Abstract
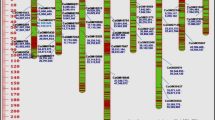
A priority in the management and use of elite plant materials for breeding has been based on molecular markers or DNA sequencing of entire genomes, in order to perform genetic differentiation which is still quite costly. Chickpea (Cicer arietinum) is one of the species with genomic monotony and very low polymorphism, and its detection even with DNA markers has not been easy. In germplasm banks, the genetic distinction is a priority in order to use properly selected lines. In this study, 57 chickpea accessions from a germplasm bank were analyzed by using nrRAMP (non-radioactive Random Amplified Microsatellite Polymorphism) markers, and their genetic variability was determined. Our results showed DNA polymorphisms, which are enough to differentiate between the accessions and between C. arietinum and Cicer reticulatum (out-group); this last wild species is closely related to chickpea. We concluded that the nrRAMP technique was an effective and a highly useful method to assess the genetic diversity and variability among closely related plants, such as chickpea; in addition, this technique can be easily implemented in laboratories.



Similar content being viewed by others
References
Ahmad F, Khan AI, Awan FS, Sadia B, Sadaqat HA, Bahadur S (2010) Genetic diversity of chickpea (Cicer arietinum L.) germplasm in Pakistan as revealed by RAPD analysis. Gen Mol Res 9:1414–1420
Alanís-Martínez I, Medina-Mendoza C, Marban-Mendoza N, Valadez-Moctezuma E (2015) Técnicas moleculares clásicas para la diferenciación de formas especiales de Fusarium oxysporum. Revista Mexicana de Micología 42:1–7
Aslam M, Maqbool MA, Akhtar S, Faisal W (2013) Estimation of genetic variability and association among different physiological traits related to biotic stress (Fusarium oxysporum l.) in chickpea. J Anim Plant Sci 23:1679–1685
Avila-Treviño JA, Muñóz-Alemán JM, Pérez-Molphe-Balch E, Rodríguez-Sahagún A, Morales-Domínguez JF (2017) In vitro propagation from bud and apex explants of Moringa oleifera and evaluation of the genetic stability with RAMP marker. S Afr J Bot 108:149–156
Cheng HY, Yang WC, Hsiao JY (2001) Genetic diversity and relationship among peach cultivars based on Random Amplified Microsatellite Polymorphism (RAMP). Bot Bull Acad Sin 42:201–206
Dávila JA, Loarce Y, Ferrer E (1999) Molecular characterization and genetic mapping of random amplified mirosatellite polymosphism in barley. Theor Appl Genet 98:265–273
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: version II. Plant Mol Biol Report 1:19–21
Dice LR (1945) Measures of the amount of ecological association between species. Ecology 26:297–302
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321
Hüttel B, Winter P, Weising K, Choumane W, Weigand F, Kahl G (1999) Sequence-tagged microsatellite site markers for chickpea (Cicer arietinum L.). Genome 42:210–217
Kazan K, Muehlbauer FJ (1991) Allozyme variation and phylogeny in annual species of Cicer (Leguminosae). Plant Syst Evol 175:11–21
Keneni G, Bekele E, Imtiaz M, Dagne K, Getu E, Assefa F (2012) Genetic diversity and population structure of Ethiopian chickpea (Cicer arietinum L.) germplasm accessions from different geographical origins as revealed by microsatellite markers. Plant Mol Biol Report 30:654–665
Lihua Z, Mingyang L, Guangze C, Tianchun P, Chenghai S (2013) Assessment of the genetic diversity and genetic relationships of pomegranate (Punica granatum L.) in China using RAMP markers. Sci Hortic 151:63–67
Nguyen TT, Taylor PWJ, Redden RJ, Ford R (2003) Genetic diversity estimates in Cicer using AFLP analysis. Plant Breed 123:173–179
Olsen KM, Wendel JF (2013) A bountiful harvest: genomic insights into crop domestication phenotypes. Annu Rev Plant Biol 64:47–70
Provan J, Thomas WTB, Forster BP, Powel W (1999) Copia-SSR: a simple marker technique which can be used on total genomic DNA. Genome 42:363–366
Robertson LD, Ocampo B, Singh KB (1997) Morphological variation in wild annual Cicer species in comparison to the cultigen. Euphytica 95:309–319
Saeed A, Hovsepyan H, Darvishzadeh R, Imtiaz M, Panguluri SK, Nazaryan R (2011) Genetic diversity of Iranian accessions, improved lines of chickpea (Cicer arietinum L.) and their wild relatives by using simple sequence repeats. Plant Mol Biol Report 29:848–858
Sánchez de la Hoz MP, Dávila JA, Loarce Y, Ferrer E (1996) Simple sequence repeat primers used in polymerase chain reaction amplifications to study genetic diversity in barley. Genome 39:112–117
Simon CJ, Muehlbauer FJ (1997) Construction of a chickpea linkage map and its comparison with maps of pea and lentil. J Hered 88:115–119
Talebi R, Rokhzadi A (2013) Genetic diversity and interrelationships between agronomic traits in landrace chickpea accessions collected from ‘Kurdistan’ province, North-west of Iran. Int J Agric Crop Sci 5:2203–2209
Udupa SM, Sharma A, Sharma RP, Pai RA (1993) Narrow genetic variability in Cicer arietinum L. as revealed by RFLP analysis. J Plant Biochem Biotechnol 2:83–86
Upadhyaya HD, Dwivedi SL, Baum M, Varshney RK, Udupa SM, Gowda CLL, Hoisington D, Singh S (2008) Genetic structure, diversity, and allelic richness in composite collection and reference set in chickpea (Cicer arietinum L.). BMC Plant Biol 8:106–118
Valadez-Moctezuma E, Kahl G, Rubluo-Islas A, Arreguín de los Monteros R (2005) Optimización de las huellas de DNA obtenidas con RAPDs y MP-PCR mediante la técnica nrRAMP. Revista Chapingo Serie Horticultura 11:351–356
Winter P, Kahl G (1995) Molecular marker technologies for plant improvement. World J Microbiol Biotechnol 11:438–448
Wu K, Jones R, Danneberger L, Scolnik PA (1994) Detection of microsatellite polymorphisms without cloning. Nucleic Acid Res 22:3257–3258
Yang RW, Zhou YH, Zhang Y, Zheng YL, Ding CB (2006) The genetic diversity among Leymus species based on random amplified microsatellite polymorphism (RAMP). Genet Resour Crop Evol 53:139–144
Zhang L, Zhou Y, Wei Y, Zheng Y, Liu S (2003) Relationships among Kengyilia species based on random amplified microsatellite polymorphism (RAMP). High Technol Lett 4:28–33
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Valadez-Moctezuma, E., Cabrera-Hidalgo, A.J. Easy strategy used to detect the genetic variability in chickpea (Cicer arietinum L.). Physiol Mol Biol Plants 24, 921–928 (2018). https://doi.org/10.1007/s12298-018-0548-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-018-0548-x