Abstract
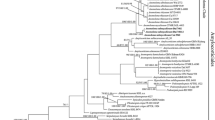
Recent studies based on morphological characteristics and molecular analyses have revealed that the characteristics of Sparassis crispa from Asia are not concordant with those of collections from Europe and North America. Consequently, the Asian isolate was redefined as Sparassis latifolia. This study is the first report of Sparassis latifolia collected in Korea. The taxonomic relationships and replacement of Sparassis species were inferred from a comparison of the morphological characteristics and by molecular sequence analysis of the internal transcribed spacer (ITS) rDNA regions. In particular, this study focused on the phylogenetic relationships inferred from the biogeographical distribution of isolates within the genus Sparassis.
Similar content being viewed by others
References
Blanco-Dios, J.B., Wang, Z., Binder, M., and Hibbett, D.S. 2006. A new Sparassis from Spain described using morphological and molecular data. Mycol. Res. 110, 1227–1231.
Dai, Y.C., Wang, Z., Binder, M., and Hibbett, D.S. 2006. Phylogeny and a new species of Sparassis (Polyporales, Basidiomycota): evidence from mitochondrial atp6, nuclear rDNA and rpb2 genes. Mycologia 98, 584–592.
Desjardin, D.E., Wang, Z., Binder, M., and Hibbett, D.S. 2004. Sparassis cystidiosa sp. nov., from Thailand is described using morphological and molecular data. Mycologia 96, 1010–1014.
Lee, S.B. and Taylor, J.W. 1990. Isolation of DNA from fungal mycelia and single spores, pp. 282–287. In Innis, M.A., Gelfand, D.H., Sninsky, J., and White, T.J. (eds.). PCR protocols. Academic Press. San Diego, California, USA.
Park, H., Lee, B.H., Oh, D.S., Ka, K.H., Bak, W.C., and Lee, H.J. 2005. Cultivation of cauliflower mushroom (Sparassis crispa) using coniferous sawdust-based media with barley flours. J. Kor. For. En. 24, 31–36.
Ronquist, F. and Huelsenbeck, J.P. 2003. MRBAYES 3: Bayesian phylogenetic inference under mixed molds. Bioinformatics 19, 1572–1574.
Ryu, S.R., Ka, K.H., Park, H., Bak, W.C., and Lee, B.H. 2009. Cultivation characteristics of Sparassis crispa strains using sawdust medium of Larix kaempferi. Korean J. Mycol. 37, 49–54.
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F., and Higgins, D.G. 1997. The CLUSTAL_X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882.
Wang, Z., Binder, M., Dai, Y.C., and Hibbett, D.S. 2004. Phylogenetic relationships of Sparassis inferred from nuclear and mitochondrial ribosomal DNA and RNA polymerase sequences. Mycologia 96, 1015–1029.
White, T.J., Bruns, T., Lee, S., and Taylor, J. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics, pp. 315–322. In Innis, M.A., Gelfand, D.H., Sninsky, J., and White, T.J. (eds.). PCR protocols. Academic Press. San Diego, California, USA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ryoo, R., Sou, HD., Ka, KH. et al. Phylogenetic relationships of Korean Sparassis latifolia based on morphological and ITS rDNA characteristics. J Microbiol. 51, 43–48 (2013). https://doi.org/10.1007/s12275-013-2503-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-013-2503-4