Abstract
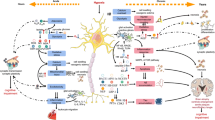
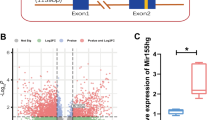
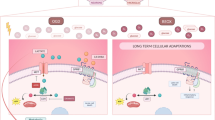
Heat-stroke is a serious form of hyperthermia with high mortality, and can induce severe central nervous system disorders. The neurovascular unit (NVU), which consists of vascular cells, glial cells, and neurons, controls blood-brain barrier (BBB) permeability and cerebral blood flow, and maintains the proper functioning of neuronal circuits. However, the detailed function of each BBB component in heat-stroke remains unknown. In order to interpret alterations caused by heat stress, we performed transcriptome comparison of neuron and astrocyte primary cultures after heat treatment. Differentially-expressed genes were then selected and underwent Gene Ontology annotation and Kyoto Encyclopedia of Genes and Genomes pathway analysis. Gene-act networks were also constructed, and the expression of pivotal genes was validated by quantitative PCR, as well as single-cell qPCR in heat-stroke rats. Our work provides valuable information on the transcriptional changes in NVU cells after heat stress, reveals the diverse regulatory mechanisms of two of these cellular components, and shows that a cell-type-specific approach may be a promising therapeutic strategy for heat-stroke treatments.







Similar content being viewed by others
References
Bouchama A, Knochel JP. Heat stroke. N Engl J Med 2002, 346: 1978–1988.
Kovats RS, Kristie LE. Heatwaves and public health in Europe. Eur J Public Health 2006, 16: 592–599.
Audet GN, Dineen SM, Quinn CM, Leon LR. Altered hypothalamic inflammatory gene expression correlates with heat stroke severity in a conscious rodent model. Brain Res 2016, 1637: 81–90.
Argaud L, Ferry T, Le QH, Marfisi A, Ciorba D, Achache P, et al. Short-and long-term outcomes of heatstroke following the 2003 heat wave in Lyon, France. Arch Intern Med 2007, 167: 2177–2183.
Zlokovic BV. Neurovascular pathways to neurodegeneration in Alzheimer’s disease and other disorders. Nat Rev Neurosci 2011, 12: 723–738.
Epstein Y, Roberts WO. The pathopysiology of heat stroke: an integrative view of the final common pathway. Scand J Med Sci Sports 2011, 21: 742–748.
Liu Z, Chopp M. Astrocytes, therapeutic targets for neuroprotection and neurorestoration in ischemic stroke. Prog Neurobiol 2016, 144: 103–120.
Citri A, Pang ZP, Sudhof TC, Wernig M, Malenka RC. Comprehensive qPCR profiling of gene expression in single neuronal cells. Nat Protoc 2011, 7: 118–127.
Andrew S. FastQC: A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/. Accessed 15 July 2016.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA–Seq. Bioinformatics 2009, 25: 1105–1111.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods 2012, 9: 357–359.
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 2012, 7: 562–578.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015, 43: e47.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26: 139–140.
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, et al. TM4: a free, open–source system for microarray data management and analysis. Biotechniques 2003, 34: 374–378.
Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 2009, 37: 1–13.
Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009, 4: 44–57.
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, et al. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 2006, 34: W293–297.
Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV, et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 2013, 14: 128.
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 2016.
Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res 2015, 43: D447–452.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 2003, 13: 2498–2504.
Horowitz M, Robinson SD. Heat shock proteins and the heat shock response during hyperthermia and its modulation by altered physiological conditions. Prog Brain Res 2007, 162: 433–446.
Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, O’Keeffe S, et al. An RNA–sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J Neurosci 2014, 34: 11929–11947.
Chang CY, Chen JY, Chen SH, Cheng TJ, Lin MT, Hu ML. Therapeutic treatment with ascorbate rescues mice from heat stroke-induced death by attenuating systemic inflammatory response and hypothalamic neuronal damage. Free Radic Biol Med 2016, 93: 84–93.
Biedenkapp JC, Leon LR. Increased cytokine and chemokine gene expression in the CNS of mice during heat stroke recovery. Am J Physiol Regul Integr Comp Physiol 2013, 305: R978–986.
Quinn CM, Audet GN, Charkoudian N, Leon LR. Cardiovascular and thermoregulatory dysregulation over 24 h following acute heat stress in rats. Am J Physiol Heart Circ Physiol 2015, 309: H557–564.
Reaux-Le Goazigo A, Van Steenwinckel J, Rostene W, Melik Parsadaniantz S. Current status of chemokines in the adult CNS. Prog Neurobiol 2013, 104: 67–92.
Miller RJ, Rostene W, Apartis E, Banisadr G, Biber K, Milligan ED, et al. Chemokine action in the nervous system. J Neurosci 2008, 28: 11792–11795.
Rostene W, Kitabgi P, Parsadaniantz SM. Chemokines: a new class of neuromodulator? Nat Rev Neurosci 2007, 8: 895–903.
Du L, Zhang Y, Chen Y, Zhu J, Yang Y, Zhang HL. Role of microglia in neurological disorders and their potentials as a therapeutic target. Mol Neurobiol 2016. doi:10.1007/s12035-016-0245-0.
Ben Haim L, Rowitch DH. Functional diversity of astrocytes in neural circuit regulation. Nat Rev Neurosci 2017, 18: 31–41.
Hu X, Yuan Y, Wang D, Su Z. Heterogeneous astrocytes: Active players in CNS. Brain Res Bull 2016, 125: 1–18.
Acknowledgements
We thank Dr. Qiumin Le (Fudan University, China), Dr. Biao Yan, and Professors Guowei Le and Yonghui Shi (Jiangnan University, China) for technical assistance. This work was supported by the 12th Five-Year Plan of the PLA (BWS11J062), the China Postdoctoral Science Foundation (2015M572806), and the Director’s Fund of the General Hospital of Jinan Military Region, China (2014ZX03).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Niu, B., Zhang, T., Hu, H. et al. Transcriptome Sequencing Reveals Astrocytes as a Therapeutic Target in Heat-Stroke. Neurosci. Bull. 33, 627–640 (2017). https://doi.org/10.1007/s12264-017-0156-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12264-017-0156-8