Abstract
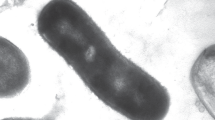
A nitrate-reducing Fe(II)-oxidizing bacterial strain, F8825T, was isolated from the Fe(II)-rich sediment of an urban creek in Pearl River Delta, China. The strain was Gram-negative, facultative chemolithotrophic, facultative anaerobic, non-spore-forming, and rod-shaped with a single flagellum. Phy-logenetic analysis based on 16S rRNA gene sequencing indicated that it belongs to the genus Ciceribacter and is most closely related to C. lividus MSSRFBL1T (99.4%), followed by C. thiooxidans F43bT (98.8%) and C. azotifigens A.slu09T (98.0%). Fatty acid, polar lipid, respiratory quinone, and DNA G + C content analyses supported its classification in the genus Ciceribacter. Multilocus sequence analysis of concatenated 16S rRNA, atpD, glnII, gyrB, recA, and thrC suggested that the isolate was a novel species. DNA-DNA hybridization and genome sequence comparisons (90.88 and 89.86%, for values of ANIm and ANIb between strains F8825T with MSSRFBL1T, respectively) confirmed that strain F8825T was a novel species, different from C. lividus MSSRFBL1T, C. thiooxidans F43bT, and C. azotifigens A.slu09T. The physiological and biochemical properties of the strain, such as carbon source utilization, nitrate reduction, and ferrous ion oxidation, further supported that this is a novel species. Based on the polyphasic taxonomic results, strain F8825T was identified as a novel species in the genus Ciceribacter, for which the name Ciceribacter ferrooxidans sp. nov. is proposed. The type strain is F8825T (= CCTCC AB 2018196T = KCTC 62948T).
Similar content being viewed by others
References
Bankevich, A., Nurk, S., Antipov, D., Gurevich, A.A., Dvorkin, M., Kulikov, A.S., Lesin, V.M., Nikolenko, S.I., Pham, S., Prjibelski, A.D., et al. 2012. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol.19, 455–477.
Chaudhuri, S.K., Lack, J.G., and Coates, J.D. 2001. Biogenic magnetite formation through anaerobic biooxidation of Fe(II). Appl. Environ. Microbiol.67, 2844–2848.
Chun, J., Oren, A., Ventosa, A., Christensen, H., Arahal, D.R., da Costa, M.S., Rooney, A.P., Yi, H., Xu, X.W., De Meyer, S., et al. 2018. Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int. J. Syst. Evol. Microbiol.68, 461–466.
Cole, J.J., Prairie, Y.T., Caraco, N.F., Mcdowell, W.H., Tranvik, L.J., Striegl, R.G., Duarte, C.M., Kortelainen, P., Downing, J.A., Mid-delburg, J.J., et al. 2007. Plumbing the global carbon cycle: integrating inland waters into the terrestrial carbon budget. Ecosystems10, 172–185.
Deng, T., Chen, X., Zhang, Q., Zhong, Y., Guo, J., Sun, G., and Xu, M. 2017. Ciceribacter thiooxidans sp. nov. a novel nitrate-reducing thiosulfate-oxidizing bacterium isolated from sulfide-rich anoxic sediment. Int. J. Syst. Evol. Microbiol.67, 4710–4715.
Gerhardt, P., Murray, R.G.E., Costilow, R.N., Nester, E.W., Wood, W.A., Krieg, N.R., and Phillips, G.B. 1981. Manual of methods for general bacteriology. American Society Microbiology, Washington, D.C., USA.
Huerta-Cepas, J., Forslund, K., Coelho, L.P., Szklarczyk, D., Jensen, L.J., von Mering, C., and Bork, P. 2017. Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol. Biol. Evol.34, 2115–2122.
Hyatt, D., Chen, G.L., Locascio, P.F., Land, M.L., Larimer, F.W., and Hauser, L.J. 2010. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics11, 119.
Kanehisa, M., Sato, Y., and Morishima, K. 2016. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol.428, 726–731.
Kathiravan, R., Jegan, S., Ganga, V., Prabavathy, V.R., Tushar, L., Sasikala, Ch., and Ramana, Ch.V. 2013. Ciceribacter lividus gen. nov. sp. nov. isolated from rhizosphere soil of chick pea (Cicerarietinum L.). Int. J. Syst. Evol. Microbiol.63, 4484–4488.
Kumar, M., Verma, M., and Lal, R. 2008. Devosia chinhatensis sp. nov., isolated from a hexachlorocyclohexane (HCH) dump site in India. Int. J. Syst. Evol. Microbiol.58, 861–865.
Lack, J.G., Chaudhuri, S.K., Kelly, S.D., Kemner, K.M., O’Connor, S.M., and Coates, J.D. 2002. Immobilization of radionuclides and heavy metals through anaerobic bio-oxidation of Fe(II). Appl. Environ. Microbiol.68, 2704–2710.
Lu, L.Y., Chen, W.F., Han, L.L., Wang, E.T., and Chen, W.X. 2009. Rhizobium alkalisoli sp. nov. isolated from Caragana intermedia growing in saline-alkaline soils in the north of China. Int. J. Syst. Evol. Microbiol.59, 3006–3011.
Martens, M., Delaere, M., Coopman, R., De Vos, P., Gillis, M., and Willems, A. 2007. Multilocus sequence analysis of Ensifer and related taxa. Int. J. Syst. Evol. Microbiol.57, 489–503.
Minnikin, D.E., O’Donnell, A.G., Goodfellow, M., Alderson, G., Athalye, M., Schaal, A., and Parlett, J.H. 1984. An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J. Microbiol. Methods2, 233–241.
Moore, D.D. and Dowhan, D. 2012. Preparation and analysis of DNA. Curr. Protoc. Mol. Biol.98, 2.0.1–2.0.3.
Nguyen, T.M. and Kim, J. 2017. A rapid and simple method for identifying bacterial polar lipid components in wet biomass. J. Microbiol.55, 635–639.
Nishijima, M., Araki-Sakai, M., and Sano, H. 1997. Identification of isoprenoid quinones by frit-FAB liquid chromatography-mass spectrometry for the chemotaxonomy of microorganisms. J. Microbiol. Methods28, 113–122.
Richter, M. and Rosselló-Móra, R. 2009. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. USA106, 19126–19131.
Richter, M., Rosselló-Móra, R., Oliver Glöckner, F., and Peplies, J. 2016. JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics32, 929–931.
Schaedler, F., Lockwood, C., Lueder, U., Glombitza, C., Kappler, A., and Schmidt, C. 2018. Microbially mediated coupling of Fe and N cycles by nitrate-reducing Fe(II)-oxidizing bacteria in littoral freshwater sediments. Appl. Environ. Microbiol.84, e02013–17.
Schippers, A. and Jørgensen, B.B. 2002. Biogeochemistry of pyrite and iron sulfide oxidation in marine sediments. Geochim. Cosmochim. Acta66, 85–92.
Senn, D.B. and Hemond, H.F. 2002. Nitrate controls on iron and arsenic in an urban lake. Science296, 2373–2376.
Siddiqi, M.Z., Choi, G.M., and Im, W.T. 2018. Ciceribacter azotifigens sp. nov. a nitrogen-fixing bacterium isolated from activated sludge. Int. J. Syst. Evol. Microbiol.68, 482–486.
Sinsabaugh, R.L., Hill, B.H., and Follstad Shah, J.J. 2009. Ecoenzymatic stoichiometry of microbial organic nutrient acquisition in soil and sediment. Nature462, 795–798.
Song, Y., Yang, R., Guo, Z., Zhang, M., Wang, X., and Zhou, F. 2000. Distinctness of spore and vegetative cellular fatty acid profiles of some aerobic endospore-forming bacilli. J. Microbiol. Methods39, 225–241.
Straub, K.L., Benz, M., Schink, B., and Widdel, F. 1996. Anaerobic, nitrate-dependent microbial oxidation of ferrous iron. Appl. Environ. Microbiol.62, 1458–1460.
Vaclavkova, S., Jørgensen, C.J., Jacobsen, O.S., Aamand, J., and Elberling, B. 2014. The importance of microbial iron sulfide oxidation for nitrate depletion in anoxic Danish sediments. Aquat. Geochem.20, 419–435.
Vinuesa, P., Silva, C., Werner, D., and Martinez-Romero, E. 2005. Population genetics and phylogenetic inference in bacterial molecular systematics: The roles of migration and recombination in Bradyrhizobium species cohesion and delineation. Mol. Phylogenet. Evol.34, 29–54.
Weber, K.A., Achenbach, L.A., and Coates, J.D. 2006a. Microorganisms pumping iron: anaerobic microbial iron oxidation and reduction. Nat. Rev. Microbiol.4, 752–764.
Weber, K.A., Urrutia, M.M., Churchill, P.F., Kukkadapu, R.K., and Roden, E.E. 2006b. Anaerobic redox cycling of iron by freshwater sediment microorganisms. Environ. Microbiol.8, 100–113.
Widdel, F., Schnell, S., Heising, S., Ehrenreich, A., Assmus, B., and Schink, B. 1993. Ferrous iron oxidation by anoxygenic phototrophic bacteria. Nature362, 834–836.
Xu, M., Zhang, Q., Xia, C., Zhong, Y., Sun, G., Guo, J., Yuan, T., Zhou, J., and He, Z. 2014. Elevated nitrate enriches microbial functional genes for potential bioremediation of complexly contaminated sediments. ISME J.8, 1932–1944.
Yassin, A.F., Galinski, E.A., Wohlfarth, A., Jahnke, K.D., Schaal, K.P., and Trüper, H.G. 1993. A new actinomycete species, Nocardiopsis lucentensis sp. nov. Int. J. Syst. Bacteriol.43, 266–271.
Zhang, M., Zheng, P., Wang, R., Li, W., Lu, H., and Zhang, J. 2014. Nitrate-dependent anaerobic ferrous oxidation (NAFO) by denitrifying bacteria: a perspective autotrophic nitrogen pollution control technology. Chemosphere117, 604–609.
Acknowledgments
We thank Korean Agricultural Culture Collection (KACC) for providing free strain C. azotifigens A.slu09T. This work was funded by the National Key R & D Program of China (2018YFD0500202), the National Natural Science Foundation of China (91851202, 51678163, 21677042), GDAS’ Special Project of Science and Technology Development (2019-GDASYL-0104005), Natural Science Foundation of Guangdong Province (2018B0303110010), Science and Technology Project of Guangzhou (201707020021), Guangdong Technological Innovation Strategy of Special Funds (Key Areas of Research and Development Program, 2018B020205003) and Guangdong MEPP Fund (GDOE2019A34).
Author information
Authors and Affiliations
Corresponding author
Additional information
Conflict of Interest
The authors declare no conflict of interest.
Supplemental material for this article may be found at http://www.springerlink.com/content/120956
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Deng, T., Qian, Y., Chen, X. et al. Ciceribacter ferrooxidans sp. nov., a nitrate-reducing Fe(II)-oxidizing bacterium isolated from ferrous ion-rich sediment. J Microbiol. 58, 350–356 (2020). https://doi.org/10.1007/s12275-020-9471-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-020-9471-2