. Grey symbols denote weighted averages scores for OTUs. The symbol size corresponds to group abundance (number of sequence reads). The legend symbols represent the following groups: Ac, Acanthostylotella cornuta; Ai, Astrosclera willeyana; Aw, Acanthochaetetes wellsi; Lh, Lamellodysidea herbacea; Ls, Lamellodysidea sp.; Pc, Petrosia corticata; Xs, Xestospongia sp.; Xt, Xestospongia testudinaria; Pa, Polymastia sp.; Sc, Stylissa carteri; Sv, Svenzea aff. devoogdae; Vi, Vetulina incrustans; and Sd, sediment
Tetractinellid species shared a similar subset of abundant OTUs compared to the other host sponge species (Fig. 8). In addition to this, OTUs 41, 42, and 50 were abundant or restricted to the tetractinellid species Aciculites ciliata and Scleritoderma sp., which clustered together in the PCO analysis. OTU-84, assigned to the Caldilineales, in turn, was recorded in all tetractinellid species with the exception of the aforementioned species.
Percentage abundance of the 50 most abundant OTUs represented by different colours based on prokaryotic phylum for the following groups: Ac, Acanthostylotella cornuta; Ai, Astrosclera willeyana; Aw, Acanthochaetetes wellsi; Lh, Lamellodysidea herbacea; Ls, Lamellodysidea sp.; Pc, Petrosia corticata; Xs, Xestospongia sp.; Xt, Xestospongia testudinaria; Pa, Polymastia sp.; Sc, Stylissa carteri; Sv, Svenzea aff. devoogdae; Vi, Vetulina incrustans; Aa, Aciculites ciliata; Ax, Asteropus cf. simplex; Ds, Discodermia sp.; Ma, Manihinea sp.; Ns, Neophrissospongia sp.; Pn, Penares sp.; Sl, Scleritoderma sp.; So, Sollasipelta ornuta; Tt, Theonella aff. timmi; Ta, Theonella sp.; and Sd, sediment. The size of each OTU circle is proportional to the mean percentage of sequences per group as shown by the symbol legend in the upper right corner of the figure
Several abundant OTUs were only recorded from single sponge species. OTU-91 was restricted to the sponge V. incrustans, OTU-97 to the sponge P. corticata, OTU-21 to A. cornuta, and OTU-104 to A. willeyana. Apart from these sponges, some sponge species harboured multiple OTUs that were unique to those species, ranging from two OTUs (5 and 88) for Lamellodysidea sp., to three OTUs for S. carteri (10, 15, and 24), four OTUs for L. herbacea (4, 12, 62, and 68), and five OTUs (2, 8, 37, 38, and 64) for A. wellsi. OTUs 4, 5, and 12, all classified as the cyanobacterial species Hormoscilla spongeliae, all exhibited close relationships to organisms obtained from sponges identified as Dysideidae sp. from Guam. OTU-88 only had 91% sequence similarity to an organism detected in a sponge identified as Lamellodysidea sp. sampled from China. OTU-62 had > 99% sequence similarity to an organism detected in a sponge identified as Phyllospongia papyracea. OTU-68 had 96% sequence similarity to an organism detected in a sponge identified as Lendenfeldia chondrodes (Supplementary data 3).
The five OTUs (2, 8, 37, 38, and 64) restricted to the sponge A. wellsi were classified as belonging to the SAR202 clade (Chloroflexi), Defluviicoccales (Alphaproteobacteria), HOC36 (Gammaproteobacteria), Actinobacteriota, and Gammaproteobacteria, respectively. OTU-2 had 95% sequence similarity to an organism detected in a sponge identified as Coelocarteria singaporensis sampled from the Great Barrier Reef. OTU-8 had 96% sequence similarity to an organism detected in a coral identified as Pocillopora damicornis. OTU-37 had 94% sequence similarity to an organism detected in a coral identified as Siderastrea stellata sampled from Brazil. OTU-38 had 94% sequence similarity to an organism detected in a sponge identified as Poecillastra compressa. OTU-64 had 96% sequence similarity to an organism detected in biofilm of an artificial substrate from the Great Barrier Reef (Supplementary Table 3).
Discussion
Here, we assessed the prokaryotic communities of several sponge species sampled from waters surrounding Orchid Island, Taiwan. In addition to sampling sponges from a karst pool and cave ecosystem, two species of sponges (X. testudinaria and S. carteri) were also sampled from nearby coral reefs. Both X. testudinaria and S. carteri have been extensively studied in previous publications as pertains to their microbial symbionts (Cleary et al. 2015, 2018, 2020, 2021, 2022; de Voogd et al. 2015, 2019; Polónia et al. 2015, 2016, 2017, 2018, 2021). Below, we go into more detail with respect to specific groups.
Sclerosponges
Two sclerosponge species, namely, A. willeyana (order Agelasida) and A. wellsi (order Clionaida) were sampled from cave and canyon habitats. Sclerosponges form a hypercalcified, coral-like skeleton surrounded by a very thin layer of live tissue embedded with siliceous spicules and exhibit superficial similarities, despite belonging to distantly related orders. Extant sclerosponges, relicts of a bygone age, bear morphological resemblance to now extinct stromatoporids that once formed extensive reefs before and alongside the emergence of coral reefs (Basile et al. 1984). They prevailed as the main reef-building marine organisms throughout the Phanerozoic, but can still be found in modern marine environments (Vacelet 1985; Asami et al. 2021 and references therein). They are long-lived, slow growing, often mushroom-shaped, and primarily inhabit caves and deeper waters; in some locations, they contribute to reef formation (Asami et al. 2020; Lang et al. 1975; Macartney et al. 2020). These characteristics, together with the fact that their exoskeleton is formed by the slow deposition of calcium carbonate in sequential layers through time, render them valuable as paleo-proxy recorders of environmental conditions at depths and environments where photosynthetic scleractinian corals are not present (Asami et al. 2020; Grottoli et al. 2020).
Acanthochaetetes wellsi is a member of the Acanthochaetetidae family, an ancient family with a predominantly extinct lineage (Rützler and Vacelet 2002). Only a handful of species persist within two extant genera, including, A. wellsi and Willardia caicosensis. The genus Acanthochaetetes was established based on the extinct fossil species A. seunesi Fischer, 1970; A. wellsi is considered a living fossil. It was originally discovered in shallow-water caves of Guam but has since been recorded in other Indo-Pacific locations in similar, cave habitats. In addition to A. wellsi, another agelasid, the non-sclerosponge Acanthostylotella cornuta (Topsent, 1897), was also observed. This mono-specific, poorly-known species (de Voogd et al. 2010) was originally described from Indonesia and was recently placed in the Agelasida (Morrow and Cárdenas 2015).
Our results demonstrated that, despite their hypercalcified skeletal similarity, the sclerosponge species harboured distinct prokaryotic communities. Species richness and evenness were higher in A. willeyana than in A. wellsi. Chloroflexi were, however, relatively abundant in both host species. Acidobacteriota, in turn, were more abundant in A. willeyana than in A. wellsi. In terms of composition, A. willeyana clustered closer to the known HMA sponge species X. testudinaria. Our results for A. willeyana align with those of Karlińska-Batres and Wörheide (2013a, 2015) for A. willeyana specimens sampled from the Great Barrier Reef and from sites across the Indo-Pacific region, respectively, namely, prokaryotic communities dominated by Chloroflexi with major Actinobacteriota, Acidobacteriota, and Proteobacteria components and minor Gemmatimonadota and Poribacteria components. In a previous study, the coralline Ceratoporella nicholsoni (order Agelasida) maintained a stable microbiome through a depth gradient (37 to 97 m) consisting mainly of Chloroflexi, Thaumarcheaota, Proteobacteria, Acidobacteria, and Actinobacteria (Macartney et al. 2020). Similar results were obtained for the coralline sponge Vaceletia crypta (order Dictyoceratida) with Chloroflexi the most abundant phylum (35%) (Karlińska-Batres and Wörheide 2013b). Some studies have suggested that the symbiotic microbial communities of coralline members may be involved in calcification (Jackson et al. 2011; Jackson and Wörheide 2014). Karlińska-Batres and Wörheide (2013a) also noted that their samples separated according to sampling location suggesting an important role of geographic distance in structuring prokaryotic community composition in line with our study for two HMA sponge species, X. testudinaria and Hyrtios erectus (Cleary et al. 2022).
Acanthochaetetes wellsi harboured the most compositionally distinct prokaryotic community of the two coralline sponges sampled, had the lowest richness of all sponge species, and harboured the most taxon-restricted abundant OTUs. These OTUs, furthermore, had relatively low sequence similarities to organisms in GenBank. Moreover, the Chloroflexi in A. wellsi were exclusively classified as Dehalococcoidia with no OTUs classified as Anaerolineae. To the best of our knowledge, this is the first next-generation assessment of the prokaryotic community of the sclerosponge Acanthochaetetes wellsi.
Lithistid sponges
In addition to sclerosponges, we encountered a diverse array of other sponge species, among which lithistids were particularly notable. Lithistids, also known as desma-bearing or rock sponges, are characterised by their exceptionally dense (hypersilicified) skeletal structure and articulated skeletons composed of desma megascleres (Pisera and Gerovasileiou 2021). Lithistids, an ancient lineage of sponges, were integral components of Upper Cambrian reefs, and alongside microbial biofilms, they contributed to the formation of columnar structures known as stromatolites (Coulson and Brand 2016). Valeria D’Auria et al. (2002) identified them as a ‘spectacular source of new metabolites’. A diverse array of compounds, including alkaloids, pigments, novel sterols, cyclic and linear peptides, polyketides, and macrolides, have been isolated from these sponges. Drawing parallels between the structures of many of these compounds and those produced by microorganisms, Valeria D’Auria et al. (2002) suggested that some of these compounds may be synthesised by microbial symbionts residing within the sponges.
Lithistids can form dense aggregations in shallow-water caves where freshwater influx occurs (Pisera and Gerovasileiou 2021) and at greater depths on sponge mounds where they provide habitat to a diverse array of vertebrate and invertebrate species and contribute to nutrient cycling and bentho-pelagic exchange (Cathalot et al. 2015; Hawkes et al. 2019; Hourigan et al. 2017; Maldonado et al. 2015, 2020; Manconi et al. 2006; Rooks et al. 2020; Xavier et al. 2021). They were previously classified in their own order, the Lithistida. The reclassification of lithistids into different orders was driven by molecular and morphological data that suggested polyphyletic origins (Morrow and Cárdenas 2015). The tetracladinid and dicranocladinid lithistids, however, remained monophyletic and branched with the choristid demosponges. Based on molecular systematics, most lithistid sponges have been placed in the Tetractinellida order, which presently includes three suborders, the Astrophorina, Spirophorina, and Thoosina (Schuster et al. 2021). Tetractinellid sponges have a subradial or radial skeletal configuration, both monactine and triaene megascleres, aster, sigma, microxea, raphide and microrhabd microscleres, and desmas are sometimes present (Morrow and Cárdenas 2015). In contrast to the tetracladinid lithistids, rhizomorinid families were identified as polyphyletic (Kelly-Borges and Pomponi 1994). The only non-tetracladinid lithistid sampled in the present study was the bright yellow Vetulina incrustans (Vetulinidae: Sphaerocladina), recently described from deep crevices of caves in the Philippines (Schuster et al. 2018). A single specimen of S. ornata was also sampled, the only representative of the monogeneric dicranocladinid family Neopeltidae. Originally found at a depth of 236 m in the Banda Sea, Indonesia (Sollas 1888), it was the only Sollasipelta species inhabiting both deep-sea and shallow cave/dark habitats. Recently, a new species of Sollasipelta was described from a submarine cave in the Ryukyu Islands in Japan (Ise et al. 2023). The authors compared their newly observed species with three other extant species, and although the location of the new species S. subterranea is very close to Orchid Island, our species aligns with the description of S. ornata and not S. subterranea.
Lithistid species belonging to the Tetractinellida order formed two distinct clusters in the PCO analysis with one cluster encompassing both species from the Scleritodermidae family (Aciculites ciliata and Scleritoderma sp.) while the other cluster encompassed all remaining species from the other families. The Scleritodermidae is the only tetractinellid family recorded in the present study belonging to the suborder Spirophorina. All the other tetractinellid families (Ancorinidae, Corallistidae, Geodiidae, Neopeltidae, and Theonellidae) belonged to the suborder Astrophorina. This suggests a phylogenetic signal on sponge-associated prokaryotic composition. Lithistid species belonging to the Tetractinellida order generally exhibited high evenness and richness and relatively high abundances of HMA-indicator taxa, such as Chloroflexi, Actinobacteriota, and Acidobacteriota (Moitinho-Silva et al. 2017). The microbial community of the tetractinellid cold water Geodia barretti (Tetractinellida, Demospongiae) was shown to be dominated by Chloroflexi (SAR202), Poribacteria, and Acidobacteria (Radax et al. 2012). Proteobacteria, particularly Gammaproteobacteria, constituted the most abundant taxon, followed by Chloroflexi, Firmicutes, and Alphaproteobacteria in the lithistid sponge Discodermia spp. sampled across a depth range of 24 to 161 meters in the Bahamas Archipelago (Brück et al. 2012).
The tetractinellid species shared a large number of dominant OTUs, many of which were not recorded in other sponge taxa in the present study with the exception of Polymastia sp. (Order: Polymastiida), which clustered intermediate to the tetractinellid species and the other sponge species. Unfortunately, our sampling for this species was limited to a single specimen. Despite their markedly distinct composition, the dominant OTUs recorded in the tetractinellids had relatively high sequence similarities to organisms in GenBank, in contrast to the sclerosponge species A. wellsi. Samples of the non-tetractinellid, lithistid V. incrustans clustered distinct from the tetractinellid, lithistids and only shared a single abundant OTU with them.
Petrosiidae
Petrosiid sponges exhibit a wide geographical distribution, thrive in a range of depths, and tolerate a wide range of water temperatures, from cold to warm (Desqueyroux-Faúndez and Valentine 2002). They typically form massive, bulbous structures, but can also exhibit branching or encrusting growth forms, possess a stony, brittle texture and a smooth surface, and may assume a vase-like shape reminiscent of a volcano crater. The family encompasses four genera and two subgenera. The Petrosia genus is characterized by a network of free spicules composed of oxea, exhibiting three distinct size classes within the subgenus Petrosia and strongyle megascleres within the subgenus Strongylophora. Their distribution spans the North, Central, and South Pacific, Central Atlantic, and Indian Oceans (Desqueyroux-Faúndez and Valentine 2002).
Three sampled host sponge species belonged to the family Petrosiidae including the known HMA species X. testudinaria as well as Xestospongia sp. and P. corticata. Previous studies have identified several species within the Petrosiidae family as HMA species (Gloeckner et al. 2014; Moitinho-Silva et al. 2017). For example, X. testudinaria has consistently been shown to harbour a HMA-type prokaryotic community characterised by a predominance of HMA-indicator taxa (Cleary et al. 2018, 2019a, b, 2020). Likewise, in a compositional analysis, the sponge species Petrosia elephantotus collected from the Red Sea clustered together with other known HMA species such as X. testudinaria, and had a relatively high abundance of Poribacteria compared to non-HMA species sampled from the same area (Cleary et al. 2020). In Mayotte, a species tentatively identified as Petrosia aff. spheroida clustered together with the known HMA species H. erectus and X. testudinaria (de Voogd et al. 2019). The Mediterranean Petrosia ficiformis was also recognised as a HMA species (Ribes et al. 2015). Interestingly, P. ficiformis occurs in two distinct colour morphs. The white/pink morph was found in dark/shady environments, including cave interiors and entrances, and lacked phototrophic symbionts. A violet morph, in contrast, inhabiting illuminated habitats such as rocky cliffs, harboured a diverse assemblage of intracellular cyanobacteria. The presence of the cyanobacterium Synechococcus feldmannii has, furthermore, been confirmed (with the aid of TEM observation and Chlorophyll a measurements) in the pink and violet morphs, but not in the white morph (Burgsdorf et al. 2014). A 16S rRNA pyrosequencing analysis of these two different colour morphs also revealed a stable bacterial community dominated by Chloroflexi, Gammaproteobacteria, and Acidobacteria in all colour morphs. Geographical variation was also shown to be a more important structural component of variation in microbial diversity than host-genetic variability (Burgsdorf et al. 2014).
Phototrophic sponges
Phototrophic sponges were another notable component of the sponge fauna in the present study. As the name implies, they depend on photosynthetic symbionts for a substantial portion of their energy needs. Previous research has documented varying distributions of photo- and heterotrophic sponges with phototrophic sponges more abundant in the Indo-Pacific region (e.g., the Great Barrier Reef) and primarily inhabiting outer reefs, located further from the shoreline, while heterotrophic sponges are relatively more prevalent in the Caribbean (Bell et al. 2018, 2020; Erwin and Thacker 2007). Phototrophic sponges also predominantly occur in clearer, oligotrophic environments.
Cyanobacteria are among the most abundant photosynthetic organisms present in symbiotic associations with sponges in general and particularly in phototrophic sponges (Konstantinou et al. 2018). Lamellodysidea species, for example, have been consistently shown to host the cyanobacterial symbiont, Hormoscilla spongeliae (previously known as Oscillatoria spongeliae), and appear unable to survive without the species. This obligate relationship has also made it hitherto impossible to culture H. spongeliae owing to the symbiont's apparent inability to survive independently (Schorn et al. 2019; Usher 2008). Hormoscilla spongeliae is known to produce high amounts of several halogenated compounds and polybrominated diphenyl ethers (PBDEs), which are chemically identical to anthropogenic pollutants, but with potential antimicrobial and antipredator properties (Agarwal et al. 2017; Flatt et al. 2005). Alongside Cyanobacteria, the microbial community of Lamellodysidea members and more specifically L. herbacea also consisted of Bacteroidetes, Alphaproteobacteria, Gammaproteobacteria, and Oligoflexia members (Podell et al. 2020). Both phototrophic sponges in this study exhibited low evenness and richness. Unlike many of the other sponge species, both phototrophic sponge species were only found in the high-light intensity pool environments growing over rock surfaces. In addition to L. herbacea, H. spongeliae, recorded as Oscillatoria spongeliae, has also been found to be a prominent member of the microbiomes of the phototrophic sponges Phyllospongia papyracea and Lendenfeldia chondrodes (Ridley et al. 2005).
SAR202 were enriched in cave sponges
The bacterial phylum Chloroflexi exhibited the highest overall abundance and Chloroflexi members were only rare in species from the high-light intensity pools or adjacent coral reefs, namely, the phototrophic species or S. carteri, a known LMA sponge species. Several studies have documented a high abundance of Chloroflexi members in HMA species (Cleary et al. 2020, 2021; Cleary et al2019a; Moitinho-Silva et al. 2017; Schmitt et al. 2011; Swierts et al. 2018). Despite the scarcity or absence of Chloroflexi members in numerous LMA species, they can flourish in others. For example, we previously observed Chloroflexi members in the sponge species Paratetilla bacca from Mayotte, Acanthella cavernosa from Taiwan, and Ectyoplasia coccinea, Cinachyrella sp., and Topsentia aqabaensis from the Red Sea, all of which were otherwise compositionally similar to samples from known LMA species from the same locations (Cleary et al. 2019a; Cleary et al. 2020; de Voogd et al. 2019). Chloroflexi abundance was high in both the agelasid A. cornuta and the sclerosponge A. wellsi. In A. wellsi, this was mainly due to high levels of Dehalococcoidia and in A. cornuta high levels of Dehalococcoidia (mainly SAR202) and TK17. In the Silva 138 database, the SAR202 clade is an order of the Dehalococcoidia. In both sponge species, levels of Anaerolineae were very low in contrast to the known HMA species X. testudinaria sampled from nearby coral reefs, which had the highest Anaerolineae levels. SAR202 members have been shown to be abundant in bathypelagic waters and Anaerolinae members in a wide range of habitats varying from arctic permafrost to the mammalian gastrointestinal tract (Campbell et al. 2014; Hug et al. 2013; Varela et al. 2008). In humans, Anaerolinae members scavenge material from human tissue and lysed bacterial cells and may perform a similar function in sponges given their rapid cell turnover.
Prior research indicates that HMA sponges are specialised for the uptake of dissolved organic matter (DOM), whereas LMA sponge species are specialised for the uptake of particulate organic matter (Bayer et al. 2018; McMurray et al. 2018). Chloroflexi, in particular, are postulated to play a key role in DOM dynamics. SAR202, for example, have been implicated in the degradation of recalcitrant or refractory DOM (Colatriano et al. 2018; Landry et al. 2017). Landry et al. (2017) identified enzymes involved in recalcitrant compound oxidation within the genomes of deep-sea Chloroflexi members, some of which were also detected in SAR202 genomes from sponge symbionts (Bayer et al. 2018). The aforementioned evidence suggests a potential role for SAR202 members in DOM degradation within sponges, particularly in environments with elevated DOM levels, such as areas close to rivers or submarine groundwater discharge (Kim and Kim 2017). The overwhelming abundance of Chloroflexi in cave-dwelling sponges including tetractinellids, other putative HMA species, and species such as A. cornuta and A. wellsi hints that DOM-rich groundwater discharge from Orchid Island may constitute the primary environmental factor shaping the cave sponge communities. While further research is needed to substantiate this hypothesis, the implications are intriguing and suggest that the sponge-filled cave systems surrounding Orchid Island and other similar areas could potentially function as DOM filters with important repercussions for nutrient dynamics across the land-sea interface.
In summary, this study delves into a poorly studied ecosystem with a number of conspicuous sponge species potentially new to science. In addition to the species in the present study, a rich calcareous sponge fauna was also encountered. While these specimens have been collected, their species-level identification remains pending. Formal descriptions of the potentially new species are forthcoming. In addition to sponges, other groups of organisms including corals and algae were observed. Surveys should also be undertaken to discover potentially similar systems in adjacent insular areas with rocky coasts. Intriguing and otherwise rare species have already been identified in cave systems in Japan and the Philippines (Schuster et al. 2018; Asami et al. 2020, 2021). Aside from the present karst cave system, these other cave systems potentially harbour a wealth of new and rare species. For instance, we collected three different specimens of the bright, yellow coloured, encrusting V. incrustans. This species was recently described based on a single specimen from a crevice in the Philippines (Schuster et al. 2018). Species belonging to this genus are considered relict species, representing the sole surviving members of a once diverse Mesozoic group. Previously, they were only recorded from the Caribbean region (Pisera et al., 2017). Ise et al. (2023) recently described a new lithistid species, Sollasipelta subterranea from an anchialine cave in the Ruykyus Islands, and an earlier report showed that many more species await description including new species from the particular cave in Japan (Ise 2019). In the Mediterranean, caves support a large part of total poriferan diversity in addition to the large number of deep-sea, relict, and living fossils that are recorded in these caves (Gerovasileiou and Voultsiadou 2012); marine cave systems, however, remain understudied in other regions of the world. With regard to the prokaryotic communities, we we observed a marked enrichment of Dehalococcoidia members (primarily SAR202) in all cave sponge species.
References
Agarwal V, Blanton JM, Podell S, Taton A, Schorn MA, Busch J, Lin Z, Schmidt EW, Jensen PR, Paul VJ, Biggs JS, Golden JW, Allen EE, Moore BS (2017) Metagenomic discovery of polybrominated diphenyl ether biosynthesis by marine sponges. Nat Chem Biol 13(5):537–543. https://doi.org/10.1038/nchembio.2330
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Asami R, Kinjo A, Ohshiro D, Naruse T, Mizuyama M, Uemura R, Shinjo R, Ise Y, Fujita Y, Sakamaki T (2020) Evaluation of geochemical records as a paleoenvironmental proxy in the hypercalcified demosponge Astrosclera willeyana. Prog Earth Planet Sci 7:1–18
Asami R, Matsumori T, Shinjo R, Uemura R, Miyaoka Y, Mizuyama M, Ise Y, Sakamaki T (2021) Reconstruction of ocean environment time series since the late nineteenth century using sclerosponge geochemistry in the northwestern subtropical Pacific. Prog Earth Planet Sci 8:38. https://doi.org/10.1186/s40645-021-00434-7
Bakran-Petricioli T, Vacelet J, Zibrowius H, Petricioli D, Chevaldonné P, Rađa T (2007) New data on the distribution of the ‘deep-sea’ sponges Asbestopluma hypogea and Oopsacas minuta in the Mediterranean Sea. Mar Ecol 28:10–23. https://doi.org/10.1111/j.1439-0485.2007.00179.x
Basile LL, Cuffey RJ, Kosich DF (1984) Sclerosponges, pharetronids, and sphinctozoans (relict cryptic hard-bodied Porifera) in the modern reefs of Enewetak Atoll. J Paleontol 58:636–650
Bayer K, Jahn MT, Slaby BM, Moitinho-Silva L, Hentschel U (2018) Marine sponges as Chloroflexi hot spots: genomic insights and high-resolution visualization of an abundant and diverse symbiotic clade. mSystems 3(6):e00150-18. https://doi.org/10.1128/mSystems.00150-18
Bell JJ, Rovellini A, Davy SK, Taylor MW, Fulton EA, Dunn MR, Bennett HM, Kandler NM, Luter HM, Webster NS (2018) Climate change alterations to ecosystem dominance: how might sponge-dominated reefs function? Ecology 99:1920–1931. https://doi.org/10.1002/ecy.2446
Bell JJ, McGrath E, Kandler NM, Marlow J, Beepat SS, Bachtiar R, Shaffer MR, Mortimer C, Micaroni V, Mobilia V, Rovellini A, Harris B, Farnham E, Strano F, Carballo JL (2020) Interocean patterns in shallow water sponge assemblage structure and function. Biol Rev Camb Philos Soc 95:1720–1758. https://doi.org/10.1111/brv.12637
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodríguez AM, Chase J, Cope EK, Da Silva R, Diener C, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibbons SM, Gibson DL, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley GA, Janssen S, Jarmusch AK, Jiang L, Kaehler BD, Kang KB, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu YX, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton JT, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson MS 2nd, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, van der Hooft JJJ, Vargas F, Vázquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang Y, Zhu Q, Knight R, Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Brück WM, Reed JK, McCarthy PJ (2012) The bacterial community of the lithistid sponge Discodermia spp. as determined by cultivation and culture-independent methods. Mar Biotechnol 14:762–773
Burgsdorf I, Erwin PM, López-Legentil S, Cerrano C, Haber M, Frenk S, Steindler L (2014) Biogeography rather than association with cyanobacteria structures symbiotic microbial communities in the marine sponge Petrosia ficiformis. Front Microbiol 5:529. https://doi.org/10.3389/fmicb.2014.00529
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583
Campbell AG, Schwientek P, Vishnivetskaya T, Woyke T, Levy S, Beall CJ, Griffen A, Leys E, Podar M (2014) Diversity and genomic insights into the uncultured Chloroflexi from the human microbiota. Environ Microbiol 16:2635–2643. https://doi.org/10.1111/1462-2920.12461
Cárdenas CA, González-Aravena M, Font A, Hestetun JT, Hajdu E, Trefault N, Malmberg M, Bongcam-Rudloff E (2018) High similarity in the microbiota of cold-water sponges of the Genus Mycale from two different geographical areas. PeerJ 6:e4935. https://doi.org/10.7717/peerj.4935
Cathalot C, Van Oevelen D, Cox TJS, Kutti T, Lavaleye M, Duineveld G, Meysman FJR (2015) Cold-water coral reefs and adjacent sponge grounds: hotspots of benthic respiration and organic carbon cycling in the deep sea. Front Mar Sci 2:37. https://doi.org/10.3389/fmars.2015.00037
Chao S-M (2002) The shallow-water echinoderms from Lanyu, Taiwan. Collect Res 15:1–7
Chao S-M, Lee S-C (2002) Marine shells from Lanyu, Taiwan. Collect Res 15:9–33
Chen CH, Shieh YN, Lee T, Chen C-H, Mertzman SA (1990) Nd-Sr-O isotopic evidence for source contamination and an unusual mantle component under Luzon Arc. Geochim Cosmochim Acta 54:2473–2483
Chen CH, Yang TF, Tien JL, Lee T (1992) Eruption ages of North Luzon Arc (Taiwan): based on fission-track dating. Acta Geol Taiwan 30:149–156
Chen CA, Wei NV, Dai CF (2022a) Genotyping the clonal population structure of a gorgonian coral, Junceella fragilis (Anthozoa: Octocorallia: Ellisellidae) from Lanyu, Taiwan, using simple sequence repeats in ribosomal intergenic spacer. Zool Stud 41:295–302
Chen YH, Chen HJ, Yang CY, Shiu JH, Hoh DZ, Chiang PW, Chow WS, Chen CA, Shih TH, Lin SH, Yang CM, Reimer JD, Hirose E, Iskandar BH, Huang H, Schupp PJ, Tan CHJ, Yamashiro H, Liao MH, Tang SL (2022b) Prevalence, complete genome, and metabolic potentials of a phylogenetically novel cyanobacterial symbiont in the coral-killing sponge, Terpios Hoshinota. Environ Microbiol 24(3):1308–1325. https://doi.org/10.1111/1462-2920.15824
Chen Y-G (1993) Neotectonics and sea level change in southern part of Taiwan region since Late Pleistocene. Doctoral dissertation, National Taiwan University, Taipei, Taiwan, p 159
Cleary DFR, de Voogd NJ, Polónia ARM, Freitas R, Gomes NCM (2015) Composition and predictive functional analysis of bacterial communities in seawater, sediment and sponges in an Indonesian coral reef environment. Microb Ecol 70:889–903. https://doi.org/10.1007/s00248-015-0632-5
Cleary DFR, Polónia ARM, Becking LE, de Voogd NJ, Purwanto GH, Gomes NCM (2018) Compositional analysis of bacterial communities in seawater, sediment and high and low microbial abundance sponges in the Misool coral reef system, Indonesia. Mar Biodivers 48:1889–1901. https://doi.org/10.1007/s12526-017-0697-0
Cleary DFR, Polónia ARM, Huang YM, Gomes NCM, de Voogd NJ (2019a) A comparison of prokaryote communities inhabiting sponges, bacterial mats, sediment and seawater in Southeast Asian coral reefs. FEMS Microbiol Ecol 95:fiz169. https://doi.org/10.1093/femsec/fiz169
Cleary DFR, Swierts T, Coelho FJRC, Polónia ARM, Huang YM, Ferreira MRS, Putchakarn S, Carvalheiro L, van der Ent E, Ueng JP, Gomes NCM, de Voogd NJ (2019b) The sponge microbiome within the greater coral reef microbial metacommunity. Nat Commun 10:1644. https://doi.org/10.1038/s41467-019-09537-8
Cleary DFR, Polónia ARM, Reijnen BT, Berumen ML, de Voogd NJ (2020) Prokaryote communities inhabiting endemic and newly discovered sponges and octocorals from the Red Sea. Microb Ecol 80:103–119. https://doi.org/10.1007/s00248-019-01465-w
Cleary DFR, Polónia ARM, de Voogd NJ (2021) Composition and diversity of Prokaryotic communities sampled from sponges and soft corals in Maldivian waters. Mar Ecol 42:e12638. https://doi.org/10.1111/maec.12638
Cleary DFR, Polónia ARM, Swierts T, Coelho FJRC, de Voogd NJ, Gomes NCM (2022) Spatial and environmental variables structure sponge symbiont communities. Mol Ecol 31:4932–4948. https://doi.org/10.1111/mec.16631
Colatriano D, Tran PQ, Guéguen C, Williams WJ, Lovejoy C, Walsh DA (2018) Genomic evidence for the degradation of terrestrial organic matter by pelagic Arctic Ocean Chloroflexi bacteria. Commun Biol 1:90. https://doi.org/10.1038/s42003-018-0086-7
Coulson KP, Brand LR (2016) Lithistid sponge-microbial reef-building communities construct laminated, Upper Cambrian (Furongian) “stromatolites.” Palaios 31:358–370
de Voogd NJ, Cleary DFR, Polónia ARM, Gomes NCM (2015) Bacterial communities of four different biotopes and their functional genomic nitrogen signature from the thousand-island reef complex, West-Java, Indonesia. FEMS Microbiol Ecol 91:019. https://doi.org/10.1093/femsec/fiv019
de Voogd NJ, Gauvin-Bialecki A, Polónia ARM, Cleary DFR (2019) Assessing the bacterial communities of sponges inhabiting the remote western Indian Ocean island of Mayotte. Mar Ecol 39:e12517. https://doi.org/10.1111/maec.12517
de Voogd N, Erpenbeck D, Hooper JNA, van Soest RWM (2010) Acanthostylotella, the forgotten genus. Affinities of the family Agelasidae (Porifera, Demospongiae, Agelasida): chemical, molecular and morphological evidence. In: Ancient Animals New Challenges, Uriz MJ, Maldonado M, Becerro M, Turon X (eds). Book of Abstracts for the VIII World Sponge Conference, Girona, p. 172.
Desqueyroux-Faúndez R, Valentine C (2002) Family Petrosiidae Van Soest, 1980. In: Hooper JNA, Van Soest RWM, Willenz P (eds) Systema Porifera. Springer, Boston, MA. https://doi.org/10.1007/978-1-4615-0747-5_95
Erwin PM, Thacker RW (2007) Incidence and identity of photosynthetic symbionts in Caribbean coral reef sponge assemblages. J Mar Biol Assoc UK 87:1683–1692. https://doi.org/10.1017/S0025315407058213
Flatt PM, Gautschi JT, Thacker RW, Musafija-Girt M, Crews P, Gerwick WH (2005) Identification of the cellular site of polychlorinated peptide biosynthesis in the marine sponge Dysidea (Lamellodysidea) herbacea and symbiotic cyanobacterium Oscillatoria spongeliae by CARD-FISH analysis. Mar Biol 147:761–774. https://doi.org/10.1007/s00227-005-1614-9
Gerovasileiou V, Voultsiadou E (2012) Marine caves of the Mediterranean Sea: a sponge biodiversity reservoir within a biodiversity hotspot. PLoS ONE 7(7):e39873. https://doi.org/10.1371/journal.pone.0039873
Gerovasileiou V, Chintiroglou C, Vafidis D, Koutsoubas D, Sini M, Dailianis T, Issaris Y, Akritopoulou E, Dimarchopoulou D, Voultsiadou E (2015) Census of biodiversity in marine caves of the Eastern Mediterranean Sea. Mediterr Mar Sci 16:245–265. https://doi.org/10.12681/mms.1069
Gerovasileiou V, Chintiroglou CC, Konstantinou D, Voultsiadou E (2016) Sponges as “living hotels” in Mediterranean marine caves. Sci Mar 80(3):279–289. https://doi.org/10.3989/scimar.04403.14B
Gloeckner V, Wehrl M, Moitinho-Silva L, Gernert C, Schupp P, Pawlik JR, Lindquist NL, Erpenbeck D, Wörheide G, Hentschel U (2014) The HMA-LMA dichotomy revisited: an electron microscopical survey of 56 sponge species. Biol Bull 227:78–88. https://doi.org/10.1086/BBLv227n1p78
Grottoli AG, Chapron L, Gava D, Olesik JW (2020) Natural variability of skeletal elemental phosphorus (P/Ca), lead (Pb/Ca), and barium (Ba/Ca) in the western Pacific sclerosponges Acanthoceatetes wellsi and Astrosclera welleyana. Geochem Geophys 21:e2020GC009245
Harmelin JG (1997) Diversity of bryozoans in a Mediterranean sublittoral cave with bathyal-like conditions: role of dispersal processes and local factors. Mar Ecol Prog Ser 153:139–152. https://doi.org/10.3354/meps153139
Hawkes N, Korabik M, Beazley L, Rapp HT, Xavier JR, Kenchington E (2019) Glass sponge grounds on the Scotian Shelf and their associated biodiversity. Mar Ecol Prog Ser 614:91–109. https://doi.org/10.3354/meps12903
Hentschel U, Hopke J, Horn M, Friedrich AB, Wagner M, Hacker J, Moore BS (2002) Molecular evidence for a uniform microbial community in sponges from different oceans. Appl Environ Microbiol 68:4431–4440
Hill M, Hill A, Lopez N, Harriott O (2006) Sponge-specific bacterial symbionts in the Caribbean sponge, Chondrilla nucula (Demospongiae, Chondrosida). Mar Biol 148:1221–1230
Hourigan TF, Etnoyer PJ, Cairns SD (2017) Introduction to the state of deep-sea coral and sponge ecosystems of the United States. In: Hourigan TF, Etnoyer PJ, Cairns SD (eds) The State of Deep-Sea Coral and Sponge Ecosystems of the United States, pp 1–38. NOAA Technical Memorandum NFS-OHC-4. Silver Spring: NOAA
Hsieh C-F (2002) Composition, endemism and phytogeographical affinities of the Taiwan flora. Taiwania 47:298–310
Hug LA, Castelle CJ, Wrighton KC, Thomas BC, Sharon I, Frischkorn KR, Williams KH, Tringe SG, Banfield JF (2013) Community genomic analyses constrain the distribution of metabolic traits across the Chloroflexi phylum and indicate roles in sediment carbon cycling. Microbiome 1:1–22. https://doi.org/10.1186/2049-2618-1-22
Iliffe TM, Kornicker LS (2009) Worldwide diving discoveries of living fossil animals from the depths of anchialine and marine caves. In: Lang MA, Macintyre IG, Rützler K (eds.) Proceedings of the Smithsonian Marine Science Symposium. Smithsonian Contributions to the Marine Sciences, pp 269–280
Ise Y (2019) Preliminary report of submarine cave sponges in Shimoji Island, Miyako Islands, Okinawa. Taxa 46:13–17
Ise Y, Vacelet J, Mizuyama M, Fujita Y (2023) New lithistid sponge of the genus Sollasipelta (Porifera, Demospongiae, Tetractinellida, Neopeltidae) from submarine caves of the Ryukyu Islands, southwestern Japan, with redescription of S. sollasi. Zootaxa 5285(2):293–310. https://doi.org/10.11646/zootaxa.5285.2.4
Jackson DJ, Wörheide G (2014) Symbiophagy and biomineralization in the “living fossil” Astrosclera willeyana. Autophagy 10:408–415
Jackson DJ, Macis L, Reitner J, Wörheide G (2011) A horizontal gene transfer supported the evolution of an early metazoan biomineralization strategy. BMC Evol Biol 11:1–7
Kao WH, Chang SY, Liu HC, Chen MH (2007) The mollusks from intertidal and marine shallow water of Lanyu (Orchid Island), Taiwan. Platax: 69–92. https://doi.org/10.29926/PLATAX.200712.0007
Karlińska-Batres K, Wörheide G (2013a) Phylogenetic diversity and community structure of the symbionts associated with the coralline sponge Astrosclera willeyana of the Great Barrier Reef. Microb Ecol 65:740–752. https://doi.org/10.1007/s00248-013-0212-5
Karlińska-Batres K, Wörheide G (2013b) Microbial diversity in the coralline sponge Vaceletia crypta. Antonie Van Leeuwenhoek 103:1041–1056
Karlińska-Batres K, Wörheide G (2015) Spatial variability of microbial communities of the coralline demosponge Astrosclera willeyana across the Indo-Pacific. Aquat Microb Ecol 74:143–156. https://doi.org/10.3354/ame01732
Kelly-Borges M, Pomponi SA (1994) Phylogeny and classification of lithistid sponges (Porifera: Demospongiae): a preliminary assessment using ribosomal DNA sequence comparisons. Mol Mar Biol Biotechnol 3:87–103
Kim J, Kim G (2017) Inputs of humic fluorescent dissolved organic matter via submarine groundwater discharge to coastal waters off a volcanic island (Jeju, Korea). Sci Rep 7:7921. https://doi.org/10.1038/s41598-017-08518-5
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41(1):e1. https://doi.org/10.1093/nar/gks808
Konstantinou D, Gerovasileiou V, Voultsiadou E, Gkelis S (2018) Sponges-Cyanobacteria associations: Global diversity overview and new data from the Eastern Mediterranean. PLoS ONE 13(3):e0195001. https://doi.org/10.1371/journal.pone.0195001
Kuo J, Yang YT, Lu MC, Wong TY, Sung PJ, Huang YS (2019) Antimicrobial activity and diversity of bacteria associated with Taiwanese marine sponge Theonella swinhoei. Ann Microbiol 69:253–265
Landry Z, Swan BK, Herndl GJ, Stepanauskas R, Giovannoni SJ (2017) SAR202 genomes from the dark ocean predict pathways for the oxidation of recalcitrant dissolved organic matter. mBio 8:e00413-17. https://doi.org/10.1128/mBio.00413-17
Lang JC, Hartman WD, Land LS (1975) Sclerosponges: primary framework constructors on the Jamaican fore-reef. J Mar Res 332:223–231
Lee SC (1980) Intertidal fishes of the rocky pools at Lanyu (Botel Tobago), Taiwan. Bull Inst Zool Acad Sin 19:1–13
Lee SC, Lee CH (1981) A collection of subtidal fishes from Lanyu (Botel Tobago), Taiwan with a note on a newly recorded serranid, Ypsigramma lineata. Bull Inst Zool Acad Sin 20:87–89
Lurgi M, Thomas T, Wemheuer B, Webster NS, Montoya JM (2019) Modularity and predicted functions of the global sponge-microbiome network. Nat Commun 10:992. https://doi.org/10.1038/s41467-019-08925-4
Macartney KJ, Pankey MS, Slattery M, Lesser MP (2020) Trophodynamics of the sclerosponge Ceratoporella nicholsoni along a shallow to mesophotic depth gradient. Coral Reefs 39:1829–1839
Maldonado M, Aguilar R, Blanco J, Garcia S, Serrano A, Punzon A (2015) Aggregated clumps of lithistid sponges: a singular, reef-like bathyal habitat with relevant paleontological connections. PLoS ONE 10:e0125378
Maldonado M, López-Acosta M, Sitjà C, García-Puig M, Galobart C, Ercilla G, Leynaert A (2020) Sponge skeletons as an important sink of silicon in the global oceans. Nat Geosci 12:815–822. https://doi.org/10.1038/s41561-019-0430-7
Manconi R, Serusi A (2008) Rare sponges from marine caves: discovery of Neophrissospongia nana nov. sp. (Demospongiae, Corallistidae) from Sardinia with an annotated checklist of Mediterranean lithistids. ZooKeys 4:71–87. https://doi.org/10.3897/zookeys.4.39
Manconi R, Serusi A, Pisera A (2006) A new Mediterranean ‘lithistid’ sponge, Aciculites mediterranea sp. nov. (Porifera: Demospongiae) from a dark marine cave in Sardinia. J Mar Biol Assoc UK 86:691–698
McMurdie PJ, Holmes S (2013) phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217
McMurray S, Stubler A, Erwin P, Finelli C, Pawlik J (2018) A test of the sponge-loop hypothesis for emergent Caribbean reef sponges. Mar Ecol Prog Ser 588:1–14. https://doi.org/10.3354/meps12466
Meroz-Fine E, Shefer S, Ilan M (2005) Changes in morphology and physiology of an East Mediterranean sponge in different habitats. Mar Biol 147:243–250. https://doi.org/10.1007/s00227-004-1532-2
Moitinho-Silva L, Steinert G, Nielsen S, Hardoim CCP, Wu Y-C, Mccormack GP, López-Legentil S, Marchant R, Webster N, Thomas T, Hentschel U (2017) Predicting the HMA-LMA status in marine sponges by machine learning. Front Microbiol 8:752. https://doi.org/10.3389/fmicb.2017.00752
Morrow C, Cárdenas P (2015) Proposal for a revised classification of the Demospongiae (Porifera). Front Zool 12:1–27
Oksanen J, Simpson GL, Blanchet FG, Kindt R, Legendre P, Minchin PR, O'Hara RB, Solymos P, Stevens MHM, Szoecs E, Wagner H, Barbour M, Bedward M, Bolker B, Borcard D, Carvalho G, Chirico M, De Caceres M, Durand S, Evangelista HBA, FitzJohn R, Friendly M, Furneaux B, Hannigan G, Hill MO, Lahti L, McGlinn D, Ouellette M-H, Cunha ER, Smith T, Stier A, Ter Braak CJF, Weedon J (2022) Vegan: Community Ecology Package. R package version 2.6–4. https://www.rdocumentation.org/packages/vegan/versions/2.6-4. Accessed 28 Nov 2023
Pisera A, Gerovasileiou V (2021) Lithistid demosponges of deep-water origin in marine caves of the North-Eastern Mediterranean Sea. Front Mar Sci 8:630900. https://doi.org/10.3389/fmars.2021.630900
Pita L, Turon X, López-Legentil S, Erwin PM (2013) Host rules: spatial stability of bacterial communities associated with marine sponges (Ircinia spp.) in the Western Mediterranean Sea. FEMS Microbiol Ecol 86:268–276. https://doi.org/10.1111/1574-6941.12159
Podell S, Blanton JM, Oliver A, Schorn MA, Agarwal V, Biggs JS, Moore BS, Allen EE (2020) A genomic view of trophic and metabolic diversity in clade-specific Lamellodysidea sponge microbiomes. Microbiome 8:97. https://doi.org/10.1186/s40168-020-00877-y
Polónia ARM, Cleary DFR, Freitas R, de Voogd NJ, Gomes NCM (2015) The putative functional ecology and distribution of archaeal communities in an Indonesian coral reef environment. Mol Ecol 24:409–423. https://doi.org/10.1111/mec.13024
Polónia ARM, Cleary DFR, Freitas R, Coelho FJRC, de Voogd NJ, Gomes NCM (2016) Comparison of archaeal and bacterial communities in two sponge species and seawater from an Indonesian coral reef environment. Mar Genom 29:69–80. https://doi.org/10.1016/j.margen.2016.04.014
Polónia ARM, Cleary DFR, Freitas R, Gomes NCM, de Voogd NJ (2017) Archaeal and bacterial diversity, composition and predicted function in Xestospongia testudinaria and sediment in a coral reef environment. J Sea Res 119:37–53. https://doi.org/10.1016/j.seares.2016.10.007
Polónia ARM, Cleary DFR, Coelho FJRC, Becking LE, de Voogd NJ, Toha HA, Gomes NCM (2018) Compositional analysis of archaeal communities in seawater, sediment and high and low microbial abundance sponges in the Misool coral reef system, Indonesia. Mar Biol Res 14:537–550. https://doi.org/10.1080/17451000.2018.1498977
Polónia ARM, Cleary DFR, Gauvin-Bialecki A, de Voogd NJ (2021) Archaeal communities of low and high microbial abundance sponges inhabiting the remote western Indian Ocean island of Mayotte. Antonie Van Leeuwenhoek 114:95–112. https://doi.org/10.1007/s10482-020-01503-5
R Core Team (2022) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/
Radax R, Rattei T, Lanzen A, Bayer C, Rapp HT, Urich T, Schleper C (2012) Metatranscriptomics of the marine sponge Geodia barretti: tackling phylogeny and function of its microbial community. Environ Microbiol 14:1308–1324. https://doi.org/10.1111/j.1462-2920.2012.02714.x
Reigle NJ (1963) Notes on the mollusks of Lanyu, Taiwan. Quart J Taiwan Mus 16:81–87
Reveillaud J, Maignien L, Eren AM, Huber JA, Apprill A, Sogin ML, Vanreusel A (2014) Host-specificity among abundant and rare taxa in the sponge microbiome. ISME J 8:1198–1209. https://doi.org/10.1038/ismej.2013.227
Ribes M, Dziallas C, Coma R, Riemann L (2015) Microbial diversity and putative diazotrophy in high- and low-microbial-abundance Mediterranean sponges. Appl Environ Microbiol 81:5683–5693. https://doi.org/10.1128/AEM.01320-15
Richard M, Bellon H, Maury RC, Barrier E, Juang WS (1986) Miocene to recent calc-alkalic volcanism in eastern Taiwan: K-Ar ages and petrography. Tectonophysics 125:87–102
Ridley CP, John Faulkner D, Haygood MG (2005) Investigation of Oscillatoria spongeliae-dominated bacterial communities in four dictyoceratid sponges. Appl Environ Microbiol 71:7366–7375. https://doi.org/10.1128/AEM.71.11.7366-7375.2005
Rooks C, Fang JKH, Mørkved PT, Zhao R, Rapp HT, Xavier JR, Hoffmann F (2020) Deep-sea sponge grounds as nutrient sinks: denitrification is common in Boreo-Arctic sponges. Biogeosciences 17:1231–1245. https://doi.org/10.5194/bg-17-1231-2020
Rützler K, Vacelet J (2002) Family Acanthochaetetidae Fischer, 1970. In: Hooper JNA, van Soest RWM (eds) Systema Porifera: A Guide to the Classification of Sponges. Kluwer Academic/Plenum Publishers, New York, pp 275-278
Schmitt S, Deines P, Behnam F, Wagner M, Taylor MW (2011) Chloroflexi bacteria are more diverse, abundant, and similar in high than in low microbial abundance sponges. FEMS Microbiol Ecol 78:497–510. https://doi.org/10.1111/j.1574-6941.2011.01179.x
Schorn MA, Jordan PA, Podell S, Blanton JM, Agarwal V, Biggs JS, Allen EE, Moore BS (2019) Comparative genomics of cyanobacterial symbionts reveals distinct, specialized metabolism in tropical Dysideidae sponges. mBio 10(3):e00821-19. https://doi.org/10.1128/mBio.00821-19
Schuster A, Pisera A, Kelly M, Bell LJ, Pomponi SA, Wörheide G, Erpenbeck D (2018) New species and a molecular dating analysis of Vetulina Schmidt, 1879 (Porifera: Demospongiae: Sphaerocladina) reveal an ancient relict fauna with Tethys origin. Zool J Linn Soc 184:585–604. https://doi.org/10.1093/zoolinnean/zlx114
Schuster A, Pomponi SA, Pisera A, Cárdenas P, Kelly M, Wörheide G, Erpenbeck D (2021) Systematics of ‘lithistid’ tetractinellid demosponges from the Tropical Western Atlantic-implications for phylodiversity and bathymetric distribution. PeerJ 9:e10775. https://doi.org/10.7717/peerj.10775
Sket B (1996) The ecology of anchialine caves. Trends Ecol Evol 11:221–225. https://doi.org/10.1016/0169-5347(96)20031-X
Sollas WJ (1888) Report on the Tetractinellida collected by HMS Challenger, during the years 1873–1876. Zoology 25(part 63):1–458
Swierts T, Cleary DFR, de Voogd NJ (2018) Biogeography of prokaryote communities in closely related giant barrel sponges across the Indo-Pacific. FEMS Microbiol Ecol. 94:fiy194. https://doi.org/10.1093/femsec/fiy194
Taylor MW, Schupp PJ, de Nys R, Kjelleberg S, Steinberg PD (2005) Biogeography of bacteria associated with the marine sponge Cymbastela concentrica. Environ Microbiol 7:419–433. https://doi.org/10.1111/j.1462-2920.2004.00711.x
Usher KM (2008) The ecology and phylogeny of cyanobacterial symbionts in sponges. Mar Ecol 29:178–192
Vacelet J (1985) Coralline sponges and the evolution of Porifera. In: Conway Morris S, George JD, Gibson R, Platt HM (eds) The origins and relationships of lower invertebrates. Clarendon Press, Oxford, pp 1–13
Vacelet J (1996) Deep-sea sponges in a Mediterranean cave. Biosystem Ecol Ser 11:299–312
Vacelet J, Boury-Esnault N, Harmelin JG (1994) Hexactinellid cave, a unique deep-sea habitat in the scuba zone. Deep-Sea Res 41:965–973. https://doi.org/10.1016/0967-0637(94)90013-2
Valeria D’Auria MV, Zampella A, Zollo F (2002) The Chemistry of Lithistid Sponge: a Spectacular Source of New Metabolites. Stud Nat Prod Chem 26:1175. https://doi.org/10.1016/S1572-5995(02)80027-8
Varela MM, van Aken HM, Herndl GJ (2008) Abundance and activity of Chloroflexi-type SAR202 bacterioplankton in the meso- and bathypelagic waters of the (sub)tropical Atlantic. Environ Microbiol 10:1903–1911. https://doi.org/10.1111/j.1462-2920.2008.01627.x
Wang CH, Burnett WC (1990) Holocene mean uplift rates across an active plate-collision boundary in Taiwan. Science 248:204–206. https://doi.org/10.1126/science.248.4952.204
Xavier JR, Rees DJ, Pereira R, Colaço A, Pham CK, Carvalho FC (2021) Diversity, distribution and phylogenetic relationships of deep-sea lithistids (Porifera, Heteroscleromorpha) of the Azores Archipelago. Front Mar Sci 8:479
Yang TF, Lee T, Chen C-H, Cheng S-N, Knittel U, Punongbayan R, Rasdas AR (1996) A double island arc between Taiwan and Luzon: consequence of ridge subduction. Tectonophysics 258:85–101
Yu HP (1985) Notes on the land hermit crabs (Crustacea, Decapoda, Coenobitidae) from Lan-yu Island in the southern Taiwan. J Taiwan Mus 38:59–64
Acknowledgements
This work was made possible with the help of Julian, Floris, and Kate. We would also like to thank Tung Lung Diving for their help and support. We thank Rob Langelaan for his support in the lab. We also express our gratitude to two anonymous reviewers who helped to improve the manuscript.
Funding
Open access funding provided by FCT|FCCN (b-on). This work is part of the research programme NWO-VIDI with project number 16.161.301 by the Netherlands Organization for Scientific Research (NWO). This work was also supported by European Funds through COMPETE [FCOMP-01–0124-FEDER-008657] and by National Funds through the Portuguese Foundation for Science and Technology (FCT) within the LESS CORAL [PTDC/AAC-AMB/115304/2009] project. We also acknowledge financial support to CESAM from FCT/MCTES (UIDP/50017/2020 + UIDB/50017/2020 + LA/P/0094/2020), through national funds.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
No animal testing was performed during this study.
Sampling and field studies
All necessary permits for sampling and observational field studies have been obtained by the authors from the competent authorities.
Data availability
Sequences used in this study have been uploaded to the NCBI ShortRead Archive (BioProject nr: PRJNA865713 and BioSample nr: SUB11909922).
Author contribution
DFRC collected specimens in the field, analysed the data, and wrote the manuscript. NJdeV collected specimens in the field, identified the sponge specimens, and helped to write the manuscript. MvdP performed the laboratory analyses. ARMP and NCMG helped to write the manuscript.
Additional information
Communicated by M. Klautau
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12526_2023_1387_MOESM1_ESM.xls
Supplementary file1 (XLS 24 KB) Supplementary data 1. Supplementary data 1. List of samples used in the present study including the sample code (Sample), biotope code (Bio), Biotope, Species, sampling date (Date), sampling site (Site), Country, Depth, Latitude (Lat), Longitude (Lon) and location (Country).
12526_2023_1387_MOESM3_ESM.xls
Supplementary file3 (XLS 22 KB) Supplementary data 3. List of 50 most abundant OTUs including OTU number and associated ASV code, taxonomy, and results of the BLAST analysis.
12526_2023_1387_MOESM4_ESM.xlsx
Supplementary file4 (XLSX 49 KB) Supplementary data 4. Results of emmeans analysis showing pairwise comparisons of differences in diversity indices.
12526_2023_1387_MOESM5_ESM.xlsx
Supplementary file5 (XLSX 68 KB) Supplementary data 5. Results of emmeans analysis showing pairwise comparisons of differences in the relative abundances of selected phyla.
12526_2023_1387_MOESM6_ESM.xlsx
Supplementary file6 (XLSX 68 KB) Supplementary data 6. Results of emmeans analysis showing pairwise comparisons of differences in the relative abundances of the most abundant classes.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cleary, D.F.R., Huang, Y.M., Polónia, A.R.M. et al. Sponges and their prokaryotic communities sampled from a remote karst ecosystem. Mar. Biodivers. 54, 8 (2024). https://doi.org/10.1007/s12526-023-01387-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12526-023-01387-4