Abstract
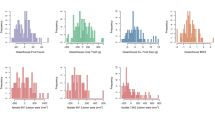
Enhanced bioavailability of cis-isomers of lycopene, accumulated in orange-fruited tangerine mutant has broadened the scope of nutritional enrichment in tomato. At the same time, advancements in the field of marker assisted selection (MAS) have made the stacking of multiple desirable alleles through molecular breeding to develop superior tomato genotypes possible. Here we report seedling stage MAS from 146 F2 plants, to identify 3 superior performing, root knot disease resistant orange-fruited segregants. In the selected segregants, fruit weight ranged from 39.2 to 54.6 g, pericarp thickness ranged from 4.56 to 6.05 mm and total soluble solid content ranged from 3.65 to 4.87° Brix. Presence of parental diversity allowed identification of the other desirable alleles of the genes governing late blight and mosaic disease resistance, growth habit (determinate and indeterminate) as well as fruit elongation and firmness. Resistance to root knot disease of the selected 3 segregants was also validated through a unique method employing in vitro rooted stem cuttings subjected to artificial inoculation, where the resistant parent and the selected segregants developed no galls in comparison to ~ 24 galls developed in the susceptible parent. The selected segregants form the base for development of multiple disease resistant, nutritionally enriched orange-fruited determinate/indeterminate tomato lines with superior fruit quality. The study also highlights the utility of early generation MAS for detailed characterization of segregants, through which multiple desirable alleles can be precisely targeted and fixed to develop superior tomato genotypes.








Similar content being viewed by others
Data availability
All necessary data are incorporated in the manuscript itself. Additional data, if any, are available from the corresponding author on reasonable request.
References
Anthon G, Barrett DM (2007) Standardization of a rapid spectrophotometric method for lycopene analysis. https://fruitandvegetable.ucdavis.edu/files/217095.pdf
Arens P, Mansilla C, Deinum D, Cavellini L, Moretti A, Rolland S, van der Schoot H, Calvache D, Ponz F, Collonnier C, Mathis R, Smilde D, Caranta C, Vosman B (2010) Development and evaluation of robust molecular markers linked to disease resistance in tomato for distinctness, uniformity and stability testing. Theor Appl Genet 120:655–664. https://doi.org/10.1007/s00122-009-1183-2
Bendezu IF (2004) Detection of the Mi 1.2 tomato gene by PCR using non-organic DNA purification. Nematropica 34(1):23–30
Bhagyashree, Maurya D, Chattopadhyay T (2020) Simultaneous detection of Mi1.2 and Ph3 resistance alleles for root knot and late blight diseases in tomato through duplex polymerase chain reaction. J Crop Weed 16(1):223–228. https://doi.org/10.22271/09746315.2020.v16.i1.1297
Bhavana P, Singh AK, Kumar R, Prajapati GK, Thamilarasi K, Manickam R, Maurya S, Choudhary JS (2019) Identification of resistance in tomato against root knot nematode (Meloidogyne incognita) and comparison of molecular markers for Mi gene. Australas Plant Pathol 48:93–100. https://doi.org/10.1007/s13313-018-0602-8
Black LL, Wang TC, Hanson PM, Chen JT (1996) Late blight resistance in four wild tomato accessions: effectiveness in diverse locations and inheritance of resistance. Phytopathology 86:S24
Burri BJ, Chapman MH, Neidlinger TR, Seo JS, Ishida BK (2009) Tangerine tomatoes increase total and tetra-cis-lycopene isomer concentrations more than red tomatoes in healthy adult humans. Int J Food Sci Nutr 60(Suppl 1):1–16. https://doi.org/10.1080/09637480701782084
Chattopadhyay T, Hazra P, Akhtar S, Maurya D, Mukherjee A, Roy S (2021) Skin colour, carotenogenesis and chlorophyll degradation mutant alleles: genetic orchestration behind the fruit colour variation in tomato. Plant Cell Rep 40:767–782. https://doi.org/10.1007/s00299-020-02650-9
Cooperstone JL, Ralston RA, Riedl KM, Haufe TC, Schweiggert RM, King SA, Timmers CD, Francis DM, Lesinski GB, Clinton SK, Schwartz SJ (2015) Enhanced bioavailability of lycopene when consumed as cis-isomers from tangerine compared to red tomato juice, a randomized, cross-over clinical trial. Mol Nutr Food Res 59(4):658–669. https://doi.org/10.1002/mnfr.201400658
Devran Z, Elekçioğlu IH (2004) The screening of F2 plants for the root-knot nematode resistance gene, Mi by PCR in tomato. Turk J Agric for 28:253–257
Devran Z, Söğüt MA, Mutlu N (2010) Response of tomato rootstocks with the Mi resistance gene to Meloidogyne incognita race 2 at different soil temperatures. Phytopathol Mediterr 49:11–17. https://doi.org/10.14601/Phytopathol_Mediterr-3116
El Mehrach K, Mejía L, Gharsallah-Couchane S, Salus MS, Martin CT, Hatimi A, Vidavski F, Williamson V, Maxwell DP (2005) PCR-based methods for tagging the Mi-1 locus for resistance to root-knot nematode in begomovirus-resistant tomato germplasm. Acta Hortic 695:263–270. https://doi.org/10.17660/ActaHortic.2005.695.29
El-Sappah AH, Islam MM, El-Awady HH, Yan S, Qi S, Liu J, Cheng GT, Liang Y (2019) Tomato natural resistance genes in controlling the root-knot nematode. Genes 10(11):925. https://doi.org/10.3390/genes10110925
Foolad MR, Panthee DR (2012) Marker-assisted selection in tomato breeding. Crit Rev Plant Sci 31(2):93–123. https://doi.org/10.1080/07352689.2011.616057
Galpaz N, Wang Q, Menda N, Zamir D, Hirschberg J (2008) Abscisic acid deficiency in the tomato mutant high-pigment 3 leading to increased plastid number and higher fruit lycopene content. Plant J 53(5):717–730. https://doi.org/10.1111/j.1365-313X.2007.03362.x
Garcia BE, Mejía L, Salus MS, Martin CT, Seah S, Williamson VM, Maxwell DP (2007) A co-dominant SCAR marker, Mi23, for detection of the Mi-1.2 gene for resistance to root-knot nematode in tomato germplasm. http://invirlab.plantpath.wisc.edu/GeminivirusResistantTomatoes/Markers/MAS-Protocols/Mi23-SCAR.pdf
Goggin FL, Jia L, Shah G, Hebert S, Williamson VM, Ullman DE (2006) Heterologous expression of the Mi-1.2 gene from tomato confers resistance against nematodes but not aphids in eggplant. Mol Plant Microbe Interact 19(4):383–388. https://doi.org/10.1094/MPMI-19-0383
Hazra P, Longjam M, Chattopadhyaya A (2018) Stacking of mutant genes in the development of “purple tomato” rich in both lycopene and anthocyanin contents. Sci Hortic 239:253–258. https://doi.org/10.1016/j.scienta.2018.05.039
Isaacson T, Ronen G, Zamir D, Hirschberg J (2002) Cloning of tangerine from tomato reveals a carotenoid isomerase essential for the production of β-Carotene and xanthophylls in plants. Plant Cell 14(2):333–342. https://doi.org/10.1105/tpc.010303
Jiang K, Liberatore KL, Park SJ, Alvarez JP, Lippman ZB (2013) Tomato yield heterosis is triggered by a dosage sensitivity of the florigen pathway that fine-tunes shoot architecture. PLoS Genet 9:e1004043. https://doi.org/10.1371/journal.pgen.1004043
Jung J, Kim HJ, Lee JM, Oh CS, Lee H-J, Yeam I (2015) Gene-based molecular marker system for multiple disease resistances in tomato against tomato yellow leaf curl virus, late blight, and verticillium wilt. Euphytica 205:599–613. https://doi.org/10.1007/s10681-015-1442-z
Kachanovsky DE, Filler S, Isaacson T, Hirschberg J (2012) Epistasis in tomato color mutations involves regulation of phytoene synthase 1 expression by cis-carotenoids. Proc Natl Acad Sci U S A 109(46):19021–19026. https://doi.org/10.1073/pnas.1214808109
Khachik F, Carvalho L, Bernstein PS, Muir GJ, Zhao D-Y, Katz NB (2002) Chemistry, distribution, and metabolism of tomato carotenoids and their impact on human health. Exp Biol Med 227(10):845–851. https://doi.org/10.1177/153537020222701002
Kolotilin I, Koltai H, Tadmor Y, Bar-Or C, Reuveni M, Meir A, Nahon S, Shlomo H, Chen L, Levin I (2007) Transcriptional profiling of high pigment-2dg tomato mutant links early fruit plastid biogenesis with its overproduction of phytonutrients. Plant Physiol 145(2):389–401. https://doi.org/10.1104/pp.107.102962
Krieger U, Lippman ZB, Zamir D (2010) The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato. Nat Genet 42:459–463. https://doi.org/10.1038/ng.550
Kumar V, Kumar P, Chattopadhyay T (2017) A rapid and reproducible method for isolating genomic DNA from a few crop plants suitable for polymerase chain reaction-based genotyping. J Appl Nat Sci 9:1119–1122. https://doi.org/10.31018/jans.v9i2.1332
Kumar A, Jindal SK, Dhaliwal MS, Sharma A, Kaur S, Jain S (2019) Gene pyramiding for elite tomato genotypes against ToLCV (Begomovirus spp.), late blight (Phytophthora infestans) and RKN (Meloidogyne spp.) for northern India farmers. Physiol Mol Biol Plants 25(5):1197–1209. https://doi.org/10.1007/s12298-019-00700-5
Levin I, Frankel P, Gilboa N, Tanny S, Lalazar A (2003) The tomato dark green mutation is a novel allele of the tomato homolog of the DEETIOLATED1 gene. Theor Appl Genet 106(3):454–460. https://doi.org/10.1007/s00122-002-1080-4
Lin T, Zhu G, Zhang J, Xu X, Yu Q, Zheng Z, Zhang Z, Lun Y, Li S, Wang X, Huang Z, Li J, Zhang C, Wang T, Zhang Y, Wang A, Zhang Y, Lin K, Li C, Xiong G, Xue Y, Mazzucato A, Causse M, Fei Z, Giovannoni JJ, Chetelat RT, Zamir D, Städler T, Li J, Ye Z, Du Y, Huang S (2014) Genomic analyses provide insights into the history of tomato breeding. Nat Genet 46(11):1220–1226. https://doi.org/10.1038/ng.3117
Liu J, van Eck J, Cong B, Tanksley SD (2002) A new class of regulatory genes underlying the cause of pear-shaped tomato fruit. Proc Natl Acad Sci U S A 99:13302–13306. https://doi.org/10.1073/pnas.162485999
Liu Y, Roof S, Ye Z, Barry C, van Tuinen A, Vrebalov J, Bowler C, Giovannoni J (2004) Manipulation of light signal transduction as a means of modifying fruit nutritional quality in tomato. Proc Natl Acad Sci U S A 101(26):9897–9902. https://doi.org/10.1073/pnas.0400935101
Liu J, Zhang J, Miao H, Jia C, Wang J, Xu B, Jin Z (2017) Elucidating the mechanisms of the tomato ovate mutation in regulating fruit quality using proteomics analysis. J Agric Food Chem 65:10048–10057. https://doi.org/10.1021/acs.jafc.7b03656
Maurya D, Bhagyashree, Akhtar S, Chattopadhyay T (2020) Exploring allelic status of selected disease resistance genes in a set of tomato genotypes using gene-linked molecular markers. J Crop Weed 16(1):236–241. https://doi.org/10.22271/09746315.2020.v16.i1.1299
Maurya D, Mukherjee A, Akhtar S, Chattopadhyay T (2021) Development and validation of the OVATE gene-based functional marker to assist fruit shape selection in tomato. 3 Biotech 11:474. https://doi.org/10.1007/s13205-021-03029-7
Maurya D, Mukherjee A, Bhagyashree, Sangam S, Kumar R, Akhtar S, Chattopadhyay T (2023) Marker assisted stacking of Ty3, Mi1.2 and Ph3 resistance alleles for leaf curl, root knot and late blight diseases in tomato. Physiol Mol Biol Plants 29(1):121–129. https://doi.org/10.1007/s12298-022-01277-2
Milligan SB, Bodeau J, Yaghoobi J, Kaloshian I, Zabel P, Williamson VM (1998) The root-knot resistance gene Mi from tomato is a member of the leucine zipper, nucleotide binding, leucine-rich repeat family of plant genes. Plant Cell 10(8):1307–1319. https://doi.org/10.1105/tpc.10.8.1307
Mukherjee A, Akhtar S, Chattopadhyay T (2021) Development of tetra-primer amplicon refractory mutation system (T-ARMS) strategy for identification of the dark green mutant allele in tomato. 3 Biotech 11:182. https://doi.org/10.1007/s13205-021-02732-9
Mustilli AC, Fenzi F, Ciliento R, Alfano F, Bowler C (1999) Phenotype of the tomato high pigment-2 mutant is caused by a mutation in the tomato homolog of DEETIOLATED1. Plant Cell 11(2):145–157. https://doi.org/10.1105/tpc.11.2.145
Nombela G, Williamson VM, Muñiz M (2003) The root-knot nematode resistance gene Mi-1.2 of tomato is responsible for resistance against the whitefly Bemisia tabaci. Mol Plant Microbe Interact 16(7):645–649. https://doi.org/10.1094/MPMI.2003.16.7.645
Perveen R, Suleria HAR, Anjum FM, Butt MS, Pasha I, Ahmad S (2015) Tomato (Solanum lycopersicum) carotenoids and lycopenes chemistry; metabolism, absorption, nutrition and allied health claims-a comprehensive review. Crit Rev Food Sci Nutr 55(7):919–929. https://doi.org/10.1080/10408398.2012.657809
Pnueli L, Carmel-Goren L, Hareven D, Gutfinger T, Alvarez J, Ganal M, Zamir D, Lifschitz E (1998) The SELF-PRUNING gene of tomato regulates vegetative to reproductive switching of sympodial meristems and is the ortholog of CEN and TFL1. Development 125(11):1979–1989. https://doi.org/10.1242/dev.125.11.1979
Przybylska S (2020) Lycopene–a bioactive carotenoid offering multiple health benefits: a review. Int J Food Sci Tech 55(1):11–32. https://doi.org/10.1111/ijfs.14260
Rodríguez GR, Munos S, Anderson C, Sim S-C, Michel A, Causse M, Gardener BBM, Francis D, van der Knaap E (2011) Distribution of SUN, OVATE, LC, and FAS in the tomato germplasm and the relationship to fruit shape diversity. Plant Physiol 156:275–285. https://doi.org/10.1104/pp.110.167577
Seah S, Williamson VM, Garcia BE, Mejıa L, Salus MS, Martin CT, Maxwell DP (2007) Evaluation of a co-dominant SCAR marker for detection of the Mi-1 locus for resistance to root-knot nematode in tomato germplasm. https://tgc.ifas.ufl.edu/vol57/html/vol57seah.htm
Sujatha R, Irene VP, Manivannan N, Sivam KM (2017) Screening of tomato genotypes for root knot nematode (Meloidogyne incognita Kofoid and White Chitwood). Int J Curr Microbiol App Sci 6(3):1525–1533. https://doi.org/10.20546/ijcmas.2017.603.175
Yoo HJ, Park WJ, Lee G-M, Oh C-S, Yeam I, Won D-C, Kim CK, Lee JM (2017) Inferring the genetic determinants of fruit colors in tomato by carotenoid profiling. Molecules 22(5):764. https://doi.org/10.3390/molecules22050764
Zhang C, Liu L, Zheng Z, Sun Y, Zhou L, Yang Y, Cheng F, Zhang Z, Wang X, Huang S, Xie B, Du Y, Bai Y, Li J (2013) Fine mapping of the Ph-3 gene conferring resistance to late blight (Phytophthora infestans) in tomato. Theor Appl Genet 126:2643–2653. https://doi.org/10.1007/s00122-013-2162-1
Zhang C, Liu L, Wang X, Vossen J, Li G, Li T, Zheng Z, Gao J, Guo Y, Visser RG, Li J, Bai Y, Du Y (2014) The Ph-3 gene from Solanum pimpinellifolium encodes CC-NBS-LRR protein conferring resistance to Phytophthora infestans. Theor Appl Genet 127:1353–1364. https://doi.org/10.1007/s00122-014-2303-1
Acknowledgements
AK and VL acknowledge BAU Sabour and SS acknowledge University Grants Commission, Government of India for providing fellowship. Financial support in terms of project grant (code: SNP/CI/Rabi/2018-5) provided by BAU Sabour is highly acknowledged. This article bears BAU COMMUNICATION NO. 1460/230628
Funding
This work is supported by the university-funded project entitled: Marker assisted breeding for different disease resistance alleles in tomato, Code: SNP/CI/Rabi/2018–5.
Author information
Authors and Affiliations
Contributions
TC acquired grants and designed the experiments. AK and VL contributed equally in this study. Molecular biology experiments were performed by AK, VL, SS and TC. VL, AK, MK and SA recorded the biochemical and morphological fruit quality parameters. TC, AK, VL, SS and SA were involved in crop management and morphological selection. TNG, AK, VL and TC performed the root knot assay. Analyses of molecular, morphological and biochemical data were performed by TC, AK, VL and TNG. TC, AK and VL prepared the manuscript draft. All the authors took part in revision and finalization of the draft manuscript and approved the final version of it.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12298_2023_1361_MOESM1_ESM.tif
Fig. Sf1 Amplification from SP locus using SP-dCAPS marker. Inverse image of 2.5% gel showing undigested amplicons of ~ 147 bp generated by SP-dCAPS F and SP Out R primers. Positions of relevant bands of the 100 bp ladder are mentioned. L = 100 bp DNA ladder (BioLit, SRL) (TIF 526 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumar, A., Lakshmi, V., Sangam, S. et al. Marker assisted early generation identification of root knot disease resistant orange tomato segregants with multiple desirable alleles. Physiol Mol Biol Plants 29, 1179–1192 (2023). https://doi.org/10.1007/s12298-023-01361-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-023-01361-1