Abstract
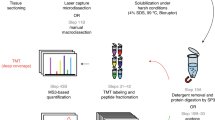
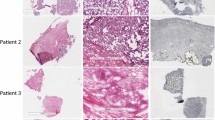
Distant metastasis is the major cause of cancer-related death in virtually all tumour types. We used spontaneous metastasis xenograft mouse models to study differences between primary tumours and corresponding metastases at the proteome level. Sampling of tissues for proteomics with mechanical homogenization is critical, because by this process many proteins are getting lost. For improving the protein yield infrared laser ablation was applied for sampling.
Similar content being viewed by others
Literatur
Lambert AW, Pattabiraman DR, Weinberg RA (2017) Emerging biological principles of metastasis. Cell 168:670–691
Schumacher U, Stürken C (2016) Das Rätsel der Metastasierung. Biologie in unserer Zeit 46:350–356, doi: 10.1002/biuz.201610606
Lange T, Ullrich S, Müller I et al. (2012) Human prostate cancer in a clinically relevant xenograft mouse model: identification of β(1,6)-branched oligosaccharides as a marker of tumor progression. Clin Cancer Res 18:1364–1373
Lange T, Kupfernagel M, Wicklein D et al. (2014) Aberrant presentation of HPA-reactive carbohydrates implies selectinindependent metastasis formation in human prostate cancer. Clin Cancer Res 20:1791–1802
Hänel L, Gosau T, Maar H et al. (2018) Differential proteome analysis of human neuroblastoma xenograft primary tumors and matched spontaneous distant metastases. Sci Rep 8:13986
Kwiatkowski M, Wurlitzer M, Omidi M et al. (2015) Ultrafast extraction of proteins from tissues using desorption by impulsive vibrational excitation. Angew Chem Int Ed Engl 54:285–288
Kwiatkowski M, Wurlitzer M, Krutilin A et al. (2016) Homogenization of tissues via picosecond-infrared laser (PIRL) ablation: giving a closer view on the in-vivo composition of protein species as compared to mechanical homogenization. J Proteomics 134:193–202
Aebersold R, Agar JN, Amster IJ et al. (2018) How many human proteoforms are there? Nat Chem Biol 14:206–214
Jungblut PR, Holzhütter HG, Apweiler R et al. (2008) The speciation of the proteome. Chem Cent J 2:16
Schlüter H, Apweiler R, Holzhütter HG et al. (2009) Finding one’s way in proteomics: a protein species nomenclature. Chem Cent J 3:11
Author information
Authors and Affiliations
Corresponding author
Additional information
Hartmut Schlüter 1981–1991 Chemiestudium und Promotion in Biochemie an der Universität Münster, dort 1991–1996 Habilitation im Fach Pathobiochemie. 1996–2000 Ruhr-Universität Bochum. 2000–2008 Charité, Berlin. 2003 Apl. Professur. Seit 2008 Professur für Massenspektrometrische Proteomanalytik am Universitätsklinikum Hamburg-Eppendorf. Seit 2017 Mitglied im Institut für Biochemie, Universität Hamburg.
Tobias Lange 2003–2009 Humanmedizinstudium und Promotion am Universitätsklinikum Hamburg-Eppendorf (UKE), dort 2010–2013 wissenschaftlicher Mitarbeiter am Institut für Anatomie und Experimentelle Morphologie und 2011–2014 MD/PhD-Studium und Promotion. Seit 2013 Juniorprofessor für Translationale Krebsforschung am UKE, 2015 Facharzt für Anatomie. Seit 2017 stellvertretender Institutsdirektor des Instituts für Anatomie und Experimentelle Morphologie, UKE.
Rights and permissions
About this article
Cite this article
Lange, T., Schlüter, H. Probennahme per Laserablation von Krebsgeweben für Proteomics. Biospektrum 25, 32–34 (2019). https://doi.org/10.1007/s12268-019-1000-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12268-019-1000-7